Knot theory in DNA, quantum field theory and quantum computing
Abstract
Knot theory, historically a purely abstract subfield of geometric topology, has emerged as a fundamental mathematical framework for understanding complex physical and biological systems. This report provides an exhaustive, cross-disciplinary synthesis of topological principles, tracing their theoretical mathematical foundations to their empirical manifestations in DNA replication, quantum field theory (QFT), and topological quantum computing (TQC). The analysis establishes the critical distinction between ideal mathematical knots and physical knots governed by thermodynamic and volumetric constraints. It subsequently details the computational complexity of evaluating knot invariants, notably the #P-hardness of the Jones polynomial, and explores the categorification of these polynomials via Khovanov homology. In the physical realm, the report examines the role of Chern-Simons theory and the Yang-Baxter equation in linking topology to statistical mechanics and quantum phenomena. Biologically, the topological constraints of DNA - quantified by linking number, twist, and writhe - are analyzed alongside the mechanochemical action of DNA topoisomerases and the recent advancements in structural biology driven by artificial intelligence models like AlphaFold 3. Finally, the report contrasts the theoretical elegance of anyonic braiding with the formidable experimental challenges of hardware realization in TQC, highlighting breakthroughs such as Quantinuum's trapped-ion processors and Microsoft's Majorana 1 architecture. Through this synthesis, the pervasive principle that topological constraints dictate physical behavior and protect physical states from localized decoherence is thoroughly elucidated.
Introduction: The Dichotomy of Mathematical and Physical Knots
Topology investigates properties of space that are preserved under continuous deformations, such as stretching, twisting, and bending, without tearing or gluing operations 1. Within this domain, knot theory studies the embedding of closed one-dimensional manifolds (circles) into three-dimensional Euclidean space ($\mathbb{R}^3$) or the 3-sphere ($S^3$) 23. The discipline finds its deepest roots in the late 19th-century hypotheses of Lord Kelvin, who proposed that atoms were knotted vortices in the luminiferous aether. While physics quickly moved past the aether theory, the mathematics of knots developed into a rigorous, abstract field that eventually found its way back into the core of modern physical sciences.
A foundational distinction must be made between mathematical knots and physical knots. A mathematical knot is an infinitely thin, one-dimensional idealized object devoid of physical attributes such as volume, friction, bending stiffness, or thermal energy 34. Two mathematical knots are considered equivalent (ambient isotopic) if one can be transformed into the other via a sequence of continuous deformations without the curve ever intersecting itself 56. Conversely, physical knots - such as those formed by DNA molecules, filamentous polymers, macroscopic ropes, or magnetic flux tubes - are subject to strict metric and energetic realities 3. They possess finite thickness (a defined tube radius), excluded volume interactions, and thermodynamic constraints 47. While mathematical knot theory relies on discrete topological invariants to classify embeddings, the analysis of physical knots necessitates quantitative metric data, energetic considerations, and non-equilibrium dynamics to understand how spatial constraints dictate systemic function 38.
The translation of mathematical knot theory to the physical world has profound implications across length scales. In quantum field theory, the trajectory of particles through spacetime can be modeled as knotted or braided worldlines, introducing fractional statistics 9. In biological systems, the vast length of genomic DNA tightly packed into cellular compartments necessitates a topological management system to resolve the severe entanglement that naturally arises during transcription and replication 910. Bridging the gap between the purely topological, scale-invariant mathematics of knots and the highly constrained, scale-dependent physical universe provides a universal language for phenomena ranging from subatomic particle exchanges to the macroscopic behavior of long-chain polymers.
Theoretical Foundations of Knot Theory
Reidemeister Moves and Topological Equivalence
The central problem of knot theory is determining whether two distinct two-dimensional projections (diagrams) represent the same underlying three-dimensional knot. In 1926, J.W. Alexander and G.B. Briggs, and independently Kurt Reidemeister in 1927, proved that two knot diagrams describe the same knot if and only if they can be transformed into one another by a finite sequence of three local operations, known as Reidemeister moves, combined with planar isotopy 5611.
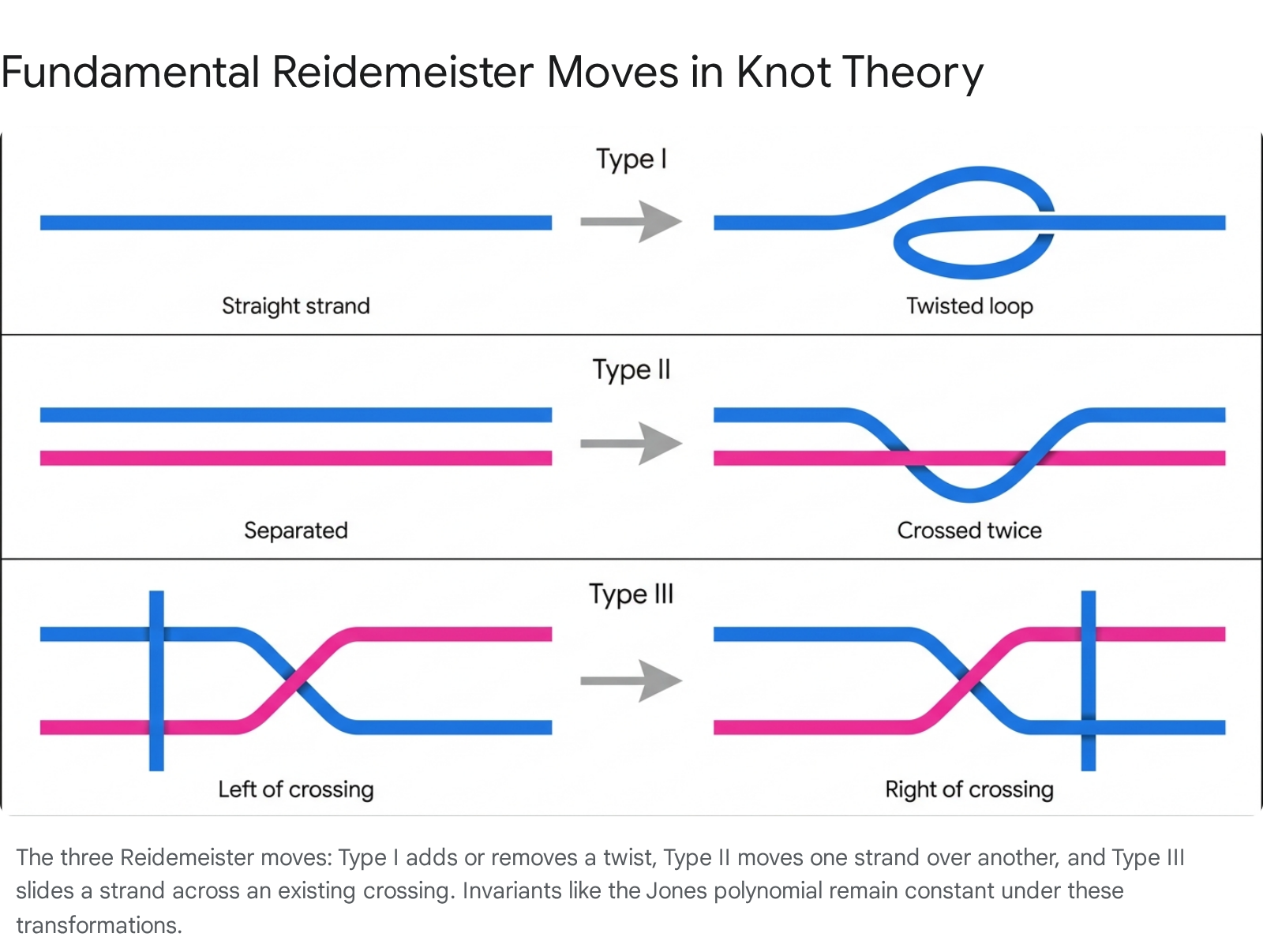
- Type I Move (Twist): Involves adding or removing a simple twist in a single strand. This is the only move that alters the writhe (the number of positive crossings minus negative crossings) of the diagram 26.
- Type II Move (Poke): Involves sliding one loop completely over or under another strand, adding or removing two crossings simultaneously without altering the overall writhe 612.
- Type III Move (Slide): Involves sliding a single strand completely over or under an existing crossing of two other strands. This move alters the relative positioning of the crossings but preserves the total crossing number and the writhe 612.
For framed links or physical ribbons, a modified Type I move (Type I') must be utilized, as standard twisting alters the framing of the manifold. This mathematical framing corresponds directly to physical torsion in applications like DNA supercoiling or the rotational phase of quantum particle worldlines 69.
Knot Polynomials and Categorification
To distinguish knots, mathematicians utilize topological invariants - properties that remain unchanged regardless of the sequence of Reidemeister moves applied to a diagram 612. A trivial example is tricolorability; however, polynomial invariants have proven to be exceptionally powerful tools for knot classification.
The Alexander polynomial, introduced in 1928, was the first significant polynomial invariant. It is computable via the determinant of a matrix derived from the crossings of the knot diagram and reflects properties of the knot group and the Alexander module 1415. Despite its computational efficiency, the Alexander polynomial fails to distinguish many knots, notably failing to differentiate between chiral pairs (a knot and its mirror image) 15.
The Jones polynomial, discovered by Vaughan F. Jones in 1984, revolutionized the field and earned him the Fields Medal. It is a Laurent polynomial in a single variable ($t$ or $q$) derived from the Kauffman bracket 21513. Because the Jones polynomial satisfies a specific skein relation, it successfully distinguishes many chiral knots, though it still fails to detect certain unknots 21513. Following this discovery, the HOMFLY (or HOMFLY-PT) polynomial was developed in 1985 as a two-variable generalization that encompasses both the Alexander and Jones polynomials 1415.
In the late 1990s, Mikhail Khovanov introduced a profound conceptual advancement: Khovanov homology. This theory acts as a "categorification" of the Jones polynomial. Instead of assigning a polynomial to a knot, it assigns a cochain complex of graded vector spaces (the Khovanov bracket) to the knot diagram 213. The polynomial acts merely as the shadow of this higher-dimensional algebraic structure. Specifically, the graded Euler characteristic of this homological complex recovers the unnormalized Jones polynomial 23. Crucially, Khovanov homology contains strictly more information than the Jones polynomial. It has been proven by Kronheimer and Mrowka to detect the unknot, it bounds the slice genus via the s-invariant, and it successfully distinguishes mutant knots that share identical Jones polynomials 1316.
Table 1: Comparison of Major Knot Invariants
| Invariant | Type | Historical Era | Categorification / Generalization | Application Domain & Capabilities |
|---|---|---|---|---|
| Alexander | Polynomial | 1920s (1928) | Knot Floer Homology | Algebraic topology. Fails to distinguish chirality. Easy to compute (determinant-based). 141514 |
| Jones | Polynomial | 1980s (1984) | Khovanov Homology | Connects to statistical mechanics/QFT. Detects chirality. Fails to detect all unknots. 21514 |
| HOMFLY(-PT) | 2-Variable Polynomial | 1980s (1985) | Khovanov-Rozansky Homology | Generalizes both Alexander and Jones polynomials. Distinguishes a vast number of knots. 151415 |
| Khovanov | Homology (Cochain Complex) | Late 1990s (1999) | Categorification of Jones | Detects the unknot. Bounds the slice genus (s-invariant). Connects to Floer homology. 231316 |
The Computational Complexity of Knot Invariants
While polynomial invariants are mathematically elegant, their exact algorithmic evaluation poses severe computational limits, serving as a bottleneck for large-scale data analysis of physical and mathematical knots. The computation of the Jones polynomial is formally classified as #P-hard 14171819.
The #P complexity class is the enumerative analogue of the NP decision class. A problem in #P counts the number of accepting paths of a non-deterministic Turing machine; therefore, a #P-hard problem is at least as computationally intractable as counting the exact number of satisfying assignments to a Boolean formula (#SAT) 14. Jaeger, Vertigan, and Welsh established the #P-hardness of the Jones polynomial by demonstrating its deep connection to the Tutte polynomial $T(M; x, y)$ of a planar graph. They proved that evaluating the Tutte polynomial is #P-hard at almost all points in the $(x, y)$-plane, except for a few specific curves 1418. Because the Jones polynomial of an alternating link corresponds to the Tutte polynomial of its associated planar graph evaluated along the hyperbola $xy = 1$, calculating the Jones polynomial inherits this extreme computational difficulty 1418.
The exact computation of the Jones polynomial requires exponential time, scaling at $O(2^c)$ where $c$ is the number of crossings in the knot or link diagram 1920. Even attempting to learn a linear number of the most significant bits of the Jones polynomial for a plat closure of an $n$-strand braid remains #P-hard 17. While highly parallelized algorithms have recently been proposed to reduce processing time by an exponential factor dependent on the number of available processors, the fundamental asymptotic limitation persists 1920.
However, this intractability in classical computing opens a direct avenue to quantum computation. Freedman, Kitaev, and Wang, followed by Aharonov, Jones, and Landau, established that an efficient quantum algorithm can additively approximate the Jones polynomial evaluated at any principal root of unity 1718. Because this approximation is universal for quantum computation at a non-lattice, principal root of unity, the problem of distinguishing values of the Jones polynomial is intrinsically tied to the power of quantum computers, demonstrating that topological invariants are foundational to the complexity class BQP (Bounded-Error Quantum Polynomial-Time) 18.
Knot Theory in Quantum Field Theory and Statistical Mechanics
Chern-Simons Theory and Wilson Loops
The profound connection between pure knot theory and the physical universe was rigorously formalized in 1989 by physicist Edward Witten. Witten demonstrated that the Jones polynomial could be derived intrinsically from a three-dimensional topological quantum field theory (TQFT) known as Chern-Simons gauge theory 212223.
In a standard quantum field theory, observables depend heavily on the local metric of spacetime (distances and angles). In a TQFT, however, the physical observables are topological invariants of the manifold in which the theory operates, entirely independent of the chosen spacetime metric 2224. The classical Chern-Simons action $S$ for a gauge connection $A$ over a 3-manifold $M$ is defined using exterior derivatives and wedge products of the gauge field, completely bypassing the need for a metric tensor 2224.
Witten demonstrated that the non-local observables in this theory, known as Wilson loops, correspond perfectly to mathematical knots and links 212224. A Wilson loop essentially measures the path-ordered phase acquired by a particle (transforming in some representation $R$ of a gauge group $G$) as it moves along a closed trajectory $K$ in spacetime 24. By evaluating the vacuum expectation values (vevs) of these Wilson loops within a functional path integral formulation using the non-Abelian gauge group SU(2), Witten miraculously recovered the precise algebraic structure of the Jones polynomial 2123. Furthermore, perturbative expansions of the Chern-Simons action in powers of the inverse coupling constant $1/k$ revealed that the coefficients of the resulting Feynman diagram series exactly correspond to Vassiliev (finite-type) knot invariants 21.
The Yang-Baxter Equation and Vertex Models
In the realm of statistical mechanics, the topology of knots arises through the formulation of exactly solvable two-dimensional lattice models via the Yang-Baxter equation 52825. In a 2D vertex model, physical interactions at lattice intersections - such as electron scattering or spin exchanges - can be mathematically encoded in an $R$-matrix 528. If the physical system is integrable, this $R$-matrix satisfies the Yang-Baxter equation: $R_{12} R_{13} R_{23} = R_{23} R_{13} R_{12}$.
Remarkably, this algebraic equation is geometrically identical to the Type III Reidemeister move (the slide move) 530. Consequently, the $R$-matrices of integrable statistical mechanical models provide linear representations of the braid group 2825. A braid represents a set of strands that intertwine as they move monotonically in one direction; by tying the start and end points of a braid together (the Markov closure), one generates a standard knot or link 5. Because the partition function of an integrable vertex model is invariant under the Markov moves applied to these braided worldlines, physicists can systematically derive knot invariants like the Jones and HOMFLY polynomials directly from the partition functions of the statistical system 52128.
Topological Dynamics in Biological Systems
The abstraction of knot theory encounters its most dynamic and vital physical realization within the nucleus of living cells. Deoxyribonucleic acid (DNA) is a plectonemic double helix, meaning its two antiparallel strands are intertwined. Because cellular DNA is either circular (in bacteria and plasmids) or anchored in complex chromatin loops (in eukaryotes), its ends are functionally fixed in space, preventing the free rotation of the strands 41031.
DNA Topology: Twist, Writhe, and Linking Number
The topological constraints of closed DNA are governed by the fundamental equation of DNA topology: $Lk = Tw + Wr$ 3132. * Twist (Tw): The total number of times the two strands of the DNA double helix cross each other around the central helical axis. In a relaxed state, biological DNA has roughly one twist every 10.4 base pairs 3132. * Writhe (Wr): The number of times the central double-helical axis crosses over itself in three-dimensional space. This metric physically manifests as supercoiling - the coiling of the coil itself 43132. * Linking Number (Lk): A strict topological invariant that defines the total number of times one strand wraps around the other. It must always be an integer 731.
As long as the sugar-phosphate backbone of the DNA remains intact, $Lk$ cannot change 31. Consequently, any local unwinding of the twist (which is absolutely required for molecular machines to read the genetic code during replication or transcription) forces a compensatory change in writhe to keep $Lk$ constant 910. This leads to severe torsional stress and the formation of positive supercoils ahead of the advancing replication fork or RNA polymerase 910. Supercoiling induces severe bending and torsional deformations. It is highly energetically unfavorable, as the free energy ($\Delta G$) associated with superhelical density increases quadratically 31. If left unresolved, this accumulated topological tension would physically stop the polymerases, stalling replication and causing cell death 3126.
Table 2: Mapping Topological Concepts to Physical and Biological Analogs
| Pure Topological Concept | Physical/QFT Analog | Biological (DNA) Analog |
|---|---|---|
| Knot / Link | Wilson Loop in Chern-Simons Theory 22 | Catenated Plasmids / Circular DNA 810 |
| Strand Crossing (Node) | Particle Interaction / Vertex Model 528 | Helix-Helix Crossover (Node) 4 |
| Topological Invariant | Non-local Observable (Metric Independent) 2224 | Linking Number ($Lk$) 731 |
| Braid Group Representation | Anyon Worldlines in (2+1)D Spacetime 92728 | Replication Fork Tangles 1029 |
| Crossing Change (Surgery) | Quantum Tunnelling / Reconnection | Topoisomerase Strand Passage 112630 |
| Reidemeister Type III Move | Yang-Baxter Equation / SU(3) Slide 530 | Localized Helix Deformations 1130 |
DNA Topoisomerases as Topological Machines
To resolve these existential topological crises, biological evolution produced DNA topoisomerases. These ubiquitous, highly conserved enzymes homeostatically manage topology by enacting discrete, mathematically equivalent "crossing changes" to the DNA strands 9263132.
Type I Topoisomerases perform single-strand breaks. They utilize an active-site tyrosine residue as a nucleophile to attack the phosphodiester backbone, forming a transient covalent enzyme-DNA intermediate known as a cleavage complex (TOPcc) 32263031. This covalent attachment preserves the energy of the broken bond. Type IA enzymes (which bind the 5' phosphate) pass the intact DNA strand through the break via an enzyme-bridging mechanism before resealing 2633. In contrast, Type IB enzymes (which bind the 3' phosphate) allow the free end to swivel or undergo controlled rotation around the intact strand, driven purely by the release of torsional stress 26. These enzymes are generally monomeric, ATP-independent, and alter the linking number $Lk$ strictly in increments of 1 2641.
Type II Topoisomerases (such as DNA gyrase and Topo IV) act as complex, multi-domain nanomachines that catalyze double-strand breaks. Operating as homodimers or heterotetramers, they bind a segment of duplex DNA (the G-segment or Gate segment), cleave both strands simultaneously, and actively pass another intact duplex (the T-segment or Transport segment) completely through the transient break 2630.
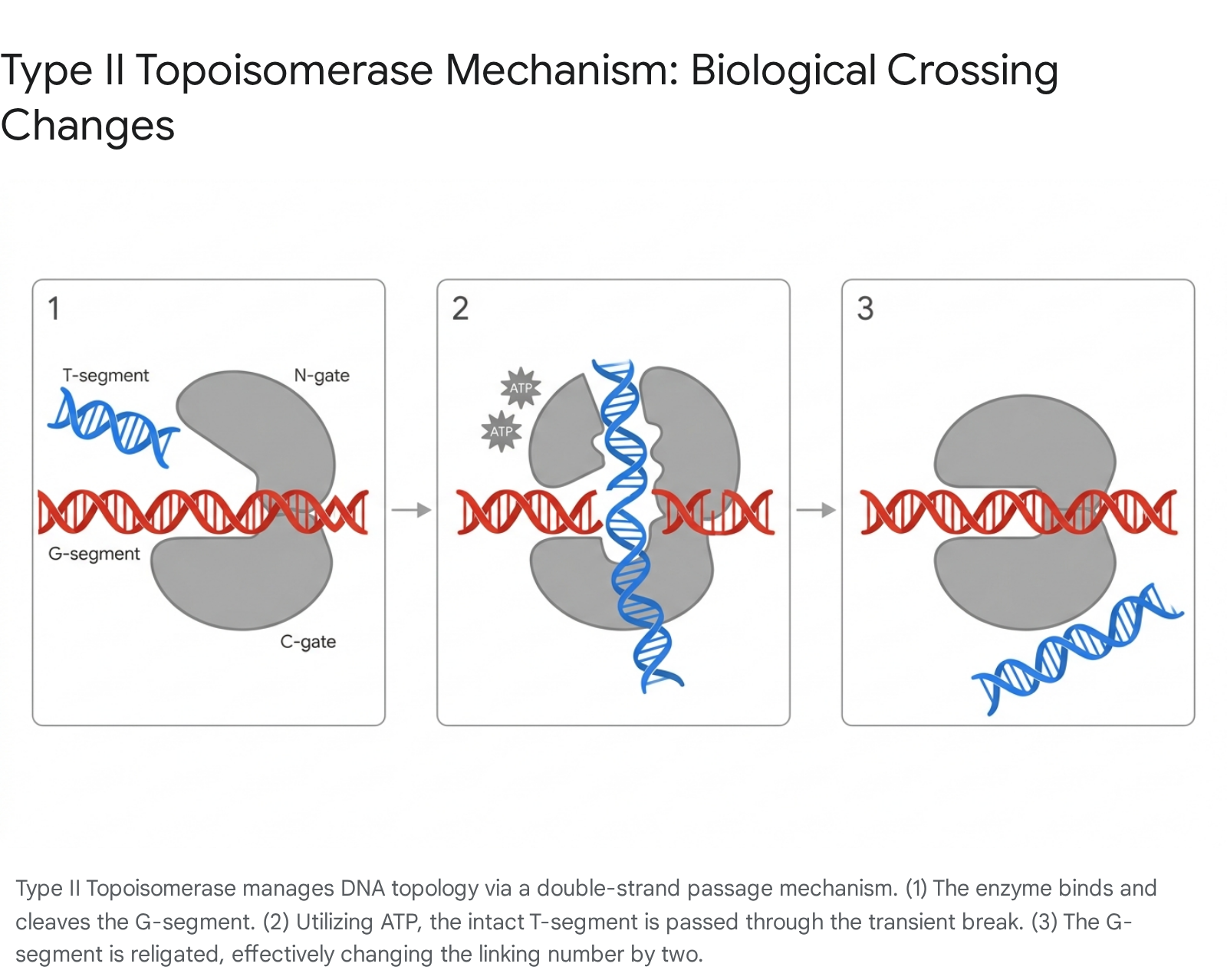
This process is energetically driven by the binding and hydrolysis of two ATP molecules 2630. This double-strand passage mechanism alters $Lk$ in increments of 2 and is the primary biological mechanism for knot resolution (unknotting) and chromosome decatenation (unlinking daughter chromosomes after replication) 3226.
Topoisomerase II functions effectively as a biological "Maxwell's demon." Theoretical models and single-molecule experiments indicate that type II enzymes relax DNA topologies to fractions far below what thermodynamic equilibrium would dictate (a phenomenon known as topology simplification) 83435. They achieve this directional, non-equilibrium sorting by inducing incredibly sharp bends (~130°) in the G-segment of the DNA upon binding 34. Because the enzyme has a specific geometric orientation relative to the bend it creates, it ensures unidirectional strand passage, actively shifting the system out of topological equilibrium using chemical energy 834.
Recent empirical breakthroughs have allowed researchers to witness these nanomachines in unprecedented detail. Using advanced cryo-electron microscopy and deep learning, researchers from KAUST recently captured the simian virus 40 (SV40) Large Tumor Antigen helicase in the act of DNA unraveling 3637. These models demonstrate that nanomachines do not pry DNA apart in one fluid motion; rather, ATP hydrolysis functions like a snapping spring, cycling through 15 distinct atomic conformational states that progressively reduce physical constraints, increase entropy, and allow the enzyme to traverse the DNA lattice, melting the topology base-pair by base-pair 3637.
Artificial Intelligence in Structural Topology: The AlphaFold 3 Era
The topology of biomolecules extends beyond the simple double helix to the complex three-dimensional folding of proteins and their interactions with ligands. The advent of artificial intelligence, notably DeepMind's AlphaFold 3 (AF3) utilizing Diffusion Transformer architectures, has revolutionized structural biology by extending prediction capabilities from pure protein sequences to protein-ligand complexes, DNA, RNA, and post-translational modifications 464748.
However, predicting and capturing deep topological complexity presents severe limitations for current AI architectures. Systematic benchmarking reveals a fundamental flaw: while AF3 excels at predicting static protein-ligand interactions with minimal conformational changes (significantly outperforming traditional physics-based docking in side-chain accuracy), it appears to memorize atomic positions from its training set rather than learning the generalized physical principles of molecular recognition 464749. In out-of-sample pose reproduction tests on over 8,000 complexes deposited after AF3's training cutoff, accuracy strongly correlated with training-set similarity 47. AF3 struggles profoundly with complexes involving significant conformational changes (>5Å RMSD) and demonstrates a persistent bias toward active GPCR conformations regardless of the introduced ligand 4647.
Mechanistic interpretability analysis proves that AF3 relies predominantly on comparative evolutionary context - the Multiple Sequence Alignment (MSA) - rather than raw sequence or thermodynamic physics. When MSAs are heavily degraded or removed, prediction accuracy collapses entirely, regardless of how familiar the sequence was in the training set 4950.
Particularly in topological domains like chirality, AF3 exhibits critical failures. In extensive predictions (over 3,200 experiments) involving D-peptides (mirrored chiral centers targeting biological L-proteins), AF3 recorded a staggering 51% chirality violation rate. Essentially, the model operates no better than random chance (randomly picking L or D for each residue) in preserving heterochiral folding, frequently generating structures with incorrect folds and physically impossible binding poses 51.
To overcome these topological blind spots, modern computational workflows require hybrid pipelines that merge deep learning with physical constraints. For instance, the AlphaFold 3x (AF3x) pipeline explicitly models distance restraints derived from cross-linking mass spectrometry (XL-MS) as covalently bound ligands within the AF3 framework. This physically forces the AI to comply with topological realities 4852. Similarly, the AlphaUnfold framework couples static AF3 predictions with short-time (5 ns) high-pressure molecular dynamics simulations (baric stress) using NAMD3. By subjecting the AI's models to thermodynamic stress, AlphaUnfold identifies structurally fragile "not well-forged" regions. The pipeline revealed a significant inverse correlation between AF3's static confidence score (pLDDT) and the Root Mean Square Deviation (RMSD) after molecular dynamics, proving that low-confidence AI predictions rapidly undergo structural drift when subjected to physical physics 53. Consequently, deep learning represents the beginning, not the end, of structure-guided discovery, requiring complementary physics-based refinement 47.
Topological Quantum Computing: From Theory to Hardware
Anyonic Braiding and Theoretical Mathematics
Topological quantum computing (TQC) seeks to encode and manipulate quantum information in a way that is intrinsically protected from local environmental decoherence by the global topological properties of the system 275455. The physical substrate for TQC relies on two-dimensional quasi-particles known as anyons 52854.
In standard 3D space, fundamental particles are strictly categorized as either bosons (yielding a phase of $+1$ upon exchange) or fermions (yielding a phase of $-1$). This dichotomy is dictated by the symmetry or antisymmetry of their wavefunctions upon particle exchange 52728. However, when particles are tightly constrained to a 2D spatial plane (making a (2+1)-dimensional spacetime), the rules of exchange change dramatically. The exchange of identical particles in 2D is no longer described by the permutation group, but by the braid group. The paths of the particles cannot pass through one another in 2D without intersecting, meaning their worldlines in spacetime weave together to form a permanent physical braid 52554.
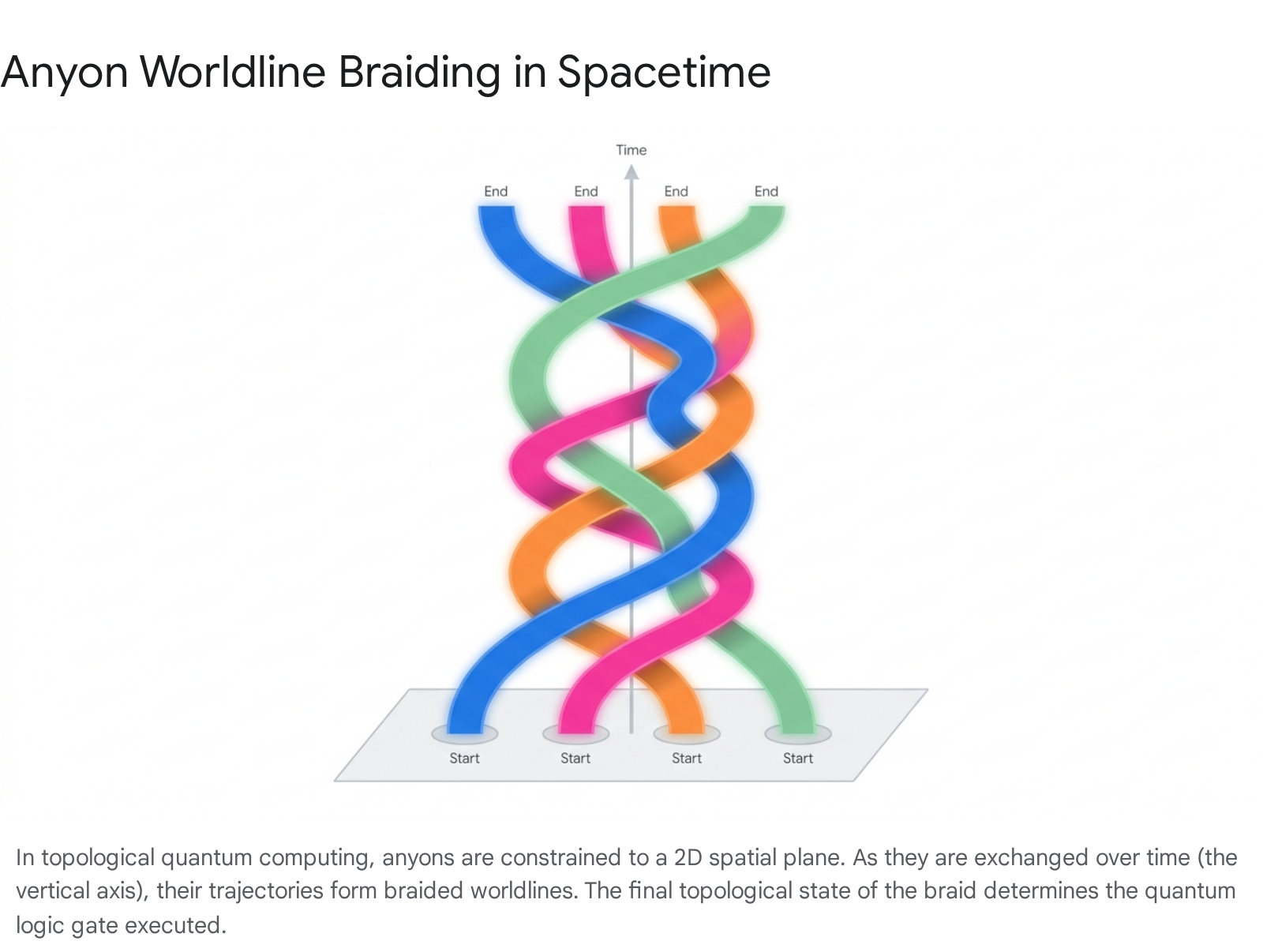
This topological reality allows for the emergence of anyons - particles that can accrue any arbitrary fractional phase upon exchange (Abelian anyons). More profoundly, it allows for non-Abelian anyons. When non-Abelian anyons are exchanged (braided), the order of the exchange matters ($A \times B \neq B \times A$). Their worldlines intertwine, and the global wavefunction of the system undergoes a complex, non-commutative unitary rotation within a highly degenerate ground state manifold 275456. Because these quantum logic operations depend purely on the topology of the braid (which particles moved around which) and are utterly invariant to minor physical perturbations, trajectory distortions, or timing fluctuations, TQC offers a theoretical pathway to fault tolerance built directly into the hardware 275455.
Experimental Challenges and Recent Breakthroughs in Hardware
Translating the elegant mathematics of SU(2) Witten-Chern-Simons theory and modular functors into functional hardware has proven to be an exceptional engineering challenge 252854. For decades, the only naturally occurring candidates thought to host non-Abelian anyons were fractional quantum Hall (FQH) states (e.g., the $\nu = 5/2$ state in 2D electron gases subjected to extreme magnetic fields and cryogenic temperatures) 27283839. However, isolating and controlling these quasiparticles experimentally remained elusive. The period of 2024 - 2026 has witnessed unprecedented, paradigm-shifting milestones in the synthesis of topological orders.
Quantinuum's Trapped-Ion Anyons: In 2024, researchers from Quantinuum, collaborating with Harvard and Caltech, utilized their H2 trapped-ion processor to simulate the first unambiguous realization of non-Abelian topological order. Using a kagome lattice of 27 qubits and a shallow adaptive circuit, they prepared the ground state wavefunction of $D_4$ topological order with a fidelity exceeding 98.4% per site 596040. Utilizing mid-circuit measurements, the team dynamically created and moved anyons along Borromean rings in spacetime. Anyon interferometry successfully detected the intrinsically non-Abelian braiding process 6040. The ability of the H-Series architecture - characterized by all-to-all connectivity and Quantum Charge-Coupled Device (QCCD) routing - to perform rapid mid-circuit feed-forward operations was critical to simplifying the pathway to these exotic states without being overwhelmed by circuit error rates 626364.
Microsoft's Majorana 1 and Topoconductors: In February 2025, Microsoft achieved a completely distinct paradigm of hardware-level TQC with the unveiling of Majorana 1, the world's first quantum processor built on a topological core 416768. The chip utilizes a Tetron design comprising parallel indium arsenide (InAs) semiconductor nanowires proximitized by an ultra-thin aluminum (Al) superconducting layer. Microsoft termed this novel materials stack a "topoconductor" 68424344. When cooled near absolute zero and subjected to precise magnetic fields (approx. 1.8 T), the device enters a state of topological superconductivity, fostering Majorana Zero Modes (MZMs) at the open ends of the nanowires 427245.
Because an MZM mathematically acts as its own antiparticle, quantum information can be non-locally encoded in the shared fermionic parity of four coupled Majoranas distributed across the vacuum gaps of the Tetron 564243. Instead of physically moving the Majoranas - a process known as analog braiding, which is highly susceptible to thermal jitter and material disorder - Microsoft achieved effective braiding through digital, measurement-based control 72. By performing orthogonal Pauli $Z$ and $X$ measurements via microwave reflectometry, they demonstrated non-commuting operations that mathematically alter the quantum state, proving the viability of measurement-based braiding 4245.
This architecture allows Microsoft to project scaling up to 1 million qubits on a single, palm-sized chip. The intrinsic hardware-level protection provided by the topological constraints means that their custom quantum error correction (QEC) codes reduce overhead roughly tenfold compared to the standard surface codes utilized by competitors like Google and IBM 41674574. Despite some skepticism within the physics community regarding the clarity of the MZM signal amidst environmental noise (such as spacetime torsion effects or surface roughness generating spurious signals), the Defense Advanced Research Projects Agency (DARPA) has subsequently backed Microsoft under the US2QC program to build a fault-tolerant prototype (FTP) within years, not decades 4246.
Discussion: A Cross-Disciplinary Synthesis
The pervasive appearance of knot theory across discrete scientific disciplines underscores a profound universal principle: topology protects state.
In pure mathematics, topological invariants filter out the noisy, irrelevant details of continuous geometric distortion to classify the fundamental, immutable essence of a space. In statistical mechanics and QFT, the algebraic invariance of the Yang-Baxter equation and the explicit metric-independence of the Chern-Simons action ensure that macroscopic observables remain robust against microscopic, localized perturbations 52228.
This theoretical robustness is physically exploited by biological evolution inside the nucleus, where topoisomerases maintain the homeostatic linking number of DNA. Just as mathematical knots rely on discrete crossing changes (Reidemeister moves) to alter their invariants, topoisomerases consume ATP to drive non-equilibrium topological surgeries, ensuring genomic stability amidst the chaotic, high-stress unspooling required for DNA replication 82634.
Finally, human engineers are now attempting to harness this exact principle to build fault-tolerant quantum computers. Whether simulating synthetic anyons in a QCCD ion trap or engineering Majorana Zero Modes in a hybrid semiconductor-superconductor topoconductor, the goal is identical to the biological management of DNA: to store information non-locally. By distributing a quantum state across a topological manifold, computational systems can be rendered immune to localized environmental decoherence, mirroring the exact topological protections that nature has utilized for billions of years 3154.
Conclusion
Knot theory has transitioned from a specialized enclave of pure mathematics into a vital operational framework across physics, biology, and computer science. The extreme difficulty of evaluating fundamental invariants like the Jones polynomial highlights the profound complexity inherent in topologically entangled systems - a complexity that acts as a severe barrier in classical algorithmic prediction but serves as a vital protective mechanism in the physical world. In biology, topoisomerases act as molecular guardians, executing topological surgeries to resolve the life-threatening entanglement of DNA driven by its plectonemic structural constraints. In computational biology, while AI models like AlphaFold 3 push the boundaries of spatial prediction, they require targeted biochemical and thermodynamic constraints (like those introduced by AlphaUnfold) to resolve complex topological ambiguities accurately. Ultimately, the translation of theoretical braiding statistics into experimental reality via non-Abelian anyons and Majorana Zero Modes represents a paradigm shift. By encoding logic into the immutable fabric of topology, researchers are on the threshold of bypassing the fragility of classical quantum states, paving the way for the fault-tolerant foundation era of quantum computing.