Validation and clinical utility of epigenetic clocks in 2026
Introduction: The Shift from Chronological to Biological Age
The pursuit of quantifying human aging has undergone a profound paradigm shift over the past decade. Historically, the medical establishment and regulatory bodies have relied almost exclusively on chronological age - the linear accumulation of time since birth - as the primary covariate for modeling disease risk, forecasting mortality, and anticipating physiological decline. However, a wealth of longitudinal gerontological research has decisively demonstrated that chronological age fails to capture the immense inter-individual heterogeneity inherent in functional preservation and disease vulnerability. Two individuals of identical chronological age may exhibit vastly different aging trajectories at the cellular, tissue, and systemic levels due to the complex interplay of genetic endowment, cumulative environmental exposures, lifestyle factors, and socioeconomic determinants. To address this critical gap in clinical prognostication, the field of geroscience has catalyzed the development of biological age biomarkers, designed to quantitatively measure the physiological and molecular degradation that drives the pathogenesis of age-related conditions.
At the absolute forefront of these emerging diagnostics are epigenetic clocks. These sophisticated algorithms leverage genome-wide DNA methylation patterns to estimate an individual's biological age and the underlying physiological pace of aging. DNA methylation, which specifically involves the covalent addition of methyl groups to the 5-carbon position of cytosine residues within CpG dinucleotides, undergoes highly predictable and systemic alterations over the human lifespan. A stereotypic molecular signature of aging emerges through global genomic hypomethylation coupled with localized hypermethylation at specific gene promoter regions. By applying advanced machine learning algorithms - ranging from penalized elastic net regression models to modern, transformer-based neural networks - researchers have successfully distilled the staggering complexity of over a million CpG sites into actionable, quantitative aging scores 12. The economic and scientific implications of these biomarkers are vast, evidenced by the influx of capital into the longevity biotech sector, the public launch of entities like BioAge Labs, and global initiatives such as the XPRIZE Healthspan 34.
As of 2026, the landscape of epigenetic clocks has evolved dramatically from early algorithms that merely estimated chronological age into highly sophisticated, multi-omic predictors of morbidity, mortality, and organ-specific functional decline. Nevertheless, the rapid commercialization of these algorithms by direct-to-consumer longevity and biohacking companies has substantially outpaced rigorous clinical validation. The aggressive marketing of biological age testing kits to the general public has generated widespread consumer misconceptions regarding the biological stability, interpretability, and actionable clinical utility of these tests. This comprehensive report provides an exhaustive analysis of the current state of biological age biomarkers, strictly prioritizing peer-reviewed gerontological literature and consensus statements from major medical and regulatory boards. The analysis rigorously evaluates the predictive power, cross-cultural validity, and regulatory standing of third-generation and newly developed multi-omic epigenetic clocks, while systematically dissecting and clarifying the inherent limitations of consumer-grade epigenetic testing.
The Biomarker Landscape: Contextualizing Epigenetic Clocks Among Multi-Omics
While DNA methylation currently serves as the anchor and gold standard for biological age estimation, the aging process is a highly multifaceted construct that propagates across numerous biological layers. To achieve a nuanced understanding of biological age, it is necessary to contextualize epigenetic clocks alongside alternative and complementary biomarkers, including telomere length, transcriptomics, proteomics, and glycomics. Each omic layer offers a distinct temporal and mechanistic window into physiological decline.
The measurement of leukocyte telomere length was historically considered the primary molecular clock of human aging. Telomeres are the repetitive ribonucleoprotein complexes that cap and protect the ends of chromosomes, progressively shortening with each cellular division until they trigger cellular senescence or apoptosis. While critically short telomeres are implicated in specific telomere biology disorders, average telomere length exhibits a relatively weak and highly variable correlation with chronological age in the general population. Epidemiological data indicates a Pearson correlation coefficient of merely $r \approx -0.51$ in women and $r \approx -0.55$ in men 1. Furthermore, telomere length demonstrates immense inter-individual baseline variability and extremely low predictive power for tracking short-term lifestyle interventions or longitudinal changes in physical function, rendering it significantly less informative as a general health-span tracking metric when compared to advanced DNA methylation clocks 56.
Transcriptomic biomarkers evaluate the dynamic levels of gene expression (mRNA) across various tissues. Because the transcriptome governs immediate cellular function, it is highly sensitive to acute environmental changes, disease states, and circadian rhythms. While this volatility traditionally limited its utility as a stable, long-term marker of aging, recent integrations of transcriptomics with epigenetics have proven highly revelatory. Multi-cohort analyses utilizing the Multi-Ethnic Study of Atherosclerosis (MESA) and the Parkinson's Progression Markers Initiative (PPMI) have identified "multi-omic aging genes." Research demonstrates that genes exhibiting the greatest age-dependent gain in DNA methylation consistently feature a corresponding, significant loss of gene expression 3. These multi-omic markers, which bridge epigenetic regulation and transcriptomic output, show substantially stronger associations with long-term mortality risk than either omic layer evaluated in isolation 3.
The proteome and metabolome offer a real-time, highly granular snapshot of an organism's functional and metabolic state. Aging is universally accompanied by distinct phenotypic shifts in circulating proteins, such as Growth Differentiation Factor 15 (GDF-15) and inflammatory cytokines, as well as distinct metabolic profiles. While powerful for risk stratification, direct proteomic and metabolomic assays are highly susceptible to acute physiological states, including recent dietary intake, acute infections, or transient physical exertion. To circumvent this volatility, cutting-edge epigenetic models increasingly use DNA methylation data to mathematically estimate these proteomic and metabolomic states. This approach combines the inherent biological stability of the epigenome with the immediate phenotypic relevance of the downstream omic layers, creating a highly resilient biomarker profile 249.
Finally, glycomics provides a critical measure of immune-specific aging. Glycan analysis, particularly the profiling of the immunoglobulin G (IgG) glycome, tracks the enzymatic addition of complex sugar moieties to proteins. Glycan markers are robustly linked to immune aging, immune system regulation, and systemic chronic inflammation - a primary driver of biological aging often termed "inflammaging." Glycan-based tests have demonstrated significant responsiveness to sustained, structured lifestyle interventions, such as prolonged weight loss and structured exercise regimens, providing valuable, system-specific insights into immunosenescence that complement broader, genome-wide epigenetic evaluations 5.
The Evolutionary Paradigm of Epigenetic Clocks: Generations I to V
The architecture and mathematical foundation of epigenetic clocks have progressed through distinct generational iterations over the past decade. These generations are defined not merely by the number of CpG sites they analyze, but by their fundamental mathematical training targets and their resulting clinical applicability.
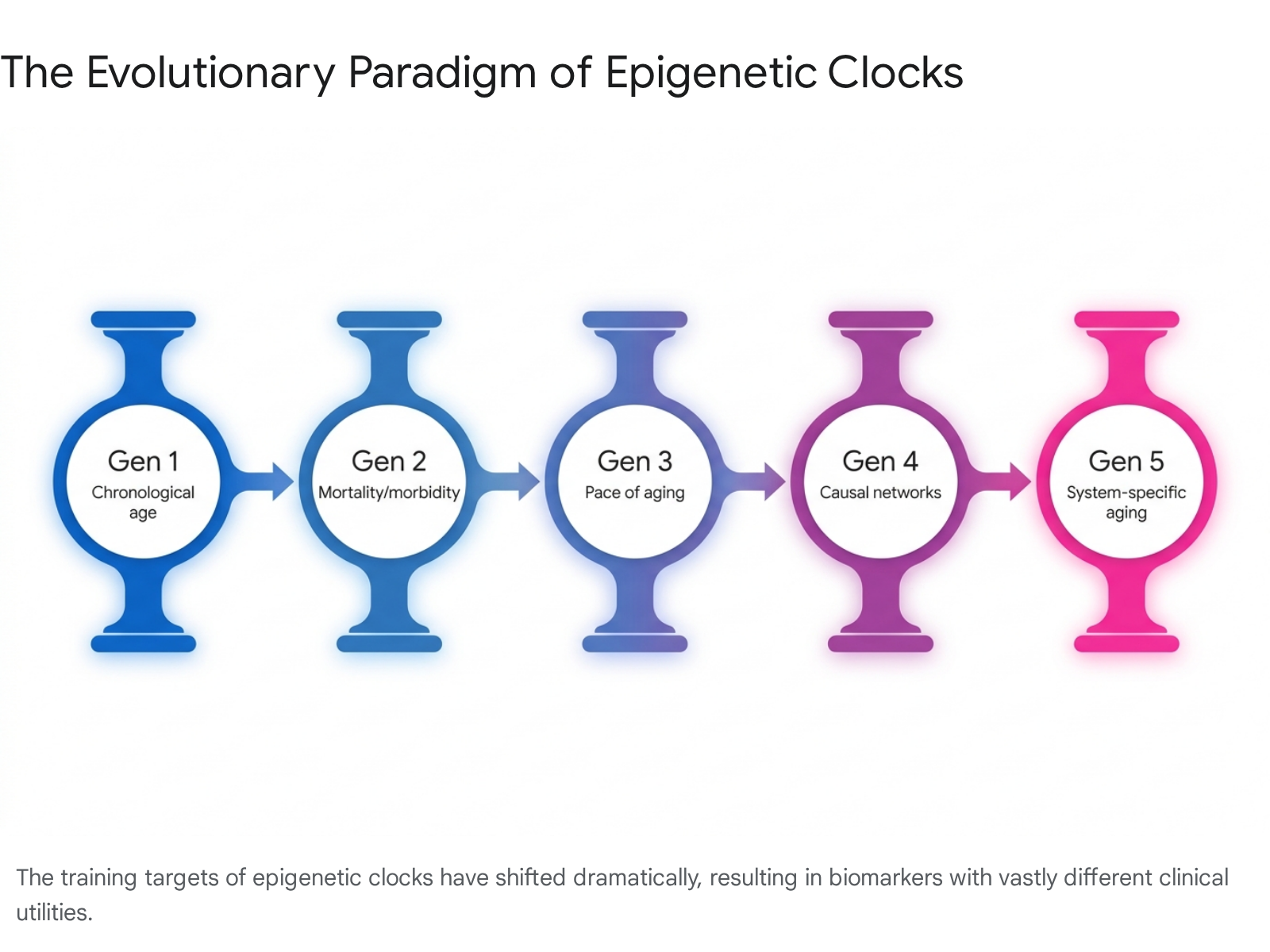
Understanding this evolution is critical to discerning why certain modern clocks are uniquely appropriate for mortality prediction and intervention tracking, while earlier models are entirely unsuited for evaluating anti-aging therapeutics.
First-generation epigenetic clocks, developed primarily between 2013 and 2015, were foundational to the field. Algorithms such as the Horvath pan-tissue clock and the Hannum blood clock were trained using supervised machine learning specifically to predict chronological age 1105. The Horvath clock utilizes a penalized regression model (elastic net regularization) to form a weighted average of 353 distinct CpG sites, achieving a remarkable predictive accuracy across diverse human tissue types, yielding a Pearson correlation coefficient of $r \approx 0.96$ with chronological calendar age 16. However, because these models were mathematically optimized to align perfectly with chronological time, they actively strip away and discard the biological variance that constitutes true aging heterogeneity. Consequently, first-generation models are highly unresponsive to actual lifestyle interventions and fail to adequately predict clinical morbidity or mortality outcomes independently of a patient's chronological age 13147.
Recognizing the profound clinical limitations of pure chronological estimators, researchers pivoted to developing second-generation clocks. These models were explicitly trained against hard phenotypic data, disease incidence, and human lifespan. The DNAm PhenoAge algorithm, for instance, incorporates 513 CpG sites selected specifically for their correlation with nine clinical blood biomarkers and chronological age. In validation studies, a one-year increase in phenotypic age acceleration via PhenoAge is associated with a 9% increase in all-cause mortality risk ($HR=1.09, p=3.8 \times 10^{-49}$) and a 10% increase in cardiovascular mortality risk 7. Building on this, the GrimAge algorithm (versions I and II) identifies DNA methylation surrogates for specific plasma proteins associated with morbidity (e.g., GDF-15, PAI-1, leptin) and integrates smoking pack-year history. GrimAge V2 further refined this accuracy by incorporating epigenetic markers for Hemoglobin A1C and C-reactive protein, cementing it as one of the most powerful predictors of time-to-death, coronary heart disease, and severe age-related morbidity available 5138.
Third-generation clocks represent a fundamental methodological shift from static age estimation to rate-of-change assessment. Rather than providing a single point-in-time biological age (e.g., "this individual is biologically 55 years old"), these models estimate the actual pace at which physiological aging is presently occurring. The flagship third-generation model, DunedinPACE, was derived from the highly detailed Dunedin longitudinal birth cohort. Researchers tracked within-individual physiological decline across 19 distinct organ-system integrity indicators over two decades 91810. Distilled into a single DNA methylation blood test, DunedinPACE outputs a specific rate: a score of 1.0 indicates normal aging (accumulating one biological year per calendar year), while a score of 1.2 indicates an individual is aging 20% faster than average. Because it captures current physiological momentum, DunedinPACE demonstrates exceptional, validated sensitivity to geroprotective interventions, sustained caloric restriction, and the long-term biological effects of early-life adversity, offering unparalleled utility for longitudinal tracking in both research and clinical trial settings 91820.
By 2024 and 2026, the field advanced into fourth and fifth-generation frameworks designed to integrate multi-omic data and deconvolute aging into specific physiological systems. The OMICmAge algorithm utilizes an elastic net regression model that integrates methylation CpG values, immune cell subsets, and 396 Epigenetic Biomarker Proxies (EBPs) derived from extensive metabolomic and proteomic datasets. Evaluating 990 core CpG sites, OMICmAge yields superior 10-year mortality prediction compared to even GrimAge. More importantly, it provides mechanistic transparency regarding why an individual is aging rapidly by highlighting specific downstream metabolite or protein regulatory deficiencies that the methylation patterns represent 2421. Concurrently, the SystemsAge (or SymphonyAge) framework shattered the "single metric" paradigm entirely. It is a biological systems-based clock that measures aging rates across 11 distinct physiological systems - including the brain, heart, inflammation, liver, kidney, and immune system - from a single whole-blood draw 132223. This framework reveals critical within-person heterogeneity, demonstrating that a patient might possess an advanced, high-risk cardiovascular age while maintaining a youthful, resilient immune age.
Table 1: Comparison of Major Epigenetic Clock Models
| Generation | Clock Model | Biological Training Target | Core Utility / Measurement Output | Mortality / Morbidity Predictive Power |
|---|---|---|---|---|
| First | Horvath (Pan-Tissue), Hannum | Chronological Age | Static age estimation across diverse tissues. Poor intervention tracking. | Low to Moderate |
| Second | PhenoAge, GrimAge (v1, v2) | Clinical phenotypes, mortality, plasma proteins | Static age estimation emphasizing disease risk and survival probability. | Very High |
| Third | DunedinPACE | 20-year longitudinal organ system decline | The rate or pace of aging (e.g., 1.05 years/year). Highly sensitive to interventions. | Very High |
| Fourth / Fifth | OMICmAge | Electronic medical records, proteomics, metabolomics | Mechanistic age estimation with downstream biomarker proxy insights. | Exceptional |
| Fourth / Fifth | SystemsAge | 11 specific physiological organ systems | Deconvoluted organ-specific aging rates; captures within-person heterogeneity. | High (System-Specific) |
Longitudinal Validation and the 2026 Predictive Power Landscape
To successfully bridge the formidable gap between academic research discovery and authorized clinical utility, the Biomarkers of Aging Consortium established rigorous standard validation criteria in 2024 and 2025. A defining advancement in this standardization effort is the deployment of the TranslAGE platform and its associated STAR score framework. The TranslAGE platform systematically harmonizes over 179 disparate DNA methylation datasets to evaluate 41 different epigenetic clocks. The STAR framework standardizes the evaluation of these clocks across four critical domains: Stability (robustness to biological and technical noise), Treatment response (sensitivity to aging interventions), Associations (cross-sectional relationships with disease phenotypes), and Risk prediction (predictive power for future functional decline, morbidity, and mortality) 2425.
When subjected to the rigors of the STAR framework and comprehensive analyses of independent longitudinal datasets - such as the Framingham Heart Study, the Normative Aging Study, and the Health and Retirement Study - second- and third-generation clocks consistently and overwhelmingly outperform both chronological age and first-generation models in clinical risk prediction 2910. DunedinPACE, for example, demonstrated independent predictive value for incident morbidity, disability, and mortality over and above the predictive power of GrimAge when applied to the Framingham Heart Study cohort, representing an exceptionally rigorous test of its prognostic capabilities 910. Similarly, the OMICmAge algorithm exhibits a robust, highly reproducible correlation with clinical outcomes, showing an intraclass correlation coefficient (ICC) exceeding 0.99 for technical replicates, alongside hazard ratios for all-cause mortality that significantly exceed baseline PhenoAge and GrimAge metrics 21.
Beyond epidemiological risk prediction, recent prospective clinical trials assessing specific pharmacological and lifestyle interventions have definitively validated the responsiveness of these advanced models. In the landmark CALERIE trial, which evaluated sustained caloric restriction in humans, researchers observed clear effects on pace-of-aging outcomes. While standard DNAm age clocks did not significantly differ between the control and intervention groups, DunedinPACE decreased moderately and significantly in the calorie restriction arm, with effect sizes of approximately $d = -0.29$ at 12 months ($p = 0.0004$) and up to $d = -0.43$ ($p = 1.4 \times 10^{-5}$) among participants achieving higher levels of target restriction 20.
Furthermore, clinical trials evaluating pharmaceutical gerotherapeutics have yielded unprecedented results. The administration of the GLP-1 receptor agonist semaglutide in middle-aged adults significantly and systematically reduced biological age acceleration across multiple advanced models. Over the trial period, semaglutide was associated with a reduction in biological age acceleration as measured by SystemsAge ($-4.17$ years/year, $p = 0.009$), PhenoAge ($-4.90$ years/year, $p = 0.004$), and OMICmAge ($-2.20$ years/year, $p = 0.009$) 11. These trials mark a critical watershed moment in geroscience, providing the first definitive human clinical trial evidence that approved, widely available therapeutics can meaningfully modulate validated epigenetic aging biomarkers across distinct organ systems, particularly demonstrating broad pleiotropic benefits in the inflammatory, cardiovascular, and hepatic system clocks 2211.
Addressing Misconceptions: Reliability, Noise, and the "Age Reversal" Fallacy
The rapid translation of epigenetic clock algorithms from epidemiological research tools into the commercial consumer wellness market has propagated significant and potentially harmful misconceptions. The most prevalent consumer error is the conflation of temporary, acute biomarker fluctuations with genuine, structural biological age reversal.
A critical 2025 longitudinal study examining the reliability of mortality-predictive clocks revealed a stark and counter-intuitive divergence between laboratory measurement precision and actual biological stability within the human body. Data indicates that while nearly all major clocks boast excellent technical reproducibility (Intraclass Correlation Coefficient [ICC] > 0.9) when replicate samples are processed under identical, controlled laboratory conditions, their biological stability over short periods varies wildly. For instance, advanced statistical models like SystemsAge and Principal Component-adjusted clocks (e.g., PCGrimAge) maintain high biological reliability and stability, clustering in the upper percentiles of performance across testing conditions. Conversely, some highly predictive clinical clocks, including standard DunedinPACE and GrimAge V2, exhibit significant biological variance and lower biological ICC following acute, short-term physiological stressors 27.
Following an acute stressor, a single high-fat meal, or transient exposure to high levels of urban pollution, several widely marketed clocks demonstrated dramatic drops in biological reliability, shifting rapidly into "moderate" or "poor" stability ranges within a single 24-hour testing period 27. This immense susceptibility to acute environmental and metabolic perturbations completely undermines the narrative pushed by direct-to-consumer biohacking companies. These companies frequently market unproven longevity supplements or intensive 3-month lifestyle protocols, pointing to a rapid, short-term drop in biological age scores as definitive proof of "age reversal." In reality, because standard algorithms are highly sensitive to systemic inflammation and acute metabolic shifts, a sudden reduction in the score over a few weeks largely reflects a transient decrease in physiological stress or a clearing of systemic inflammation, rather than a fundamental, structural rejuvenation of the cellular epigenome 2712.
Further highlighting this volatility, a study examining the effects of repeated therapeutic plasmapheresis on healthy donors found an unexpected increase in epigenetic aging markers. Across multiple sessions, mixed models indicated significant increases in multiple GrimAge family clocks (approximately $+0.16$ to $+0.26$ years per session) alongside cumulative accelerations in the DunedinPACE algorithm 20. This directly confounds simplistic consumer assumptions that clearing blood plasma inherently makes an organism biologically "younger" on an epigenetic level, emphasizing the complex, often unpredictable response of methylation sites to systemic interventions.
Additionally, epigenetic age must be rigorously understood through the lens of its standard error of measurement (SEM). Even the highest quality research-grade testing can carry an SEM of 2 to 5 years 613. If a consumer receives a test result indicating a biological age of 45, and three months later tests at 42, that 3-year reduction is entirely within the statistical margin of error and standard acute biological variability. It does not constitute proof of therapeutic efficacy. To mitigate this noise, gerontology experts strongly advise utilizing Principal Component (PC) optimized clocks (such as PCPhenoAge, which smooths technical and biological variance) and requiring longitudinal trend analysis over 12 to 24 months to confirm true trajectory modification.
A final pervasive fallacy is the treatment of biological age as a single, monolithic metric. Aging is not biologically uniform across the organism. The successful implementation and validation of the SystemsAge framework definitively proves that the human body ages heterogeneously. A patient may exhibit highly decelerated musculoskeletal aging due to a lifetime of resistance training, yet present simultaneously with highly accelerated inflammatory, metabolic, or neurocognitive aging 1322. Distilling this immense physiological complexity into a single "biological age" number obfuscates actionable clinical targets and provides a dangerously oversimplified view of a patient's true health trajectory.
Cross-Cultural and Genetic Validity: Confronting Eurocentric Biases
As epigenetic clocks transition from exploratory research tools into global clinical trials and population health management systems, the cross-cultural and genetic validity of these algorithms faces intense and necessary scrutiny. The foundational datasets utilized to train the vast majority of first and second-generation clocks - such as the Framingham Heart Study, the Lothian Birth Cohort, and early iterations of the NHANES study - were predominantly populated by individuals of Western, Caucasian, and European descent 930.
This pervasive Eurocentric bias poses a significant risk of systemic miscalculation and algorithmic bias when applying these clocks to globally diverse populations. DNA methylation patterns are heavily influenced by underlying genetics. Recent genomic analyses utilizing LD Score Regression (LDSC) have estimated the single nucleotide polymorphism (SNP)-based heritability for epigenetic age deviation at approximately $h^2 = 0.41$ ($SE = 0.37$) 31. Because common genetic variants account for over 40% of the variance in epigenetic age acceleration, applying algorithms mathematically optimized on European genomic architectures to African, Asian, or Indigenous populations can generate spurious age acceleration artifacts that do not accurately reflect true morbidity risk.
Recent advancements in 2024 and 2025 have actively sought to rectify this data inequity. The training and validation of third- and fifth-generation clocks now frequently leverage highly diverse, multi-ethnic datasets. For instance, the robust OMICmAge and SystemsAge frameworks were trained and validated against the Mass General Brigham (MGB-ABC) Biobank and the Women's Health Initiative (WHI) cohorts, explicitly ensuring representation across broader socioeconomic, racial, and ethnic lines 22123.
Despite these improvements in cohort diversity, clinical validation studies demonstrate that significant biological and socioeconomic disparities persist in the data output. A comprehensive 2026 cross-sectional analysis utilizing the Area Deprivation Index (ADI) demonstrated that neighborhood deprivation and cumulative lifetime adversity are primary drivers of accelerated epigenetic aging across multiple second-generation clocks. Crucially, the study revealed that race moderated the direct association between ADI and epigenetic aging, with significantly stronger adverse effects and higher accelerated aging (via GrimAge and PCPhenoAge) observed among Black participants compared to White participants exposed to similar deprivation levels 32. Furthermore, when clocks are applied cross-culturally across different nations, baseline epigenetic set-points vary widely. This indicates that different clinical thresholds may be required to interpret "accelerated aging" accurately among distinct ancestral groups, underscoring the urgent, unmet need for comprehensive, multi-ethnic, pan-ancestry DNA methylation normative reference databases before these tools can be equitably deployed in global healthcare 33.
Regulatory Consensus and Medical Professional Guidelines
The translation of advanced aging biomarkers from academic laboratories into regulated, actionable clinical settings represents a formidable operational and regulatory hurdle. A fundamental challenge is that aging itself is not classified as a disease by the International Classification of Diseases (ICD); therefore, pharmaceutical therapeutic trials cannot currently utilize "aging" or "healthspan extension" as a primary regulatory indication for drug approval.
Demonstrating that a therapeutic extends a healthy human lifespan inherently requires decades of continuous follow-up - a timescale that renders anti-aging drug development economically unfeasible for the biopharma industry. To solve this critical bottleneck, intensive advocacy efforts throughout 2024 and 2025 pushed the U.S. Food and Drug Administration (FDA) and the European Medicines Agency (EMA) to formally accept validated biomarkers of biological aging as surrogate endpoints in clinical trials 31436.
Catalyzed by the mandates of the FDA User Fee Reauthorization Act and the Food and Drug Omnibus Reform Act of 2022 (FDORA), the FDA and the Reagan-Udall Foundation convened multiple pivotal public workshops between 2024 and 2026. These sessions focused heavily on the advancement of novel surrogate endpoints and the integration of real-world evidence (RWE) in rare disease and oncology drug development, with significant overlap into geroscience 15161718. While the FDA now formally acknowledges the potent epidemiological correlations between epigenetic age acceleration and all-cause mortality, regulators have yet to universally clear epigenetic clocks for direct drug approval in healthy populations 36. The current regulatory consensus strictly stipulates that a true surrogate endpoint must reliably capture the full effect of the treatment on the ultimate clinical outcome (which remains overall survival). Regulatory boards currently advise that while metrics like DunedinPACE, GrimAge, and OMICmAge possess exceptionally high translational potential, their deployment as primary endpoints in phase III trials necessitates further, irrefutable proof that drug-induced modulation of the specific epigenetic clock reliably corresponds to subsequent, hard reductions in morbidity and mortality 364119.
From a direct clinical practice standpoint, medical professional boards view epigenetic clocks with cautious, measured optimism. The American Geriatrics Society (AGS) officially acknowledges that chronological age alone is an entirely inadequate representation of physiological risk and biological resilience, particularly in high-stakes clinical scenarios such as oncology, where understanding a patient's true vulnerability is critical for treatment dosing and surgical clearance 2021.
However, the 2025 and 2026 AGS consensus statements and clinical practice guidelines emphasize that, outside of controlled research settings, epigenetic clocks currently lack the standardization required for routine point-of-care clinical decision-making 4522. The geriatric medicine community already utilizes a suite of highly validated, highly predictive functional assessments - such as frailty indices, comprehensive geriatric assessments (CGA), and simple grip strength tests. These traditional phenotypic markers cost virtually nothing to administer, yet they strongly and reliably predict mortality, functional decline, and intensive care unit outcomes independently of chronological age 623.
The primary barrier to widespread clinical adoption identified by medical boards is actionable clinical utility 252425. If a 60-year-old patient receives a commercial epigenetic test result indicating an epigenetic age of 68, the attending physician must be able to utilize that specific data point to alter a treatment plan safely and effectively. While newly developed multi-system clocks like SystemsAge are beginning to provide targeted pathways (e.g., identifying isolated hepatic or renal aging), the sheer lack of interoperable electronic health record (EHR) integration, the absence of established clinical decision support algorithms, and the lack of harmonized laboratory processing standards limit their immediate utility for the average clinician 45. Thus, while biological age testing holds profound promise for the future of precision longevity medicine and clinical trial risk stratification, the prevailing professional medical consensus dictates that it should supplement - rather than replace - traditional physiological and geriatric evaluations in current clinical practice.
Commercial Direct-to-Consumer (DTC) vs. Research-Grade Diagnostics
The explosion of the direct-to-consumer longevity market has rapidly democratized public access to biological age testing. However, the chasm in quality, analytical validity, and interpretative rigor between a $150 DTC mail-in kit and a highly controlled, clinical research-grade assay is profound and clinically highly significant.
Research-grade diagnostic protocols - utilized in landmark epidemiological cohorts like the Framingham Heart Study, the MGB Biobank, or the CALERIE trial - rely almost exclusively on venous whole blood draws or sorted peripheral blood mononuclear cells (PBMCs). These samples are analyzed via comprehensive, high-density microarrays (such as the Illumina Infinium MethylationEPIC v2.0 array), which are capable of capturing and analyzing nearly 1 million distinct CpG sites across crucial regulatory regions of the genome 350. In stark contrast, to minimize logistical costs and maximize user convenience, many DTC testing companies utilize unassisted saliva samples or dried blood spots, analyzing them via proprietary, heavily reduced CpG targeting panels that sometimes evaluate fewer than 2,000 sites 135126.
Saliva exhibits a substantially higher degree of cellular heterogeneity - containing unpredictable ratios of buccal epithelial cells to varying leukocyte concentrations - than venous blood. Because DNA methylation is inherently cell-type specific, this high cellular variability in saliva results in lower measurement stability, increased technical noise, and severely diminished validation against hard clinical endpoints like long-term mortality 262754.
Furthermore, DTC platforms frequently employ older, less responsive algorithms (such as the first-generation Horvath or Hannum chronological clocks) simply because they are open-source and free from the complex commercial licensing barriers associated with advanced models. They utilize these outdated clocks despite overwhelming peer-reviewed evidence confirming their inability to track lifestyle interventions accurately 1314. Ultimately, the clinical value of biological age testing relies entirely upon the interpretation of the data by a longevity-literate physician capable of differentiating genuine systemic disease risk from transient epigenetic noise, an interpretative layer that DTC algorithmic wellness reports completely lack.
Table 2: Technical Comparison of DTC Commercial Tests vs. Research-Grade Diagnostics
| Diagnostic Parameter | Direct-to-Consumer (DTC) Biohacking Tests | Research-Grade Clinical Diagnostics |
|---|---|---|
| Primary Biospecimen | Often Saliva or patient-administered dried blood spot. | Venous whole blood, sorted immune cells, or targeted tissue biopsies. |
| Cellular Heterogeneity | High variability; poor control over buccal vs. leukocyte ratios in saliva. | Highly controlled; utilizes advanced deconvolution algorithms to adjust for immune cell subset proportions. |
| Array / Sequencing Depth | Often proprietary, restricted targeted panels (frequently evaluating <2,000 CpGs). | Comprehensive microarrays (e.g., Illumina EPIC v2), covering ~935,000 CpGs across regulatory regions. |
| Clock Algorithms Utilized | Frequently relies on Gen 1 clocks (Horvath/Hannum) or unvalidated proprietary "black-box" models. | Utilizes advanced, extensively peer-reviewed models (DunedinPACE, GrimAge V2, OMICmAge, SystemsAge). |
| Standard Error of Measurement | High (Often $\pm 3$ to $5$ years); high susceptibility to acute biological noise. | Optimized via Principal Component (PC) analysis (e.g., PC-GrimAge) to ensure high Intraclass Correlation ($ICC > 0.9$). |
| Clinical Actionability | Low to Moderate. Reports generate automated wellness and generic supplement recommendations. | High. Interpreted by physicians to optimize therapies, monitor off-target toxicity, and stratify mortality risk. |
| Regulatory Status | Unregulated wellness products; explicitly not intended to diagnose or treat disease. | Subject to rigorous clinical validation; increasingly utilized as exploratory/surrogate endpoints under FDA/EMA guidelines. |
Conclusion
The science of biological age quantification has decisively transitioned from an academic proof-of-concept into a realm of highly robust, multi-dimensional predictive modeling. In 2026, the vanguard of epigenetic analysis - led by third-generation pace-of-aging metrics like DunedinPACE, and highly sophisticated, systems-level frameworks like OMICmAge and SystemsAge - demonstrates unparalleled, validated power in predicting human morbidity, physical decline, and mortality. These advanced mathematical models transcend the profound limitations of early chronological estimators by capturing the authentic physiological wear and tear driven by genomic instability, chronic inflammaging, and systemic metabolic dysregulation.
However, the rapid and successful clinical translation of these powerful tools is actively hindered by the commercial proliferation of oversimplified, direct-to-consumer testing. The public and commercial conflation of short-term biological noise with authentic, structural epigenetic age reversal threatens to severely undermine the scientific credibility of the field. As clearly demonstrated by the stark discrepancy between technical laboratory precision and biological stability within the human body, human epigenomes are highly sensitive to acute environmental and metabolic stressors. This biological reality necessitates the use of rigorous longitudinal tracking and Principal Component-optimized algorithms to isolate genuine geroprotective signals from daily physiological noise.
For epigenetic clocks to fulfill their ultimate potential as the definitive biomarker of preventative precision medicine and longevity research, the medical and scientific communities must aggressively continue to champion rigorous standardization. This requires the immediate establishment of diverse, pan-ancestry normative databases to eradicate inherent ethnic and genetic biases, the continued, evidence-based advocacy for FDA recognition of surrogate aging endpoints in clinical trials, and the seamless integration of biological age diagnostics into the existing, highly validated framework of evidence-based geriatric care. When anchored firmly in rigorous science and interpreted by skilled, longevity-literate clinicians, advanced epigenetic clocks offer a profound, unprecedented, and data-driven window into the physiological trajectory of human health and the fundamental mechanisms of aging.