Organ-specific biological age clocks and aging discordance
Foundational Concepts in Biological Aging Measurement
Aging is characterized by a gradual, progressive loss of physiological integrity that drives functional decline and increases susceptibility to chronic diseases and mortality 12. Traditionally, aging has been measured chronologically, utilizing the passage of time as a universal metric. However, chronological age fails to capture the immense heterogeneity in physiological deterioration observed across different individuals, and it ignores the reality that human physiology does not decay at a uniform rate 34. To address this fundamental limitation, researchers have developed algorithms to quantify biological age, a metric that provides a more accurate reflection of an individual's molecular, cellular, and systemic state relative to their same-aged peers 56.
While early biological aging clocks provided a single, organism-wide estimate of age, recent high-throughput omics profiling has conclusively demonstrated that aging does not occur uniformly throughout the body 1. Different organ systems within the same individual can exhibit highly divergent aging trajectories, driven by a complex interplay of localized environmental exposures, genetic predispositions, and cellular stress responses 16. This phenomenon has driven the development of organ-specific biological age clocks, which utilize plasma proteomics, DNA methylation, transcriptomics, metabolomics, and advanced neuroimaging to isolate the distinct aging rates of specific tissues 26.
The delta between an organ's predicted biological age and an individual's chronological age is defined as the organ age gap, or clock discordance 17. A positive age gap indicates accelerated biological aging, whereas a negative gap suggests decelerated aging or physiological resilience 18. Analyzing this discordance provides critical insights into multiorgan aging networks, revealing how localized physiological decline in one organ system, such as the brain or cardiovascular system, can cascade to accelerate the aging of interconnected systems. This localized resolution predicts specific disease onset and overall mortality with far greater precision than conventional systemic clocks 91110.
Generational Evolution of Biological Aging Models
The mathematical models used to construct biological aging clocks have evolved through three distinct algorithmic generations, each defined by its training outcomes, underlying data structures, and resulting clinical utility. While early clocks captured generalized systemic decline, the latest iterations provide granular, tissue-level resolution and track the longitudinal pace of biological deterioration 131112.
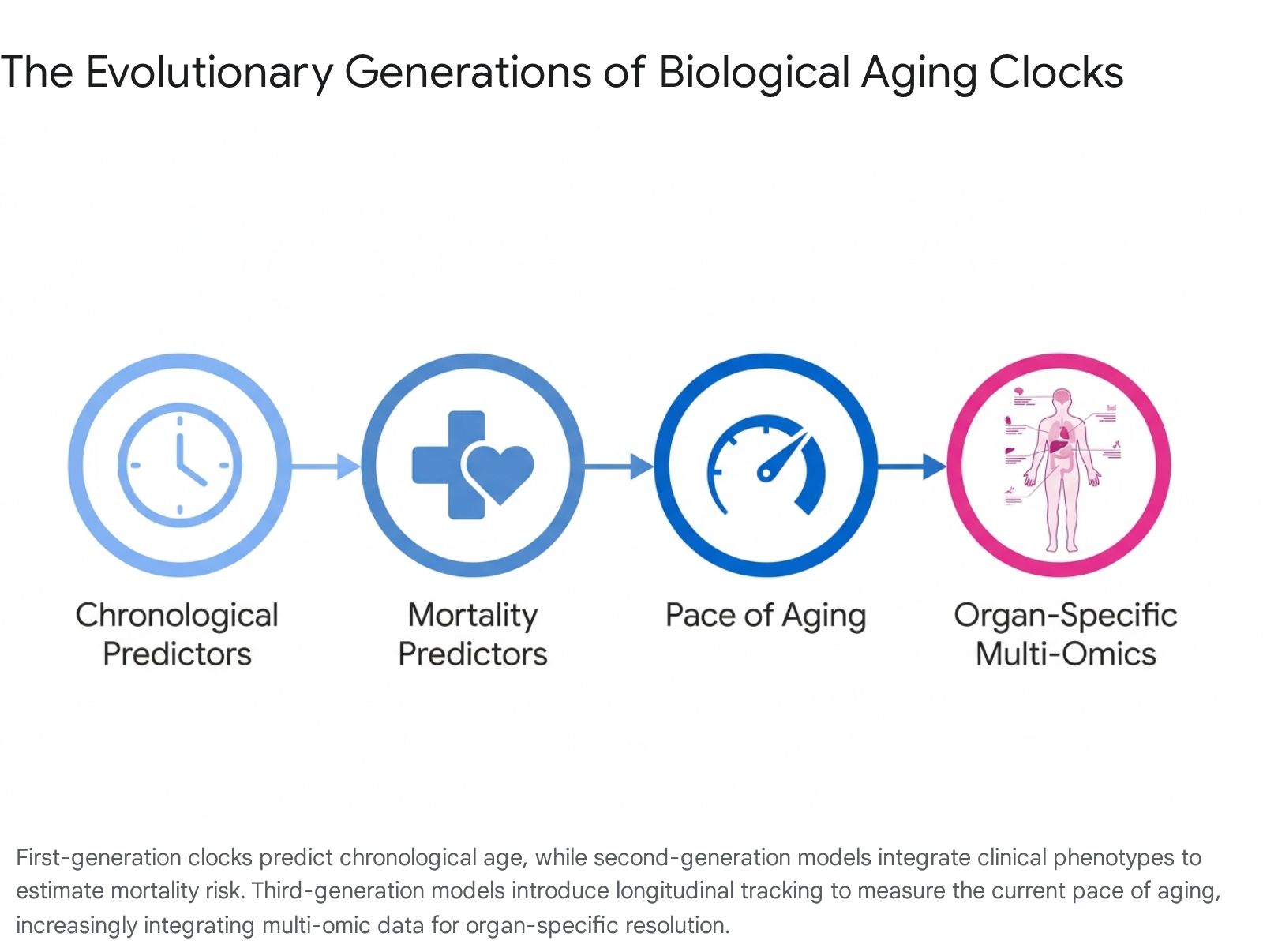

First-Generation and Second-Generation Systemic Clocks
First-generation epigenetic and proteomic clocks were trained exclusively to predict chronological age 21312. Examples include the original Horvath multi-tissue DNA methylation clock and the Hannum blood clock 2. The primary limitation of first-generation clocks is that while they correlate strongly with the linear passage of time, they often show limited sensitivity in predicting the incidence of age-related diseases or mortality in relatively healthy populations 211. Because they are designed to mirror chronological age, they inherently suppress the biological variance that defines accelerated or decelerated aging 12.
Second-generation clocks, such as PhenoAge and GrimAge, were developed to bypass this mathematical limitation by incorporating clinical biomarkers and training the algorithms directly on mortality risk and time-to-death data 131317. PhenoAge links clinical biomarkers to 513 cytosine-phosphate-guanine (CpG) sites, while GrimAge integrates plasma proteins and smoking history to identify 1,030 CpGs that predict lifespan 1714. These models yield a biological age estimate that reflects an individual's morbidity risk relative to a reference population 11. Although highly predictive of lifespan, second-generation models typically rely on blood-based biomarkers or generalized systemic phenotypes, limiting their ability to resolve isolated, organ-specific vulnerabilities 2.
Third-Generation Longitudinal Algorithms
Third-generation models shift the analytical paradigm from calculating a static biological age snapshot to measuring the ongoing pace of aging 1315. Unlike earlier models developed using cross-sectional data comparing different age cohorts, third-generation clocks such as DunedinPACE were derived from longitudinal data 1520. The DunedinPACE algorithm was calibrated using the Dunedin Longitudinal Study, which tracked a birth cohort of roughly 1,000 individuals over several decades, evaluating 19 distinct physiological biomarkers across four testing periods spaced five years apart 1521.
By tracking intra-individual changes over time, these models function as epigenetic speedometers. They quantify the rate at which an individual is accumulating physiological wear and tear per calendar year, outputting a ratio where a score of 1.0 indicates average aging, while 1.20 indicates aging 20 percent faster than the chronological norm 21. DunedinPACE has proven to be highly sensitive to interventions, showing measurable deceleration in biological aging in response to caloric restriction and lifestyle modifications 615.
Alternative approaches to capturing longitudinal dynamics from cross-sectional data involve mathematically isolating the age gap itself. The Gompertz law-based residual (GOLD-R) framework posits that mortality risk increases exponentially with chronological age. The algorithm involves a two-stage modeling process: it generates an initial residual representing the discrepancy between chronological age-based Gompertzian risk and biomarker-predicted risk, and then uses remaining biomarkers to predict the residual itself. This isolates a biologically informed signal of aging acceleration without relying on standard linear regression assumptions, refining biological age estimation for clinical utility 22.
Methodological Frameworks for Organ-Specific Proteomic Clocks
The transition from systemic to organ-specific aging models requires rigorous feature selection to ensure that the circulating biomarkers utilized accurately reflect the localized physiological state of the target organ. Blood plasma acts as an aggregate reservoir, capturing secreted proteins, leakage products from cellular damage, and extracellular matrix remodeling byproducts from all tissues simultaneously 23. Deconvoluting this complex biological mixture relies heavily on transcriptomic mapping 816.
Transcriptomic Mapping via the GTEx Database
The foundational methodology for establishing organ-specific proteomic clocks was formalized by mapping plasma proteins back to their tissues of origin using the Genotype-Tissue Expression (GTEx) human tissue bulk RNA-sequencing database 16. Researchers define an organ-enriched gene as one that exhibits at least a four-fold higher expression level in a single organ compared to all other organs 1718.
This stringent transcriptomic threshold yields mutually exclusive sets of organ-enriched RNA transcripts. Once identified, these genes are mapped to their corresponding plasma proteins measured in high-throughput assays 1018. For example, in the Wang et al. (2025) study utilizing the Olink Explore 3072 panel, 418 proteins, representing roughly 14.3 percent of the measured panel, were annotated as enriched across ten major organs: the brain, heart, lung, immune system, artery, intestine, liver, kidney, muscle, and pancreas 19. The brain and the immune system typically yield the highest number of enriched proteins, with 117 and 109 specifically identified respectively in recent cohorts 19.
Machine Learning and Predictive Modeling
With organ-specific protein sets mathematically isolated, machine learning models are trained against the chronological age of healthy individuals. The most widely adopted algorithms for this task are bagged least absolute shrinkage and selection operator (LASSO) or elastic net regularized regression models 101618. Regularization techniques penalize overly complex models by shrinking the coefficients of less predictive proteins to zero, thereby selecting a sparse, highly robust subset of biomarkers that reliably track physiological time without overfitting the data 16.
An artery-specific aging model evaluates only the plasma concentrations of artery-enriched proteins. Because these models are trained on populations spanning broad age ranges, typically from 20 to 90 years of age, they effectively define the normative trajectory of age-related proteomic shifts for that specific tissue 28. The model calculates a predicted biological age for the organ, and the difference between this prediction and chronological age serves as the actionable organ age gap 7. To prevent data leakage and ensure robustness, models are typically trained using nested cross-validation frameworks 116.
To further refine these models for specific clinical outcomes, researchers employ algorithmic techniques like Feature Importance for Biological Aging (FIBA) 1016. FIBA assesses the contribution of specific proteins within the clock to a designated phenotypic outcome, such as the association between the brain age gap and clinical cognitive decline, allowing for the derivation of cognition-optimized organ aging models 16.
Key Proteomic Biomarkers by Organ System
The feature selection process has identified highly specific proteins whose circulating concentrations shift dramatically with age, providing mechanistic insights into localized physiological deterioration. The pathways implicated highlight how structural damage, chronic inflammation, and metabolic dysfunction coalesce to drive organ-specific aging 2021.
| Organ System | Key Proteomic Biomarkers Identified | Associated Pathological Mechanisms & Observations |
|---|---|---|
| Brain | Neurofilament light chain (NEFL), Glial fibrillary acidic protein (GFAP), Kallikrein-6 (KLK6), Contactin-1/2 (CNTN1, CNTN2), Brevican (BCAN), Synaptotagmin-1 (SYT1) 1920 | Reflects synaptic loss, glial activation, and neuroaxonal damage. Approximately 69% of brain clock markers are low-abundance, non-secreted proteins, indicating structural leakage past the blood-brain barrier 2822. |
| Artery / Vascular | Elastin (ELN), Latent-transforming growth factor beta-binding protein 2 (LTBP2) 19 | Indicates advanced vascular dysfunction, extracellular matrix (ECM) remodeling, and arterial stiffness. These proteins are predominantly expressed in smooth muscle and endothelial cells 1920. |
| Kidney | Cystatin C, Creatinine, Blood Urea Nitrogen (BUN) surrogates, Uromodulin 1923 | Tracks declining glomerular filtration rates, diminished renal clearance, and vulnerability to hypertension. High creatinine directly correlates with accelerated kidney aging 2333. |
| Liver | Albumin variations, AST/ALT ratio-associated proteins 19 | Strongly associated with overall metabolic dysfunction, fatty liver disease progression, and the accumulation of baseline hepatic damage 1934. |
| Immune | 76 highly specific proteins, including Leukocyte associated immunoglobulin like receptor 1 (LAIR1), Interleukin variants, and IgG-3 21933 | Reflects profound immunosenescence and chronic low-grade systemic inflammation (inflammaging). Frequently intersects with degraded T-cell mediated immunity pathways 233. |
| Organismal (Systemic) | Growth differentiation factor 15 (GDF15), Interphotoreceptor matrix proteoglycan 2 (IGDCC4) 1920 | GDF15 acts as a powerful generalized stress signal. High systemic correlation with overall mortality and incident vascular dementia 2023. |
Neurological and Vascular Tissue Biomarkers
The brain clock relies heavily on structural proteins that ordinarily reside within the central nervous system. The presence of proteins like neurofilament light chain (NEFL) and glial fibrillary acidic protein (GFAP) in the plasma indicates neuroaxonal damage, glial activation, and compromised blood-brain barrier integrity 1920. Synaptic loss is tracked through markers like synaptotagmin-1 (SYT1) 19. The arterial clock is similarly structural, relying on elastin (ELN) and LTBP2, which map directly to vascular dysfunction and extracellular matrix remodeling within arterial smooth muscle cells 1920.
Renal, Hepatic, and Immune Signatures
Kidney and liver clocks frequently integrate classical clinical biomarkers that have been translated into high-throughput proteomic equivalents. The kidney clock leverages Cystatin C and creatinine analogs to map declining glomerular filtration, while the liver clock utilizes variations in albumin and AST/ALT ratios to track hepatocyte stress 1923. The immune clock presents the most complex signature, relying on dozens of interacting cytokines and immunoglobulin receptors to quantify immunosenescence. The dysregulation of T-cell mediated immunity and the chronic elevation of inflammatory markers form the core of the immune biological age gap, propagating signals that accelerate aging in downstream systems 233.
Organ Age Discordance and Disease Prediction
The primary clinical utility of organ-specific clocks lies in their ability to analyze discordance. A positive age gap serves as a potent, leading indicator of localized disease vulnerability, often detectable years before clinical symptoms or functional deficits manifest in standard medical evaluations 18.
Pathological Risk Profiles for Single-Organ Accelerated Aging
Biological aging is highly heterogeneous within the general population. Research indicates that nearly 20 percent of seemingly healthy adults exhibit strongly accelerated aging in at least one organ system, which is mathematically characterized as a biological age one standard deviation or more above the mean for their chronological peer group 1628. These localized accelerations track precisely with future, organ-specific morbidities across vast population cohorts.
A positive brain age gap is one of the strongest independent predictors of cognitive decline and dementia 1022. Extensive longitudinal data demonstrates that having an especially aged brain poses a hazard ratio for Alzheimer's disease of 3.1, a risk magnitude functionally comparable to carrying one copy of the APOE4 genetic variant, which is the strongest known genetic risk factor for sporadic Alzheimer's disease 24. Conversely, displaying a super-youthful brain profile provides protective resilience that mirrors carrying the protective APOE2 variant, heavily offsetting the baseline genetic risk inherent in APOE4 carriers 24.
Accelerated cardiac aging correlates strongly with higher left-ventricular mass, altered stroke volume, and an increased incidence of atrial fibrillation 110. A four-year acceleration in biological heart age increases the risk of incident heart failure by 250 percent over a 15-year follow-up period 2825. Similarly, accelerated aging in the kidney, pancreas, and intestine - three interconnected organs heavily involved in metabolic regulation - reliably predicts the onset of Type 2 diabetes and hypertension 133. The kidney age gap alone raises the hazard ratio for developing chronic kidney disease to 1.67 18.
Multi-Organ Accrual and All-Cause Mortality
While a single aged organ dramatically increases specific disease risk, the compounding accrual of multiple aged organs exponentially escalates all-cause mortality risk 2426. Individuals defined as multi-organ extreme agers, comprising roughly 1.7 percent of the population, face severe survival deficits regardless of their chronological age 16.
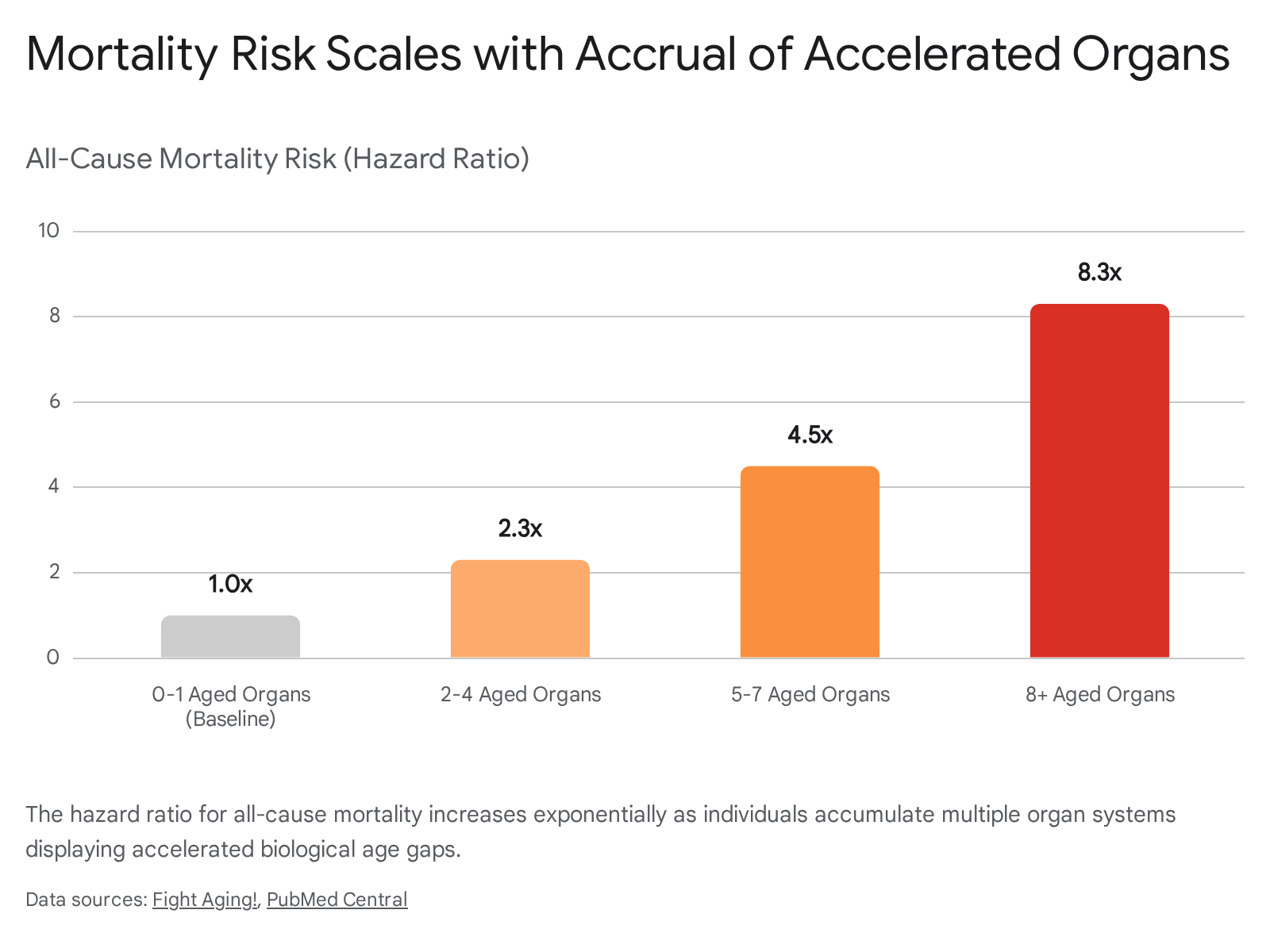
Robust 15-year longitudinal follow-ups tracking tens of thousands of participants show that mortality risk scales directly with the number of organ systems exhibiting accelerated biological age. Individuals displaying accelerated aging in 2 to 4 organs exhibit a 2.3-fold increased mortality risk. Those with 5 to 7 aged organs face a 4.5-fold increase in mortality risk. Most severely, individuals with 8 or more accelerated organs face an 8.2- to 8.3-fold higher risk of death compared to synchronous, normal agers 1024.
Amidst this systemic decline, maintaining youthful brain and immune system clocks provides unique protective effects. A youthful brain yields a hazard ratio of 0.60 for mortality, a youthful immune system yields 0.58, and maintaining both simultaneously reduces mortality risk to a remarkable 0.44 hazard ratio 24.
Multiorgan Aging Networks and Propagation Dynamics
Biological organs do not age in physiological isolation. The extensive analysis of clock discordance reveals a highly integrated multiorgan aging network, where the biological deterioration of one tissue directly accelerates the decline of others over time 93827.
Structural Equation Modeling of Aging Networks
To map these complex inter-organ relationships, researchers have applied advanced statistical frameworks to vast longitudinal and cross-sectional datasets. Utilizing brain imaging-derived phenotypes and physiological traits from the UK Biobank, mathematical models assess how the age gap of one organ relates to another 728. By regressing variables such as chronological age, biological sex, scanning sites, and total intracranial volume, analysts isolate the pure residuals representing the independent biological age gaps 7.
Using Fast-Greedy Equivalence Search (FGES) heuristics for continuous variables, causal Bayesian networks are constructed to determine the highest scoring model of multiorgan influence 11. A directed edge in these multiorgan networks signifies that the baseline age gap of one specific organ significantly influences the subsequent rate of aging or the follow-up age gap of another organ 11.
Inter-Organ Influence and Synchronicity
These network models reveal clear causal hierarchies and propagation dynamics in human physiological aging. Early acceleration in cardiorespiratory systems consistently acts as a primary catalyst for downstream metabolic aging. For instance, an accelerated biological age in the lung reliably precedes and influences the accelerated aging of the kidney, intestine, and muscle systems 1.
Furthermore, the immune and adipose systems serve as the central interconnecting hubs within this multiorgan network 1. Chronic systemic inflammation, mathematically captured by an accelerated immune age clock, propagates molecular damage across both the vascular and metabolic networks. This establishes a degraded biological environment that forces synchronous accelerated aging across the entire organism over a decade of follow-up monitoring 118. Consequently, interventions targeting immune resilience hold outsized potential for decelerating whole-body aging.
Environmental, Genetic, and Lifestyle Determinants
Organ age trajectories are shaped by a highly complex interplay of fixed genetic architecture and modifiable environmental exposures. Disentangling these factors is crucial for understanding why certain individuals exhibit massive organ clock discordance compared to their peers.
Genetic Architecture of Organ Aging
Genome-wide association studies leveraging multiorgan aging data have identified 119 specific genetic loci associated with organ aging, including 27 loci that are shared across multiple interacting organs 33. Researchers have prioritized 554 risk genes involved in organ-relevant biological pathways 33. The brain clock, for instance, is heavily influenced by specific genetic variants, with the GABBR1 and ECM1 genes exerting strong control over neurological aging trajectories 22. Genetic determinants also deeply intersect with clinical outcomes; as noted previously, the brain aging clock stratifies Alzheimer's risk across distinct APOE haplotypes, proving that genetic predispositions can be either exacerbated or mitigated by the actual biological aging rate of the organ 22.
Lifestyle and Toxin Exposure
Organ clocks are highly responsive to modifiable environmental factors, often displaying dramatic localized acceleration in response to toxins 33. Smoking initiation causes severe, localized biological age acceleration in the lung, intestine, kidney, and stomach 33. This acceleration is likely driven by the cumulative impact of inflammatory and oxidative damage inflicted directly on tissues exposed to inhaled particulates or their metabolically processed toxic byproducts 18. Heavy alcohol consumption specifically targets and accelerates pancreatic and liver aging 1.
The menopausal transition serves as one of the most profound physiological modifiers of biological aging. In longitudinal studies tracking middle-aged women, the transition from pre-menopause to post-menopause triggers synchronous, multi-organ age acceleration. This systemic shift dominates the female aging landscape and is characterized by acute spikes in immune, liver, full-body, and arterial biological age gaps, correlating directly with the sudden loss of protective estrogen signaling 1.
Dietary and Pharmacological Interventions
The malleability of organ clocks under targeted interventions confirms their utility as dynamic biomarkers. In the Comprehensive Assessment of Long-term Effects of Reducing Intake of Energy Phase 2 randomized controlled trial, participants assigned to two years of sustained caloric restriction exhibited significant biological age deceleration 41. However, the effects were profoundly heterogeneous across organ systems. Caloric and protein restriction robustly decelerated aging in the metabolic, cardiovascular, and immune systems, but left kidney biological age largely unaffected 41.
Pharmacological interventions also alter clock trajectories. The initiation of medications like RAS inhibitors or statins modifies longitudinal organ age gaps, though researchers caution that these alterations often manifest through specific drug-targeted proteins, such as renin and apolipoprotein B, rather than by reversing generalized systemic organ aging entirely 1. Supplemental interventions, including omega-3 fatty acids combined with vitamin D and exercise, have demonstrated additive protective effects, slowing the progression of epigenetic aging clocks over three-year trial periods 14.
Cross-Cohort Validation in Diverse Populations
For organ-specific clocks to be viable clinical and epidemiological tools, they must demonstrate stability and generalizability across diverse genetic, environmental, and ancestral populations 1923. Historically, many systemic biomarkers established in Western, Caucasian-majority cohorts have shown significant predictive variance when applied to other ethnicities 29. This is due to intrinsic population differences in baseline inflammatory loads, varying ancestral allele frequencies, and distinct lifelong environmental exposures 29.
External Validation in Non-Western Cohorts
Recent high-impact literature has rigorously tested the generalizability of proteomic organ clocks across distinct global cohorts 22. Wang et al. (2025) developed their comprehensive suite of organismal and organ-specific aging clocks utilizing a massive discovery cohort of 43,616 primarily European-ancestry participants from the UK Biobank 2243. To rigorously validate the models, the researchers applied them directly to independent external cohorts without recalibration. These cohorts included 3,977 Asian-ancestry adults from the China Kadoorie Biobank and 800 participants from the US Nurses' Health Study 2243.
The results demonstrated exceptional cross-cohort reliability. The proteomic organ clocks achieved Pearson correlation coefficients of r = 0.98 in the Chinese cohort and r = 0.93 in the US cohort when predicting chronological age 1922.
Implications for Universal Pathogenic Pathways
This exceptionally high cross-ancestral accuracy indicates that the specific proteomic markers capturing organ decline rely on universally conserved biological pathways rather than population-specific artifacts 19. The structural degradation implied by circulating neurofilament light chain leaking from the brain, or elastin degrading from the vascular walls, occurs through fundamental biological mechanisms shared across all human populations 19. Consequently, these validated organ-specific clocks provide a robust, biologically interpretable framework for tracking multiorgan aging and stratifying disease risk globally, bypassing many of the demographic biases that have plagued earlier single-metric aging estimates 1922.
Technological Variabilities in Proteomic Evaluation
Despite the robust biological findings, the reliability of organ-specific proteomic clocks is inherently tied to the analytical platforms used to measure the plasma proteome. The two dominant technologies in this space - SomaLogic's SomaScan and the Olink Proximity Extension Assay (PEA) - utilize fundamentally different biochemical mechanisms 30. These differences result in distinct advantages, biases, and technical noise profiles that heavily influence the construction and reproducibility of aging clocks 30.
Measurement Modalities and Target Specificity
SomaScan technology relies on slow off-rate modified DNA aptamers (SOMAmers) engineered to bind specific target proteins with high affinity. Coupled with microarray or bead-based readout systems, this platform enables broad proteome coverage, capable of quantifying upwards of 11,000 proteins simultaneously across a wide dynamic range 3045. This extreme multiplexing makes SomaScan an exceptional tool for large-scale, exploratory population proteomics 45. However, aptamer-based arrays can be more susceptible to nonspecific binding in highly complex biological matrices like plasma 30.
In contrast, the Olink PEA platform utilizes pairs of oligonucleotide-labeled antibodies. Both antibodies must simultaneously bind the target protein in close proximity to trigger DNA polymerization and subsequent quantitative amplification via real-time PCR or next-generation sequencing 30. This dual-recognition mechanism provides exceptional target specificity and very low background signal, making it highly sensitive for detecting low-abundance signaling proteins and critical immune cytokines 3045.
Assay Precision and Cross-Platform Concordance
Extensive technical benchmarking studies conducted in 2024 and 2025 reveal that these two platforms are not trivially interchangeable 3031. The platforms exhibit significantly divergent precision metrics.
| Performance Metric | SomaScan (7K / 11K) | Olink (Explore HT / 3072) | Clinical Implications for Aging Clocks |
|---|---|---|---|
| Measurement Mechanism | Modified DNA Aptamers (SOMAmers) 30 | Proximity Extension Assay (Dual-Antibody + PCR) 30 | Aptamers offer massive scale; PEA offers extreme specificity for low-abundance targets. |
| Median Coefficient of Variation (CV) | ~5.3% to ~6.8% 4532 | ~26.8% to ~35.7% (Improves when filtering out proteins below detection limits) 4532 | SomaScan presents lower technical noise across its broader panel, ensuring high reproducibility for large cohorts. |
| Data Completeness | ~95.8% to 96.2% 45 | Highly variable; robust for specific panels but requires strict thresholding 4532 | High data completeness is critical for training complex multivariable machine learning algorithms like LASSO. |
| Cross-Platform Concordance (Spearman's ρ) | N/A (Baseline comparison) | ~0.46 to 0.65 correlation against SomaScan 304532 | Aging clocks trained on one platform cannot be directly applied to the other without extensive normalization and bridging. |
| Biological Bias | Enriched for structural, metabolic, and less-studied proteins 32 | Enriched for inflammatory, signaling, and immune pathways 32 | Platform choice heavily dictates which pathogenic pathways an aging clock will ultimately identify as critical. |
When measuring identical plasma samples, cross-platform correlation is surprisingly modest. Analyses comparing overlapping proteins report median Spearman's correlations ranging from 0.46 to 0.65 3032. While highly abundant proteins correlate well across platforms, many lower-abundance markers do not 32.
These technical discrepancies introduce a critical limitation in the geroscience field: an organ aging clock trained on SomaScan data cannot be directly applied to Olink data without extensive bridging, normalization, and potential recalibration of the underlying predictive weights 32. Furthermore, platform-specific biological biases dictate that Olink panels tend to enrich for immune and signaling cascades, while SomaScan captures a broader array of structural and metabolic proteins. This measurement bias directly influences the functional pathways identified by the resulting aging clocks, necessitating careful consideration when interpreting multi-organ networks 32.
Epigenomic and Multi-Omic Architectures
While proteomics provides a real-time, highly dynamic snapshot of physiological state, other omic layers capture entirely distinct biological dimensions of aging. Proteomic clocks reflect immediate physiological stress and systemic responses, but epigenomic and transcriptomic layers reveal deeper, cumulative alterations to cellular programming. Advanced multi-omic organ clocks integrate these layers to provide a holistic assessment of cellular integrity 2.
Organ-Specific DNA Methylation Proxies
Epigenomic clocks, which analyze cytosine-phosphate-guanine (CpG) methylation patterns, have historically dominated the biological aging field 213. However, creating true organ-specific epigenomic clocks from minimally invasive peripheral blood draws is highly complex, as DNA methylation is inherently cell-type and tissue-specific. The methylation patterns of a liver cell cannot be directly read from a circulating leukocyte 2.
To bypass this limitation, recent algorithmic frameworks, such as SYMPHONYAge, have trained robust multi-system proxy models. SYMPHONYAge analyzes over 130 specific epigenetic markers from a blood draw to yield individual age estimates for 11 distinct systems, including the brain, heart, liver, metabolic, and musculoskeletal systems 4849. Validated across multiple longitudinal cohorts, including the Baltimore Longitudinal Study of Aging, these models achieve high accuracy in predicting organ-specific disease occurrences and distinguishing aging subtypes based purely on epigenetic proxies circulating in the blood 50. Similar comprehensive multi-omic models, like OMICmAge, integrate over a million methylation sites alongside clinical, proteomic, and metabolic data to improve mortality prediction accuracy significantly beyond standard first-generation methylation clocks 48.
Metabolomic and Transcriptomic Integration
The multi-organ framework is actively expanding into other high-dimensional modalities. Recent massive-scale studies have utilized plasma non-derivatized metabolites to create metabolome-based biological age gaps (MetBAGs). These metabolic clocks effectively track cardiometabolic health and mortality via small-molecule metabolic byproducts, capturing the terminal end of the physiological cascade 33. Similarly, transcriptomic clocks leverage long non-coding RNAs and microRNAs to track cellular senescence, senescence-associated secretory phenotypes (SASP), and extracellular matrix turnover at the tissue level 2.
Integrating these disparate datasets remains a profound bioinformatics challenge due to varying temporal dynamics. The plasma proteome and metabolome shift rapidly in response to acute stressors, dietary intake, and medications, whereas the epigenome changes slowly, reflecting long-term, cumulative environmental exposures 2. Future multi-omic organ clocks must harmonize preprocessing pipelines and employ advanced network-based modeling to accurately weigh the relative contributions of these interacting layers.
Conclusions and Future Directions
Organ-specific biological age clocks represent a fundamental paradigm shift in geroscience, moving the field beyond the blunt instrument of chronological age and the generalized approximations of systemic biological clocks. By leveraging high-throughput proteomics, epigenomics, and advanced transcriptomic mapping, these tools expose the deep, heterogeneous discordance of human aging, revealing that interconnected organ systems decay along vastly different trajectories.
The clinical utility of these clocks is profound. Analyzing clock discordance allows for the highly specific prediction of localized disease onset - such as the massive Alzheimer's risk associated with an accelerated brain age gap - years before traditional clinical symptoms emerge. Furthermore, mapping the multiorgan aging network has illuminated causal hierarchies, demonstrating how early cardiorespiratory or immune decline can propagate systemic failure, exponentially increasing mortality risk as accelerated organs accrue.
Despite these rapid advancements, several limitations must be addressed before widespread clinical adoption is feasible. Current foundational proteomic models are largely trained to predict chronological age rather than definitive functional decline or morbidity endpoints. Because signatures of healthy physiological aging can diverge from the signatures of age-related disease, the next generation of organ clocks must be explicitly trained on clinical outcomes and functional deficits to maximize translatability. Additionally, standardizing batch effect corrections and navigating the severe cross-platform discrepancies between technologies like SomaScan and Olink remain critical hurdles for epidemiological interoperability. Ultimately, as these algorithms refine their resolution, organ-specific clocks will provide unprecedented opportunities for early disease detection, the validation of targeted senolytic and metabolic interventions, and the precise extension of human healthspan.