Science of the exposome and disease development
Evolution of the Exposome Paradigm
Limitations of the Genomic Model
For decades, epidemiological and biomedical research operated predominantly under a genetic paradigm, systematically searching the human genome for the primary drivers of disease. However, following the completion of the Human Genome Project, the scientific consensus shifted as researchers determined that genetic variations account for only a modest proportion - estimated between 10% and 20% - of the risk for chronic, non-communicable diseases 1. The remaining 80% to 90% of disease risk is driven by non-genetic factors, specifically environmental exposures, lifestyle behaviors, and their complex interactions 123.
To address this massive gap in disease etiology, the concept of the "exposome" was introduced in 2005 by cancer epidemiologist Christopher Wild. Wild proposed the exposome as the environmental equivalent to the genome, defining it as the totality of life-course environmental exposures from the prenatal period onward 34567. By 2014, researchers expanded this definition to explicitly include the associated biological responses to these environmental influences, recognizing that internal physiological reactions are inseparable from the exposures themselves 578. More recently, the Banbury Exposomic Consortium established a consensus definition positing the exposome as an integrated compilation of all physical, chemical, biological, and psychosocial influences that impact human biology across the lifespan 9.
Refinement of Measurement Approaches
Historically, traditional environmental epidemiology operated on a "one-chemical-at-a-time" framework, attempting to isolate the direct health impacts of specific environmental toxicants 41011. Exposomics fundamentally rejects this reductionist model. Instead, it demands a systematic, agnostic, and discovery-driven evaluation of multiple concurrent exposures and their combined effects 412.
The characterization of these exposures relies on two complementary methodologies: bottom-up and top-down measurement. The bottom-up approach attempts to measure all external environmental factors an individual is subjected to by monitoring air, water, and diet 3. However, this method requires vast infrastructure and often misses critical internal variables shaped by individual physiology. Consequently, researchers increasingly favor a top-down approach, which measures internal biological markers of exposure - such as metabolites, protein adducts, and circulating inflammatory markers - that reflect the cumulative bodily changes caused by exposures after they have occurred 3.
The Three Intersecting Domains
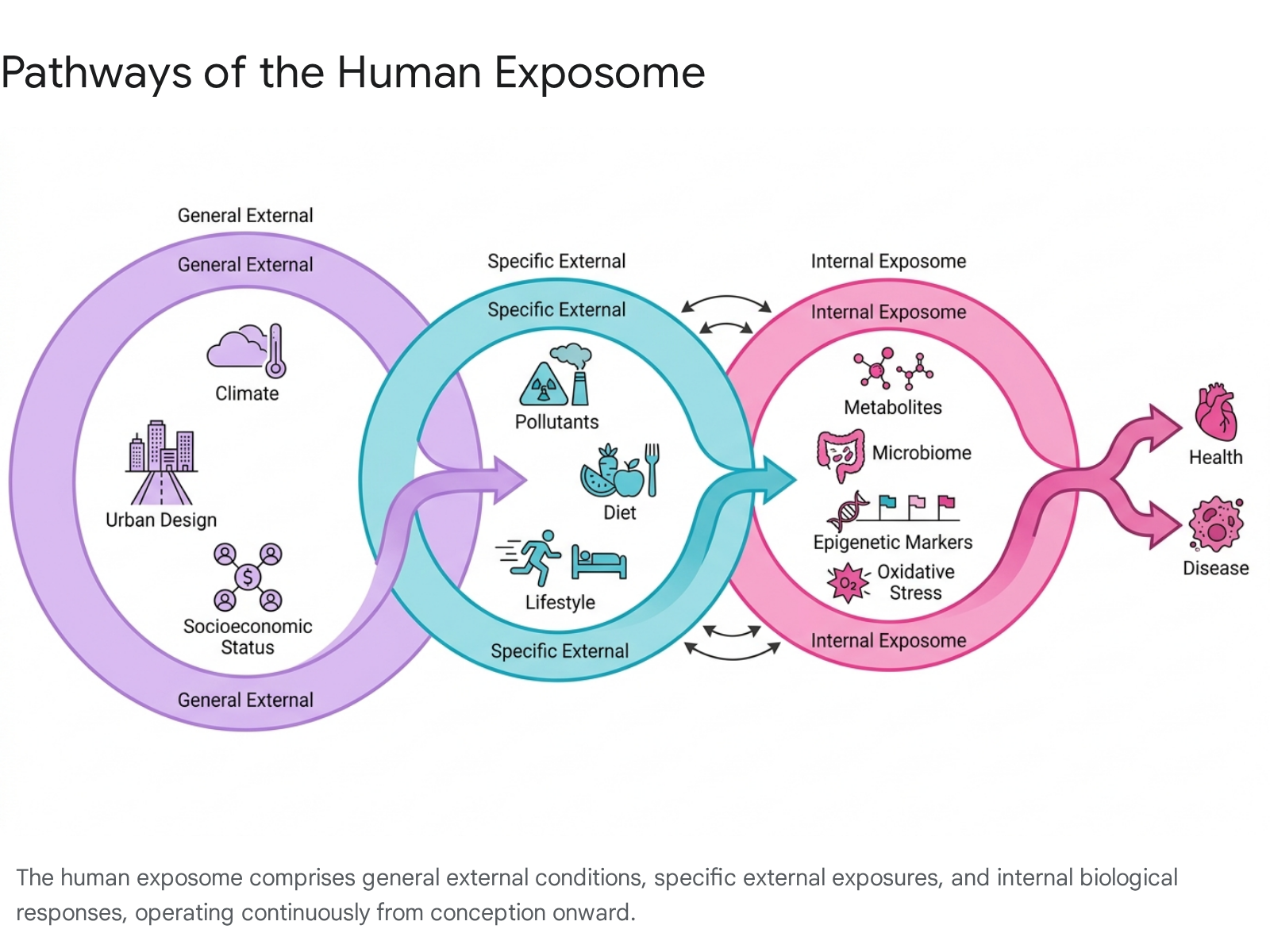
The exposome is highly dynamic, characterized by exposures that fluctuate on momentary, daily, and lifelong scales. To operationalize this vast complexity, the exposome is typically categorized into three broad, overlapping domains that map the pathway from macro-level environments to cellular biology 3456912131415.
| Exposome Domain | Description and Scope | Key Examples and Stressors |
|---|---|---|
| General External | Broad social, economic, sociopolitical, and ecological conditions that shape individual and population-level exposures. Often measured at the geographic or demographic level. | Climate, urban-rural design, socioeconomic status, psychological stress, social capital, built environment, neighborhood deprivation. 3451314 |
| Specific External | Measurable agents of harm or direct lifestyle factors that an individual physically interacts with, ingests, or inhales. | Air and water pollutants, ultraviolet radiation, infectious agents, diet, physical activity, tobacco use, occupational hazards, synthetic consumer chemicals. 345914 |
| Internal | Endogenous biological responses and metabolic processes occurring within physiological borders. Reflects how the body interacts with and alters external exposures. | Hormones, intermediate metabolism, oxidative stress, systemic inflammation, gut microbiota composition, lipidomics, epigenetic modifications. 34591014 |
The boundaries between these domains are highly permeable. General external variables, such as neighborhood deprivation, dictate the likelihood of exposure to specific external hazards, including poor ambient air quality or systemic nutritional deficits 1416. In turn, these specific external exposures drive systemic changes in the internal exposome, permanently altering metabolic homeostasis, gene expression, and immune function 41417.
Analytical Metrology and Biomarker Discovery
High-Resolution Mass Spectrometry
A critical enabler of the modern exposomic paradigm is the advancement of high-resolution mass spectrometry (HRMS) 1318192021. While earlier toxicological studies relied on low-resolution mass spectrometry to perform quantitation-centric targeted analyses of familiar compounds, untargeted HRMS is explicitly hypothesis-generating 1921. By coupling HRMS with gas chromatography (GC-HRMS), liquid chromatography (LC-HRMS), or ion mobility spectrometry (IMS), researchers can capture a broad, unbiased profile of thousands of synthetic and natural compounds, their metabolites, and transformation products in a single biological sample 13181921.
The scale of this analytical challenge separates exposomics from upstream omics fields. Genomics and proteomics sequence sequences derived from a limited set of nucleotides (four) or amino acids (twenty), whereas exposomics and metabolomics must contend with up to millions of distinct small molecules occupying a vast and unstructured chemical space 21.
Challenges in Chemical Dark Matter
Despite the heightened sensitivity of HRMS, detecting a molecular feature via mass spectrometry does not automatically identify the underlying chemical structure. A substantial portion of the chemical exposome currently exists as "dark matter" - compounds that are clearly detected in environmental or human biofluids but remain structurally unidentified and uncharacterized regarding their health risks 1822.
Overcoming this analytical bottleneck requires robust, tiered workflows. Analysts are transitioning from purely non-targeted screening to suspect screening utilizing heuristic rules, and ultimately to confirmed quantitation using commercially available chemical standards 21. Expanding the coverage of chemical space necessitates highly harmonized data pipelines that permit retrospective validation and integration with multi-omics datasets 1921.
Wearable Technologies and Environmental Sensing
Limitations of Static Monitoring
While mass spectrometry elucidates the internal exposome, precision environmental health increasingly relies on intelligent wearable sensors to map the specific external exposome 222324. Traditional environmental epidemiology relies heavily on static outdoor weather stations or geographic information systems (GIS) to estimate regional air quality or climate metrics 252627. Researchers then assign these ecological estimates to individuals based on residential zip codes. This spatial modeling routinely fails to account for micro-environments, localized indoor air quality, and the high mobility of modern human populations 2527.
Advancements in Mobile and Wearable Sensors
Flexible, multi-functional wearable sensors bridge this measurement gap by continuously acquiring, processing, and storing signals generated by physical and chemical changes in the wearer's immediate vicinity 2228. Recent technological advancements have expanded the scope of wearable capabilities far beyond rudimentary fitness trackers.
Modern sensory architectures can capture highly granular exposure metrics: * Airborne Pollutants and Gases: Wearables actively sample particulate matter counts (PM 1.0, PM 2.5, PM 4, and PM 10), volatile organic compounds (VOCs), sulfur dioxide, and nitrogen oxides (NO and NO2) 252729. * Physiological Responses: Biological sensors capture real-time cardiovascular reactivity via electrocardiograms (ECG), photoplethysmography (PPG), and ballistocardiography - a noninvasive technique that measures the mechanical recoil of the body during cardiac systole 23. Advancements also include wearable transcutaneous oxygen sensors and smart apparel capable of tracking localized physiological stress 31. * Ambient Physical Conditions: Embedded sensors document relative humidity, acute temperature fluctuations, noise exposure, ultraviolet radiation, and multi-spectral light intensity 232527.
Real-Time Health Impact Tracking
Large-scale clinical deployments are demonstrating the efficacy of these personalized monitoring approaches. The Artificial Intelligence Ready and Equitable Atlas for Diabetes Insights (AI-READI) study engineered custom sensor units placed directly in the homes of 4,000 participants. Over consecutive ten-day periods, these highly portable devices measure localized particulate matter and VOCs alongside the participants' continuous glucose monitoring and extensive retinal imaging. This generates a dense, personalized dataset mapping exactly how precise micro-environments impact ocular health in patients with type 2 diabetes 25.
Similarly, a pilot study by researchers at The City University of New York integrated smartwatches, smartphone GPS tracking, and Ecological Momentary Assessments (EMAs) to build individualized exposure profiles that move with participants throughout their day. The analysis detected immediate physiological consequences: acute exposure to heat and nitrogen dioxide directly correlated with measurable reductions in heart rate variability, a marker of physical stress recovery, while sulfur dioxide exposure precipitated acute psychological manifestations of nervousness and hopelessness 27. Furthermore, researchers at Stanford University have developed an active sampling prototype capable of collecting both particulate matter and solvent compounds, integrating subsequent data handling with proprietary genomic analysis to isolate precise disease etiologies 29.
Artificial Intelligence in Exposomics
Parsing High-Dimensional Mass Spectrometry Data
The capability of modern analytical chemistry reveals a critical paradox: the current capacity to generate high-resolution exposomic data far exceeds human capacity to interpret it for meaningful health risk assessment 1830. To bridge this gap, artificial intelligence (AI) and machine learning (ML) have become transformative forces 131820.
AI algorithms are deployed across the entire mass spectrometry preprocessing pipeline. Because real-world biological samples are extraordinarily complex, deep learning models such as peakonly, CPVA, and MetaClean are utilized for automated peak extraction, effectively reducing noise that fails to meet strict data quality requirements 18. Additionally, ML algorithms perform batch effect corrections on large-scale samples, ensuring that longitudinal exposure data remains statistically robust and reproducible 18. Once peaks are accurately identified, deep learning models for compound annotation - including computational tools such as CFM-ID, BUDDY, and SIRIUS - infer molecular structures directly from spectral data, rapidly translating chemical dark matter into recognizable xenobiotics 18.
Network Modeling and Causal Inference
Unlike controlled laboratory settings, human populations are exposed to highly correlated, complex mixtures of chemicals simultaneously. Traditional statistical models struggle with the high collinearity and sheer dimensionality inherent in these mixtures. AI platforms address this by constructing advanced models to predict additive, synergistic, or antagonistic toxicological effects 1318. Graph Neural Networks (GNNs) and regularized logistic regression capture non-linear, multi-variable relationships among environmental features, enabling researchers to calculate the combined systemic impact of multiple chemicals on disease outcomes 1318.
Furthermore, causal inference in exposomics is threatened by unmeasured confounders. Because genetics, diet, and socio-geography interact dynamically, causal structure learning and high-dimensional propensity score matching are required to separate random biological variation from genuine exposure-induced perturbations 18. Large language models (LLMs) are also accelerating this process by extracting hidden "chemical-activity" relationships from vast, unstructured toxicology databases, streamlining the generation of high-priority candidate lists for experimental validation 18.
Exposome Intelligence and Digital Twins
The convergence of these AI applications has birthed the concept of "Exposome Intelligence" (EI). In this paradigm, industrial-scale machine learning integrates untargeted mass spectrometry, wearable sensor outputs, satellite geospatial feeds, and electronic health records in near-real time 231. Deep learning harmonizes these disparate data streams, extracting multi-level patterns to shift the analytical endpoint from simple chemical identification to functional prioritization 218.
The ultimate horizon of Exposome Intelligence is the creation of human digital twins - computational replicas that integrate an individual's genomic baseline, lifetime exposome history, and clinical trajectories. These adaptive AI models aim to forecast individual disease risk precisely, simulate physiological responses to environmental interventions, and guide highly personalized public health strategies 31.
Exposome-Wide Association Studies (EWAS)
Methodological Distinctions from Genomics
For over two decades, Genome-Wide Association Studies (GWAS) served as the preeminent, hypothesis-free methodology for determining the molecular underpinnings of complex traits. However, GWAS inherently limits its scope to genetic variants - identifying either causal genes or variants in linkage disequilibrium 32333435. Because genetic factors fail to explain the majority of chronic disease incidence, researchers developed the Exposome-Wide Association Study (or Environment-Wide Association Study, EWAS) 836. Analogous to GWAS, EWAS systematically screens hundreds or thousands of environmental exposures and epigenetic markers (such as DNA methylation) for associations with health phenotypes 83236.
While methodologically inspired by genomics, EWAS faces unique structural and statistical challenges due to the fundamentally different nature of environmental data.
| Analytical Characteristic | Genome-Wide Association Studies (GWAS) | Exposome-Wide Association Studies (EWAS) |
|---|---|---|
| Stability of Measurement | Static; a single genetic sequence remains representative throughout the entire lifespan. | Highly dynamic; environmental exposures and epigenetic methylation fluctuate hourly, daily, and across developmental stages. 436 |
| Correlation Structure | Sparse linkage disequilibrium; genetic variables are largely independent across loci. | Dense and highly correlated; behavioral, dietary, and pollutant exposures cluster heavily together within populations. 1536 |
| Confounding Variables | Minimal; primarily limited to population stratification or race/ethnicity. | Severe; unmeasured social factors, lifestyle confounders, and reverse causation are ubiquitous in environmental data. 323436 |
| Causality Inference | High; identified variants are generally causally upstream of the disease phenotype. | Complex; captures causal environmental factors, but also secondary biological responses and confounding pathways simultaneously. 323334 |
Evaluating Biological Overlap and Confounding
Empirical comparisons demonstrate that GWAS and EWAS capture distinct aspects of disease biology. In a large-scale systematic evaluation of 15 complex traits - comparing GWAS results (N > 50,000) against EWAS results (N > 4,500) - researchers found that the two study types generally capture distinct gene sets 32333435. This implies that genes identified by EWAS are often not causally upstream of the trait, but rather reflect downstream confounding, environmental adaptation, or reverse causality (where the disease state alters behavior or methylation) 3234.
However, in certain specific conditions, the biological insights overlap. The aforementioned analysis revealed substantial overlap between GWAS and EWAS findings for diastolic blood pressure (gene overlap P = 5.2 * 10 - 6; gene ontology term overlap P = 0.001) 32333435. This indicates that for particular cardiovascular traits, innate genetic susceptibility and external environmental stress converge on identical mechanistic pathways.
The Cumulative Impact of Environmental Mixtures
Because EWAS evaluates multiple exposures concurrently, it is uniquely suited to model the cumulative reality of human environmental contact. A massive EWAS utilizing 20 years of data from the U.S. National Health and Nutrition Examination Survey (NHANES) tested over 115,000 associations between 619 environmental exposures and 305 clinically relevant health outcomes 37. The analysis demonstrated that while individual exposures exerted only moderate impacts on health outcomes, integrating up to 20 exposures simultaneously yielded a predictive power comparable to robust polygenic risk scores. For instance, a complex mixture of 20 specific exposures - including trans fats, vitamin E levels, and polychlorinated biphenyls (PCBs) - explained 43% of the variation in population triglyceride levels, a major risk factor for heart disease 37.
To optimize these high-dimensional EWAS analyses, researchers rely on advanced dimensionality reduction techniques. Simulation studies comparing statistical approaches for repeated exposome assessments indicate that methods such as sparse partial least-squares and deletion-substitution-addition algorithms generally outperform standard Elastic-Net or distributed lag non-linear models (DLNM) in accurately linking longitudinal exposome data to health outcomes while managing complex correlation structures 38.
Toxicology and Next-Generation Risk Assessment
Shifting from Isolated Hazards to Mixtures
Traditional regulatory toxicology operates predominantly on the "one product, one regulation" principle, relying on single-chemical animal tests or endpoint-centric analyses 3940. This historical framework routinely overlooks the reality of human exposure, in which individuals encounter concurrent physical, chemical, biological, and psychosocial stressors that interact in non-additive ways 3940.
Exposomics serves as the scientific bridge moving toxicology from isolated hazard identification toward a holistic systems biology approach 39. By detailing the internal dose of chemical mixtures and mapping their interactions with dietary and social stressors, exposome-informed toxicology reveals hidden synergistic risks. For example, toxicological models combining high-carbohydrate or high-lipid diets with ubiquitous chemical contaminants demonstrate metabolic disruptions that are not clearly observed when testing the chemicals in isolation 39.
Adverse Outcome Pathways and Mechanism Discovery
To systematically map how environmental mixtures cause disease, exposomics utilizes the Adverse Outcome Pathway (AOP) framework. An AOP is a conceptual construct that details the sequential chain of causally linked biological events starting from an initial molecular initiating event (e.g., binding of a toxicant to a cellular receptor) to an adverse health outcome at the organism level 73941.
Through the integration of analytical chemistry, computational toxicology, and cellular models, researchers elucidate these specific mechanisms. For example, recent studies utilizing high-coverage quantitative metabolomics revealed the mechanism by which magnetite (Fe3O4) nanoparticles induce cardiovascular toxicity: the particles mediate endothelial cell ferroautophagy, leading to localized iron overload and Fenton reactions, which ultimately precipitate myocardial fibrosis 42. Identifying these precise biochemical steps allows for highly targeted public health interventions. Conversely, siloed approaches often fail to map this full chain. Research on per- and polyfluoroalkyl substances (PFAS) has historically focused on isolated aspects of internal exposure, frequently resulting in statistical associations without a clear mechanistic understanding of the specific pathways driving disease risk 42.
Enhancing Regulatory Risk Assessment
The intersection of exposomics and environmental safety has given rise to Next-Generation Risk Assessment (NGRA). NGRA frameworks integrate real-world human biomonitoring data, exposure modeling, and high-throughput in vitro toxicity testing to prioritize chemical compounds for regulatory review 43.
Rather than relying on assumed external concentrations, NGRA utilizes Physiologically Based Pharmacokinetic (PBPK) models to quantify the biologically effective dose of a pollutant within the human body 2643. Researchers can predict internal blood concentrations resulting from an individual's exposome and directly compare them to the half-maximal activity concentration (AC50) generated by in vitro toxicity benchmarks 43. This exposure-driven approach guarantees that regulatory toxicology prioritizes chemicals that are actually present in the population at biologically active concentrations, rather than focusing purely on theoretical hazards 4344.
The Social Exposome and Biological Embedding
Neighborhood Deprivation and Structural Stress
While the exposome framework initially emphasized chemical and physical agents, comprehensive epidemiology quickly revealed a critical blind spot: the social environment. The "social exposome" integrates the structural, psychosocial, and sociodemographic environments into the analysis of biological health 3454647. Variables such as socioeconomic status (SES), educational attainment, neighborhood deprivation, and exposure to systemic violence are not merely demographic confounders; they are primary drivers of the general external exposome that systematically dictate physical health outcomes 4849.
Populations residing in socially disadvantaged neighborhoods face disproportionate cumulative physical exposures. A lack of social capital directly correlates with geographic proximity to industrial pollution, residence in food deserts, lack of access to green space, and chronic psychological stress stemming from financial and housing insecurity 141650. Exposome researchers quantify these macro-level influences using ecological metrics such as the Area Deprivation Index (ADI) or the Townsend Deprivation Index, which evaluate geographic census tracts based on practical indicators including unemployment rates, household overcrowding, and material wealth 4647505152.
The Microbiome as an Internal Mediator
The mechanism by which external social disadvantage and psychosocial stress become permanently encoded into human physiology is termed "biological embedding" or embodiment 4548. Recent evidence robustly implicates the human microbiome - specifically the gastrointestinal microbiota - as a primary biological interface between the social exposome and host metabolism 161752.
The gut microbiome is highly responsive to the "sociobiome" - the combination of psychosocial stress, systemic dietary quality, pollution exposure, and housing conditions dictated by SES 16. Chronic stress alters neuroendocrine signaling within the gut-brain axis, shifting the microbiome toward dysbiosis 16. Large-scale shotgun metagenomic studies of 1,390 females in the UK Twins cohort demonstrated that higher Townsend deprivation scores correlate strictly with reduced alpha microbial diversity and a severe depletion of commensal bacteria that produce short-chain fatty acids (SCFAs), which are vital for regulating systemic inflammation, energy metabolism, and immune function 16505152.
Crucially, specific microbial species have been identified as physiological mediators between social deprivation and chronic illness. Bacterial strains such as Lawsonibacter sp_NSJ_51 and Intestinimonas massiliensis are significantly depleted in individuals living in materially deprived neighborhoods 5152. Both species are prominent producers of butyrate, a compound that supports blood sugar control and mood stability. Because their absence triggers metabolic disruption, the depletion of these specific taxa statistically mediates the observed link between neighborhood deprivation and higher population rates of both type 2 diabetes and clinical anxiety 5152. Similarly, analysis of multi-ethnic cohorts in the U.S. indicates that low individual and neighborhood SES is associated with elevated abundances of Prevotella copri and decreased Dysosmobacter welbionis, confirming that social class fundamentally dictates microbial composition 53.
Social Networks and the Communicable Microbiome
Beyond structural and geographic deprivation, interpersonal social interactions directly shape the internal exposome. A comprehensive study of 1,787 adults across 18 isolated villages in Honduras mapped intricate social networks against 339,137 different bacterial strains 54. The researchers discovered that microbiome sharing extends far beyond spouses or individuals living in the same household; friends, and even second-degree social connections, exhibited substantial microbiome similarity that exceeded random chance 54.
This phenomenon, termed the "social microbiome," operates independently of shared diet, local water sources, or medications 5455. Notably, the degree of microbiome sharing proved to be the strongest statistical predictor of social relationships in the villages, superseding characteristics like wealth, religion, or education 54. This suggests that non-communicable diseases heavily influenced by the microbiome - such as metabolic syndrome or inflammatory bowel disease - may possess communicable, socially transmitted elements, reinforcing the concept that human physiology is inextricably linked to the social exposome 5455.
Global Cohorts and Population Equity
Geographic Disparities in Exposomic Data
Despite the comprehensive intent of the exposome framework, its current application suffers from severe geographic and demographic bias. The vast majority of well-funded, large-scale exposomic consortia - such as the Human Early-Life Exposome (HELIX) project, ATHLETE, EXPANSE, and EXPOsOMICS - are predominantly based in Europe and the United States 361556. Consequently, the predictive models, AI algorithms, and multi-omic reference ranges generated by these networks inherently reflect the environmental baselines and genetic profiles of high-income Western populations 5758.
There is a severe lack of longitudinal exposome data from the Global South - encompassing Africa, Latin America, South Asia, and parts of the Middle East 575960. Populations in these regions experience drastically different exposomic profiles, contending with unique environmental threats such as high burdens of infectious disease, extreme climate events (droughts, heatwaves), unregulated industrial emissions, distinct agricultural pesticide use, displacement, and severe systemic violence 57585960. Furthermore, populations in the Global South exhibit greater genetic admixture than typical U.S. or European cohorts 58. When predictive AI models trained exclusively on Western datasets are applied to these underrepresented populations, they suffer from poor generalizability and display higher misclassification rates regarding disease risk 58.
Expanding Research in the Global South
Ensuring global representation in exposomic research is both a scientific necessity and an ethical priority. Tracking specific regional exposures is crucial because populations in the Global South frequently exhibit distinct disease trajectories, including earlier onset of cardiometabolic conditions and atypical presentations of Alzheimer's disease and related dementias (AD/ADRD) 5861.
To rectify this imbalance, several major international initiatives are currently underway: * Latin America: The Mexican Health and Aging Study (MHAS) is expanding its research infrastructure by linking historical climate measures and lifetime occupational hazard data to study how environmental inequalities drive AD/ADRD in middle-income countries 62. Additionally, organizations are convening regional summits, such as the Latin American Exposome Symposium in Santiago, Chile, to prioritize research integrating environmental exposures into aging models specific to hemispheric stressors 63. * Africa: The Network for Exposomics in the U.S. (NEXUS) and the European Human Exposome Network are facilitating global coordination with African scientists 6465. Specific research grants, such as the PEARLS trial in Sub-Saharan Africa (Ghana, Kenya, South Africa), are validating urinary biomarkers like Activin A and Inhibin A to predict early-pregnancy preeclampsia under unique environmental pressures 65. Furthermore, high-level structural talks are underway in South Africa, coordinating with the Department of Science, Technology, and Innovation (DSTI) to establish a comprehensive Pan-African Exposome Network 1.
Macro-Level Determinants of Accelerated Aging
The necessity of diverse, global data is highlighted by recent studies quantifying the impact of macro-level sociopolitical exposomes. A massive cross-sectional and longitudinal dataset analyzing 161,981 healthy participants across 40 nations evaluated Biobehavioral Age Gaps (BBAGs) - the difference between an individual's estimated biological age and their chronological age 61.
The analysis demonstrated that delayed and accelerated aging are heavily influenced by regional and exposomal factors. While participants in Europe led in healthy, delayed aging, populations in lower-income countries (e.g., Egypt, South Africa) exhibited the greatest age acceleration, with Latin America and Asia falling in between 61. Accelerated aging globally was inextricably linked to physical exposures (poor air quality), social factors (socioeconomic and gender inequality, forced migration), and critical sociopolitical determinants, including restrictions on party freedom, suffrage, and democratic representation 61. Integrating diverse global populations into the exposome paradigm will not only improve precision medicine for local communities but will also expand the universal understanding of how structural inequalities and extreme environmental stressors precipitate biological aging and neurological decline.