Transposable elements in somatic aging and inflammation
Transposable elements (TEs) comprise nearly half of the mammalian genome. Historically characterized as superfluous sequences or "junk DNA," these mobile genetic units are now recognized as critical determinants of both long-term evolutionary adaptation and the physiological decline associated with organismal aging 123. Under normal physiological conditions in young, healthy cells, TEs are strictly repressed by multiple layers of epigenetic silencing, including DNA methylation and the formation of constitutive heterochromatin 45. However, the integrity of this silencing network deteriorates over the mammalian lifespan 67.
The age-related derepression of TEs, particularly Long Interspersed Nuclear Element-1 (LINE-1), results in the aberrant transcription and subsequent reverse transcription of these elements within somatic cells 78. The accumulation of TE-derived nucleic acids in the cytoplasm triggers antiviral innate immune sensors, primarily the cyclic GMP-AMP synthase-stimulator of interferon genes (cGAS-STING) pathway, leading to the secretion of type I interferons (IFN-I) and pro-inflammatory cytokines 7910. This phenomenon establishes a state of chronic, sterile inflammation commonly referred to as "inflammaging," which acts as a primary driver of age-related neurodegenerative and metabolic diseases 111213.
Classification and Evolutionary Dynamics of Transposable Elements
Transposable elements are DNA sequences capable of moving or copying themselves from one genomic location to another, independently of the host genome 1. Originally identified in maize by Barbara McClintock more than 60 years ago, TEs represent approximately 41% of the mouse genome and 48% of the human genome, a striking proportion compared to the 1.5% occupied by protein-coding sequences 1.
DNA Transposons and Retrotransposons
Transposable elements are categorized into two primary classes based on their mechanism of transposition 14. Class II elements, or DNA transposons, move via a "cut-and-paste" mechanism encoded by transposase enzymes 114. Class I elements, or retrotransposons, utilize a "copy-and-paste" mechanism involving an RNA intermediate 414.
Retrotransposons are the dominant class in humans and are further subdivided into Long Terminal Repeat (LTR) retrotransposons and non-LTR retrotransposons 47. LTR retrotransposons, such as human endogenous retroviruses (HERVs), integrate into the genome using mechanisms homologous to exogenous retroviruses 47. Non-LTR retrotransposons consist of Long Interspersed Nuclear Elements (LINEs) and Short Interspersed Nuclear Elements (SINEs) 47. LINE-1 is the most abundant autonomous retrotransposon family, encoding its own reverse transcriptase and endonuclease proteins 47. SINEs, such as Alu elements, are non-coding RNAs that lack their own enzymatic machinery and must hijack the proteins encoded by LINE-1 to achieve retrotransposition 47.
| Element Class | Transposition Mechanism | Key Subtypes in Humans | Autonomous Capability |
|---|---|---|---|
| Class I (Retrotransposons) | "Copy-and-paste" via RNA intermediate | LTRs (HERVs), Non-LTRs (LINEs, SINEs) | LINEs (Autonomous); SINEs (Non-autonomous) 47 |
| Class II (DNA Transposons) | "Cut-and-paste" via DNA intermediate | DNA Transposons | Dependent on functional transposase 114 |
Evolutionary Utility and Genomic Plasticity
While the mobilization of TEs in somatic tissues poses a severe threat to genomic integrity, their long-term activity in the germline has been a major engine of mammalian evolution and genomic plasticity 115. TEs contribute to genetic variability through multiple mechanisms. Recombination between repetitive TE sequences can generate large structural genomic rearrangements, including deletions, duplications, and chromosomal inversions 116.
Beyond passive structural disruption, TEs are frequently subjected to a process known as molecular domestication or "exaptation," wherein the host organism co-opts TE-derived sequences for critical biological functions 15. The RAG1 gene, which is essential for V(D)J somatic recombination in the adaptive immune system of jawed vertebrates, evolved through the domestication of an ancient DNA transposase known as Transib 15. Similarly, Syncytin genes, which are indispensable for placental development in mammals, derive from the envelope genes of endogenous retroviruses 1. The Arc gene, critical for neuronal plasticity and brain development, also shares a retrotransposon origin 1.
TEs further serve as abundant sources of cis-regulatory elements 116. They donate promoters, enhancers, and transcription factor binding sites that can rewire host gene regulatory networks 116. They also influence the three-dimensional architecture of chromatin, acting as insulator elements or demarcating the boundaries of topologically associating domains (TADs) 1. Under environmental stress, the transient mobilization of TEs can induce rapid genetic variability, an adaptive response observed in species such as the root-knot nematode Meloidogyne incognita, which utilizes TE movements to maintain genome plasticity and adapt to host resistance despite reproducing without sexual recombination 1718.
However, this evolutionary utility carries a profound physiological cost. The mutagenic potential of TEs necessitates highly restrictive suppression mechanisms in somatic tissues to prevent premature cellular senescence, oncogenesis, and tissue dysfunction 141920.
Epigenetic Silencing Mechanisms in Somatic Cells
To mitigate the deleterious effects of TEs, eukaryotic cells employ comprehensive epigenetic surveillance networks designed to maintain these repetitive elements in a transcriptionally inert state. During early embryonic development and throughout the somatic lifespan of young organisms, TE loci are heavily methylated and densely packed into constitutive heterochromatin 456.
DNA Methylation and Heterochromatin Maintenance
The basal suppression of TEs is enforced by DNA methylation at cytosine residues within the CpG islands of TE promoters 1419. This modification serves as a fundamental repressor of transcription 1921. In parallel with DNA methylation, specific histone modifications dictate the physical compaction of the surrounding chromatin structure 14.
The trimethylation of histone H3 at lysine 9 (H3K9me3) is the biochemical hallmark of constitutive heterochromatin 5. This modification serves as a critical docking site for Heterochromatin Protein 1 (HP1) isoforms (HP1α, HP1β, and HP1γ) 5. Upon binding to H3K9me3, HP1 induces high-order chromatin compaction, thereby restricting the access of RNA polymerase and host transcription factors to the underlying TE sequences 5.
The precision of these repressive marks relies on the Krüppel-associated box zinc-finger protein (KRAB-ZFP) family and its primary corepressor, KAP1 (also known as TRIM28) 622. Sequence-specific KRAB-ZFPs recognize distinct transposable element motifs across the genome and recruit KAP1 to the locus 22. KAP1 functions as a molecular scaffold, drawing in histone methyltransferases, such as SETDB1 and SUV39H1, alongside histone deacetylases and HP1, to establish a localized, impenetrable heterochromatic domain 52223.
The Regulatory Role of Sirtuins in Chromatin Condensation
The NAD+-dependent class III histone deacetylases known as sirtuins (SIRT1-7) act as central coordinators linking cellular metabolism, DNA repair, and epigenetic regulation 2425. Among the mammalian isotypes, SIRT1 and SIRT6 are the most extensively characterized regarding their roles as epigenetic gatekeepers of transposable elements and overall genome stability 252627. Both enzymes reside predominantly in the nucleus, where they maintain heterochromatin compaction through distinct but synergistic biochemical pathways 2528.
Sirtuin 6 (SIRT6)
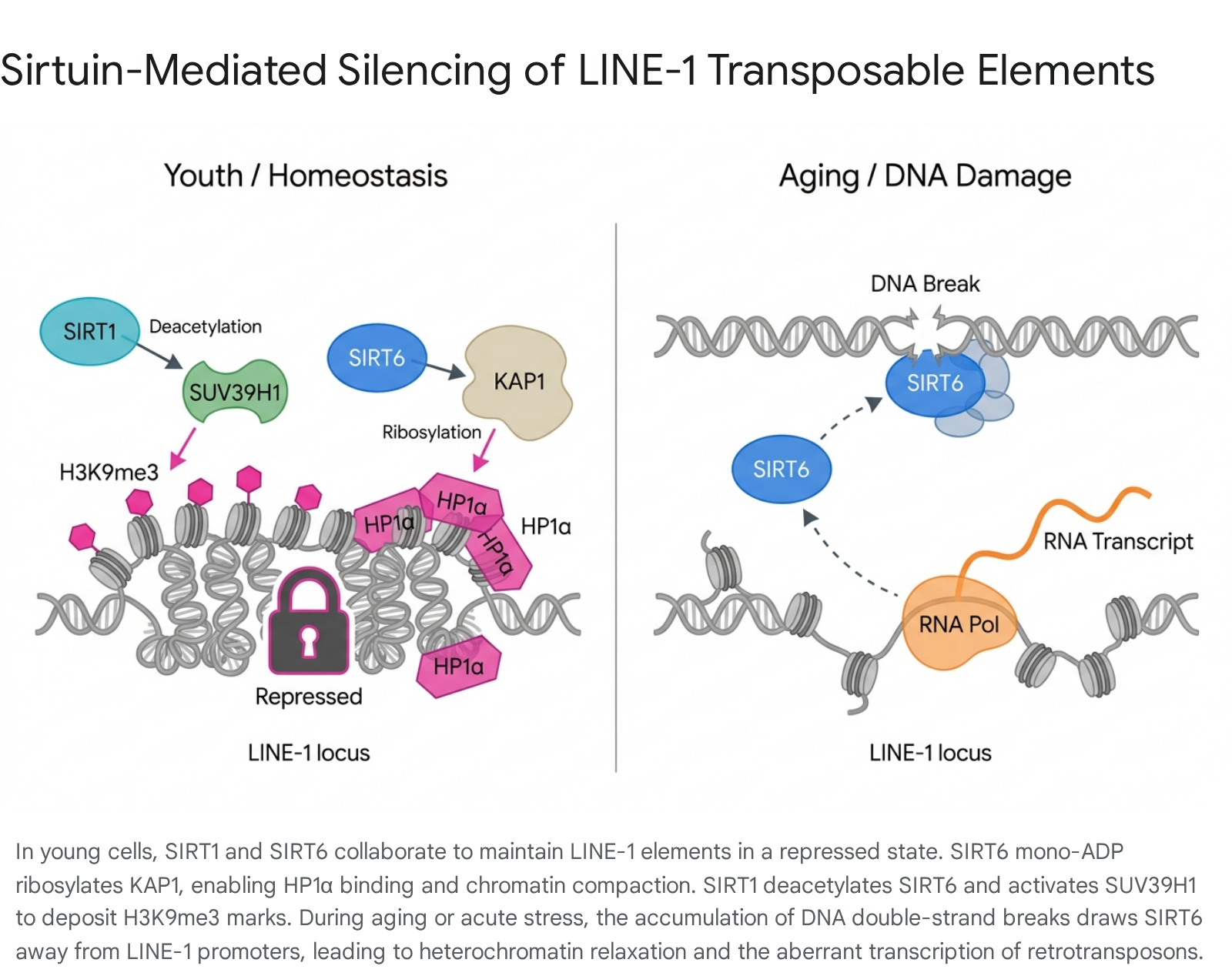
SIRT6 functions as a critical, direct suppressor of LINE-1 retrotransposons 2229. At the molecular level, SIRT6 is physically tethered to the 5' untranslated region (5'UTR) of LINE-1 loci 2230. While SIRT6 possesses histone deacetylase activity - specifically targeting H3K9ac and H3K56ac to promote chromatin condensation - its role in suppressing LINE-1 is heavily mediated by its mono-ADP ribosyltransferase activity 223132. SIRT6 mono-ADP ribosylates the KAP1 corepressor protein at these repetitive loci 22. This specific post-translational modification facilitates the stable interaction between KAP1 and HP1α, ensuring the packaging of LINE-1 elements into transcriptionally repressive heterochromatin 222933. Furthermore, SIRT6 regulates telomeric chromatin by deacetylating H3K56ac, demonstrating a broad role in genomic stability maintenance 3132.
The loss of SIRT6 leads to catastrophic genomic consequences. SIRT6-deficient murine models exhibit a severe progeroid (premature aging) phenotype, characterized by loss of subcutaneous fat, kyphosis, severe metabolic defects, genomic instability, and a markedly shortened lifespan 2233. In the absence of SIRT6, KAP1 and HP1α fail to stably associate with LINE-1 promoters, resulting in the localized depletion of H3K9me3 and hypomethylation of the CpG island within the L1 5'UTR 22. This triggers massive transcriptional derepression of LINE-1, culminating in high levels of cytoplasmic LINE-1 cDNA, DNA double-strand breaks, and severe systemic inflammation 333435. Interestingly, the modulation of specific phosphorylation sites on SIRT6 highlights the delicate balance between its functions; the phosphomimetic SIRT6 S10E mutation enhances DNA repair capacity but impairs LINE-1 silencing, ultimately leading to a shortened lifespan, indicating that loss of TE control is a dominant driver of mortality 30.
Sirtuin 1 (SIRT1)
SIRT1 shares SIRT6's capacity to deacetylate H3K9 and H3K56, modifications that antagonize chromatin relaxation 242536. SIRT1 actively drives heterochromatin formation by modifying the histone methyltransferase SUV39H1 23. By deacetylating SUV39H1 at lysine 266 (K266) within its catalytic domain, SIRT1 increases the enzyme's methyltransferase activity, directly promoting the trimethylation of H3K9 23. Furthermore, SIRT1 prevents the degradation of SUV39H1 by inhibiting its polyubiquitination via the MDM2 pathway 23. Additionally, SIRT1 decreases the activity of the p300 histone acetyltransferase, further favoring a repressive chromatin state 23.
Beyond targeting histones and methyltransferases directly, SIRT1 acts as an upstream biochemical regulator of SIRT6 3637. SIRT1 physically interacts with SIRT6 and deacetylates it at lysine 33 (K33) 3637. This specific deacetylation event promotes the polymerization of SIRT6 and is absolutely required for the rapid mobilization and retention of SIRT6 at sites of DNA double-strand breaks (DSBs) 3637. Hypoacetylated SIRT6 anchors to γH2AX at damage sites, allowing subsequent chromatin remodeling 3637. Therefore, SIRT1 and SIRT6 operate sequentially and synergistically to safeguard genomic integrity 36.
| Feature | SIRT1 | SIRT6 |
|---|---|---|
| Primary Cellular Localization | Nucleus (can shuttle to cytoplasm under stress) 2425 | Nucleus (chromatin-associated) 2425 |
| Histone Targets | H3K9ac, H3K14ac, H3K56ac, H4K16ac, H1K26 24 | H3K9ac, H3K56ac, H3K18ac 253132 |
| Non-Histone Heterochromatin Targets | SUV39H1 (activation via deacetylation), SIRT6 (deacetylation at K33), p300 (inhibition) 2336 | KAP1 (mono-ADP ribosylation) 2229 |
| Mechanisms of TE Silencing | Promotes H3K9me3 deposition via SUV39H1 activation; activates SIRT6 polymerization 2336 | Ribosylates KAP1 to recruit HP1α; directly deacetylates H3K9/H3K56 at L1 loci 222932 |
| Loss-of-Function Consequences | Defective DNA damage response, impaired SUV39H1 stabilization, cellular senescence 233638 | L1 derepression, heterochromatin failure, severe DNA damage, premature aging syndrome (progeria) 223334 |
Mechanisms of Age-Associated Transposable Element Derepression
Aging is fundamentally accompanied by a progressive erosion of the epigenome. Global DNA hypomethylation and the redistribution of heterochromatin-associated proteins leave large segments of the genome vulnerable to transcriptional activation 719. This epigenetic breakdown is not uniform; repetitive elements, including TEs, are preferentially affected by age-related hypomethylation and chromatin relaxation 1939. Evidence from the Dog Aging Project, for example, demonstrated that LINE-1 elements were the most frequently hypomethylated regions with age, accounting for 40% of all age-associated hypomethylated loci 39.
The Relocation of Chromatin Modifiers Hypothesis
A leading mechanism for this targeted epigenetic failure is the "relocation of chromatin modifiers" hypothesis 2235. Throughout the mammalian lifespan, somatic cells accumulate persistent DNA damage, particularly double-strand breaks, due to oxidative stress, metabolic byproducts, and environmental mutagens 3436. In response to this genomic damage, sirtuins - especially SIRT6 - are physically recruited away from their native heterochromatic loci, including LINE-1 promoters, and mobilize to the sites of DNA damage to facilitate rapid repair alongside factors like γH2AX, SNF2H, and PARP1 2236.
While this rapid mobilization is essential for immediate cell survival and genome repair, the chronic nature of DNA damage in aging tissues results in the sustained depletion of SIRT6 from TE loci 22. Deprived of SIRT6-mediated KAP1 ribosylation and histone deacetylation, the constitutive heterochromatin surrounding LINE-1 elements begins to unravel 2233. The 5'UTR of the LINE-1 element becomes physically accessible, leading to the robust transcription of LINE-1 RNA 722. The RNA is subsequently exported to the cytoplasm, where the LINE-1 encoded ORF2p protein utilizes its inherent reverse transcriptase activity to generate abundant copies of cytosolic LINE-1 cDNA 78. Additionally, transcription and transposition are further exacerbated by senescence-related chromatin reorganization mediated by lamin B 3440.
Innate Immune Sensing and the Inflammaging Cascade
The aberrant accumulation of LINE-1 cDNA and other TE-derived nucleic acids in the cytoplasm is a catastrophic event for cellular homeostasis. Mammalian cells are equipped with highly sensitive, germline-encoded pattern recognition receptors (PRRs) that evolved to detect microbial and viral pathogens 941. Because retrotransposons share a deep evolutionary lineage with exogenous retroviruses, their nucleic acid products are virtually indistinguishable from active viral infections by the host's immune surveillance machinery 74344. The sensing of TE nucleic acids by PRRs triggers a profound inflammatory response, providing a direct molecular link between epigenetic erosion and systemic aging.
The cGAS-STING Signaling Pathway
The primary innate immune sensor responsible for detecting cytosolic LINE-1 cDNA is the cyclic GMP-AMP synthase (cGAS) 1242.
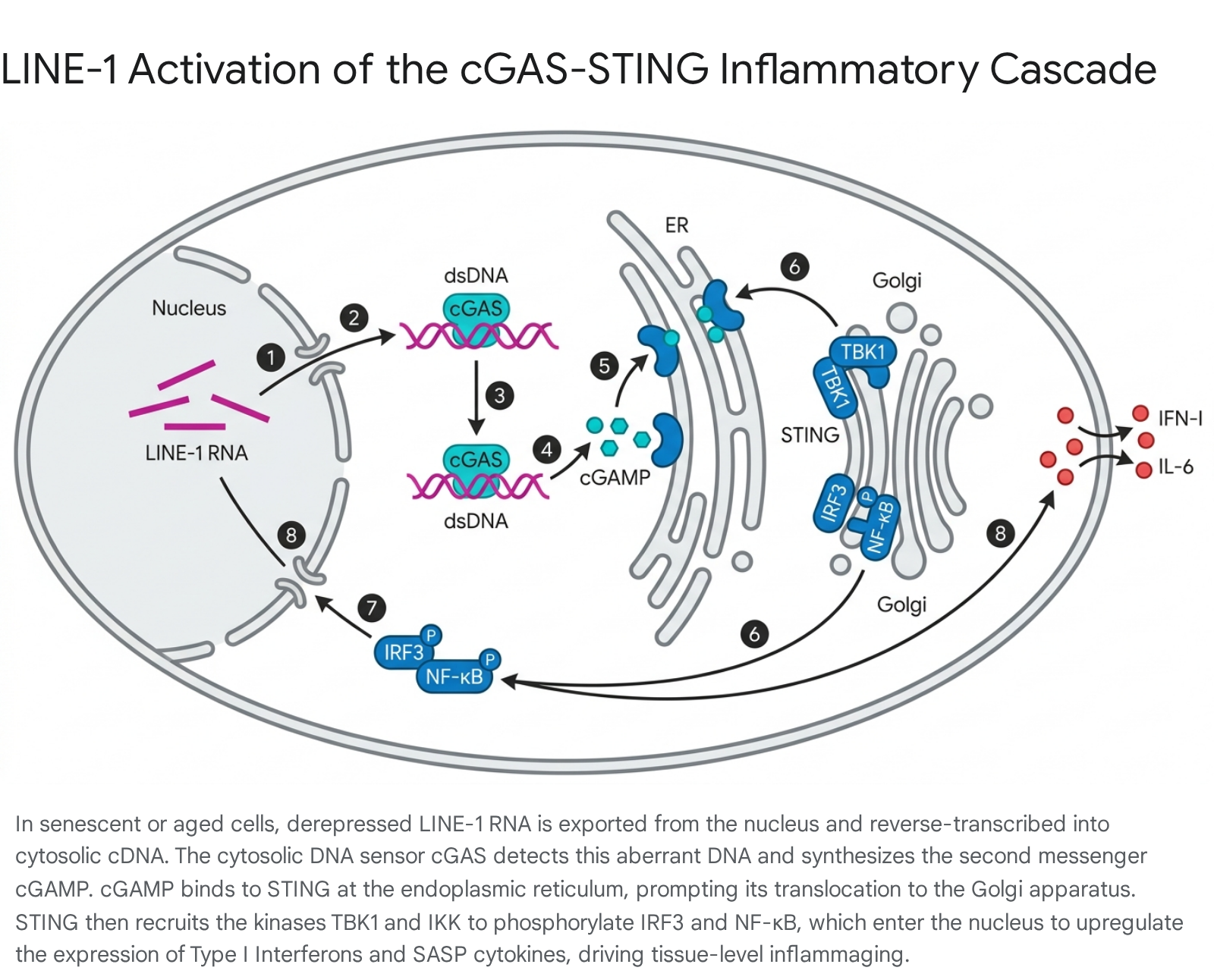
Under normal conditions, DNA is strictly confined to the nucleus and mitochondria 9. The presence of double-stranded DNA (dsDNA) or RNA:DNA hybrids in the cytoplasm signals acute cellular damage, mitochondrial leakage, or viral invasion 913.
The activation cascade proceeds through several highly coordinated spatial and biochemical steps:
- cGAS Activation: When cGAS binds to cytosolic TE-derived dsDNA, it interacts via positively charged amino acid residues with the negatively charged DNA phosphate backbone 912. cGAS dimerizes and forms DNA-linked liquid-liquid phase separations (liquid droplets) 91243. This conformational shift opens a catalytic nucleotide-binding pocket.
- Synthesis of cGAMP: The activated cGAS utilizes local ATP and GTP to synthesize 2'3'-cyclic GMP-AMP (cGAMP), a potent second messenger molecule 912.
- STING Activation: cGAMP binds directly to the Stimulator of Interferon Genes (STING), a transmembrane receptor residing in the endoplasmic reticulum (ER) 944.
- Translocation and Kinase Recruitment: Upon binding cGAMP, STING undergoes dimerization and extensive post-translational modifications. It translocates from the ER, through the ER-Golgi intermediate compartment via COPII vesicles, to the Golgi apparatus 1043. At the Golgi, STING recruits TANK-binding kinase 1 (TBK1) and IκB kinase (IKK) 1243.
- Transcriptional Cascade: TBK1 phosphorylates the transcription factor Interferon Regulatory Factor 3 (IRF3), promoting its dimerization, while IKK phosphorylates the NF-κB inhibitor IκBα, leading to the activation of the NF-κB transcription factor 943. Both activated IRF3 and NF-κB translocate into the nucleus 43. After completing its signaling role, STING is phosphorylated by ULK1/ATG1 and translocated to the lysosome for degradation via clathrin-coated vesicles (CCV) 12.
- Cytokine Secretion: In the nucleus, IRF3 drives the massive transcription of type I interferons (IFN-I), while NF-κB drives the expression of pro-inflammatory cytokines such as interleukin-6 (IL-6), IL-8, and tumor necrosis factor-alpha (TNF-α) 74245.
Secondary Sensing via RNA Receptors
While cGAS-STING is the dominant sensor for cytosolic DNA, the extensive transcription of varied TE families generates diverse nucleic acid species, including double-stranded RNA (dsRNA) and single-stranded RNA (ssRNA), which activate parallel innate immune pathways.
- RIG-I and MDA5: The retinoic acid-inducible gene I (RIG-I)-like receptors (RLRs), which include RIG-I and Melanoma Differentiation-Associated protein 5 (MDA5), are primary cytosolic RNA sensors composed of ATP-dependent DExD/H box RNA helicases 4146. Transposable elements, particularly Alu elements, are prone to forming highly structured, double-stranded RNA hairpins when transcribed 1. If these transcripts evade host RNA editing mechanisms (such as ADAR1), the resulting dsRNA binds to MDA5 (for long dsRNA >1Kbp) and RIG-I (for short stretches <300bp) 4546. Activation of RLRs engages the Mitochondrial Antiviral-Signaling (MAVS) protein, converging on the same IRF3 and NF-κB transcription factors utilized by STING to amplify IFN-I and IL-6 production 454748. Experimental evidence indicates that RIG-I deficiency can induce premature aging phenotypes in mice, suggesting complex regulatory roles for this receptor in cellular senescence 49.
- Toll-Like Receptors (TLRs): In cells with endosomal sensing capabilities, such as microglia in the central nervous system and circulating macrophages, TEs can trigger Toll-Like Receptors 4146. TLR3 recognizes dsRNA and activates a TRIF-dependent signaling pathway 4145. Meanwhile, TLR7 and TLR8 recognize ssRNA 41. Transcripts from the Human Endogenous Retrovirus (HERV-K) subfamily have been explicitly shown to bind TLR7 in murine models and TLR8 in human cells, triggering neurodegenerative and neuroinflammatory cascades 146.
The Senescence-Associated Secretory Phenotype (SASP)
The chronic activation of these innate immune pathways in aged tissues sustains the Senescence-Associated Secretory Phenotype (SASP) 4811. Senescent cells expressing SASP secrete a continuous barrage of inflammatory mediators that degrade local tissue environments via autocrine and paracrine signaling 450. This inflammatory storm inhibits local stem cell function, induces pathological cell proliferation or apoptosis, and promotes premature senescence in neighboring healthy cells, fueling a vicious cycle of tissue deterioration 45152. In the brain, plasma extracellular vesicle (EV)-derived LINE-1 RNA has been identified as a potent systemic aging factor capable of penetrating the blood-brain barrier, delivering LINE-1 RNA to microglia, initiating cGAS-STING signaling, and driving cognitive impairment 43.
Pharmacological Interventions Targeting Reverse Transcriptase
The realization that transposable elements fuel biological aging through viral-like molecular mechanisms has generated significant interest in repurposing antiviral drugs as gerotherapeutics. Because LINE-1 elements and endogenous retroviruses rely absolutely on reverse transcriptase (RT) to convert their RNA transcripts into the immunostimulatory cDNA that triggers the cGAS-STING cascade, inhibiting this enzyme blocks the upstream catalyst of inflammaging 68.
Nucleoside Reverse Transcriptase Inhibitors (NRTIs)
Nucleoside Reverse Transcriptase Inhibitors (NRTIs) are a well-established class of drugs originally developed and FDA-approved for the treatment of Human Immunodeficiency Virus (HIV) and Hepatitis B 1353. These agents act as competitive substrate inhibitors of viral reverse transcriptases, incorporating into the nascent DNA chain and causing premature chain termination 53. Preclinical and clinical evidence now demonstrates that these compounds effectively cross-react to inhibit LINE-1 RT activity, significantly reducing cytoplasmic DNA accumulation and attenuating sterile inflammation 7844. Furthermore, NRTIs have been shown to possess intrinsic anti-inflammatory properties by directly inhibiting inflammasome activation independently of their antiviral activity 133944.
In preclinical murine models, NRTI therapy has shown remarkable efficacy in rescuing age-related physiological deterioration. Treatment of SIRT6-knockout mice - which suffer from massive LINE-1 derepression - with the NRTI lamivudine (3TC) successfully abrogated the type I interferon response, significantly reduced DNA damage markers (such as γH2AX), and improved overall tissue health and lifespan, though it did not completely rescue the maximum lifespan limit 83335. Similar improvements in spatial memory and reductions in age-related weight loss have been observed in senescence-prone (SAMP8) mice treated with NRTIs 54.
Human Clinical Trials and Epigenetic Clock Reversal
Translating NRTIs to human longevity medicine is currently underway, supported by epidemiological data indicating that HIV-positive patients chronically treated with NRTIs experience a significantly reduced incidence of Alzheimer's disease and type 2 diabetes compared to those on non-NRTI antiretroviral regimens 1344. While large-scale, long-term longevity trials remain pending, two notable clinical studies have provided robust initial proof-of-concept in humans.
- Lamivudine (3TC) in Alzheimer's Disease: A Phase 2a, open-label clinical pilot study conducted at UT Health San Antonio investigated the safety and efficacy of lamivudine in 12 older adults (ages 52 to 83) with mild cognitive impairment 555657. Over a six-month treatment period, the drug was well-tolerated and was detected in the cerebrospinal fluid (CSF) 58. Post-treatment analysis revealed a reduction in CSF biomarkers associated with neuroinflammation and astrocyte activation, specifically Flt1 and GFAP, while plasma Aβ42/40 ratios nudged upward, suggesting a potential reduction in brain amyloid pathology 5558. While markers like neurofilament light and phosphotau-181 did not change, the study provides preliminary evidence that suppressing retrotransposons can safely modulate neurodegeneration 58.
- Emtricitabine/Tenofovir Alafenamide (FTC/TAF) and Epigenetic Aging: A groundbreaking 2024 human trial evaluated healthy adults (ages 18 - 50) without HIV or chronic comorbidities taking FTC/TAF (commercially known as Descovy) over a 12-week period 5462. The results indicated that the FTC/TAF regimen significantly reversed multiple epigenetic clocks. Most notably, the PhenoAge marker was reduced by an average of 6.33 years, and the DunedinPACE clock - which measures the current pace of biological aging - slowed significantly (-0.061) 546259. Furthermore, systemic proxies of inflammation, including DNAm-inferred Interleukin-6 (IL-6), showed significant reductions 62. Interestingly, a parallel cohort utilizing the older FTC/TDF (Tenofovir Disoproxil Fumarate) formulation showed no significant changes across epigenetic clocks 62. This discrepancy highlights that the superior intracellular penetration of TAF in immune cells is required to effectively halt TE activity, whereas TDF fails to reach the necessary therapeutic concentrations 5459.
| Clinical Trial / Study | NRTI Regimen | Study Population | Key Findings and Biomarker Shifts |
|---|---|---|---|
| UT Health San Antonio Phase 2a 555758 | Lamivudine (3TC) | 12 adults (ages 52-83) with mild cognitive impairment. 6-month duration. | Reductions in CSF Flt1 and GFAP; increase in plasma Aβ42/40 ratio. No change in neurofilament light or p-tau181. Safe and well-tolerated. |
| Epigenetic Aging Trial (2024) 546259 | Emtricitabine / Tenofovir Alafenamide (FTC/TAF) | 36 healthy adults (ages 18-50) without HIV. 12-week duration. | PhenoAge decreased by 6.33 years. DunedinPACE slowed (-0.061). DNAm-inferred IL-6 reduced (-0.058). Increased pool of naive CD4+ T cells. |
| Epigenetic Aging Trial (2024) 6259 | Emtricitabine / Tenofovir Disoproxil Fumarate (FTC/TDF) | 43 healthy adults (ages 18-50) without HIV. 12-week duration. | No significant changes across epigenetic clocks or inflammatory proxies. Distinct cell deconvolution pattern observed. |
Therapeutic Limitations and Toxicity Profiles
While the gerotherapeutic potential of NRTIs is substantial, chronic administration is historically associated with severe dose-limiting toxicities 5360. Older-generation NRTIs, such as stavudine (d4T), zalcitabine (ddC), and didanosine (ddI), are known to cause severe antiretroviral toxic neuropathy (ATN), myopathy, lipodystrophy, and fatal lactic acidosis 5361. These adverse effects are primarily driven by the off-target inhibition of human DNA polymerase gamma, the enzyme responsible for replicating mitochondrial DNA, leading to severe mitochondrial depletion, altered cellular autophagy, and oxidative stress 536061.
However, newer NRTIs like lamivudine, emtricitabine, and tenofovir alafenamide have a vastly improved affinity profile, exerting potent inhibition of reverse transcriptase with minimal interference on mammalian DNA polymerases 1360. Consequently, the risk of mitochondrial toxicity is dramatically lower with contemporary agents, positioning them as highly viable candidates for long-term clinical trials targeting age-associated inflammaging, provided that diligent safety monitoring is maintained 1360. Further pharmacological refinement is necessary to develop bespoke RT inhibitors optimized specifically for LINE-1 suppression rather than viral eradication.
Conclusion
The characterization of transposable elements as potent drivers of mammalian aging represents a profound paradigm shift in geroscience. No longer viewed simply as inert evolutionary relics, retrotransposons act as endogenous pathogens that exploit the epigenetic vulnerabilities of aging somatic tissues. The age-associated deterioration of DNA methylation and the failure of SIRT1- and SIRT6-mediated heterochromatin silencing unleash these genomic parasites, resulting in the rapid transcription and reverse transcription of elements like LINE-1. The subsequent accumulation of cytosolic nucleic acids activates the cGAS-STING and RIG-I pathways, bridging the gap between localized epigenetic drift and systemic, tissue-destroying sterile inflammation. By conceptualizing aging partly as an autoimmune reaction against the deteriorating genome, novel therapeutic avenues open up. The repurposing of nucleoside reverse transcriptase inhibitors offers a promising, immediately translatable strategy to quiet the genome, suppress inflammaging, and decelerate the broad spectrum of age-related degenerative diseases.