Information theory in genetics, neural coding, and evolution
The application of Claude Shannon's mathematical theory of communication to the biological sciences represents one of the most profound interdisciplinary convergences of the past century. Originally formulated in 1948 to determine the fundamental limits of signal processing and data compression in artificial telecommunications channels, information theory quickly permeated molecular and systems biology. Early adoptions were largely metaphorical, with the "central dogma" of molecular biology conceptualized as a unidirectional transfer of information from DNA to RNA to protein. However, contemporary biological research has moved far beyond analogy, employing the rigorous mathematics of entropy, mutual information, and channel capacity to quantify the absolute physical limits of life 12.
From the macro-level organization of the mammalian cortex to the micro-level dynamics of intracellular signaling cascades, organisms operate as complex information-processing engines. They must continuously sample a noisy environment, encode these physical or chemical stimuli into internal representations, and transmit these representations through heavily constrained biochemical networks to actuate survival behaviors. Because information transmission is a physical process, it is inextricably bound by the laws of thermodynamics. Acquiring information, processing it, and ultimately erasing it from cellular memory requires the expenditure of metabolic energy, establishing a fundamental trade-off between the fidelity of a biological signal and the metabolic resources an organism can afford to spend 343.
This report examines the deep legacy of information theory across multiple scales of biology. It investigates the principles of efficient neural coding, the precise measurement of channel capacities in cellular signaling networks, the thermodynamic constraints imposed by Landauer's principle on biological memory erasure, and the ongoing theoretical efforts to define biological meaning through metrics like functional information.
Mathematical Foundations of Biological Information
To apply information theory to biology, researchers must carefully select the mathematical frameworks that best capture the phenomena under investigation. Biological systems pose unique challenges to classical information theory because they are far-from-equilibrium thermodynamic systems operating with finite resources in highly noisy environments 45.
Shannon Entropy, Kolmogorov Complexity, and Mutual Information
Shannon entropy measures the fundamental uncertainty or randomness inherent in a probability distribution, effectively setting the theoretical upper limit on the capacity of a communication channel. In biological sequence analysis and physiological time-series analysis, Shannon entropy remains a foundational metric. However, because Shannon entropy is a global measure, it is often insensitive to local structural patterns. Large changes in a probability distribution over a small range have minimal impact on the overall Shannon entropy, making it sometimes insufficient for capturing the nuanced dynamics of complex biological systems 67.
To more accurately characterize system dynamics, researchers frequently employ complementary measures of structural complexity. Methods such as dispersion entropy and permutation entropy have been developed to capture ordinal structures and time-delayed mutual information in physiological signals, such as electroencephalography data 68. Furthermore, Kolmogorov complexity, a concept from algorithmic information theory, offers a different perspective by defining the complexity of a biological sequence based on the length of the shortest computational program required to generate it. While true Kolmogorov complexity is mathematically non-computable, compression algorithms are widely used to estimate the relative complexity and informational density of genomic sequences, providing a metric for the degree of redundancy and compressibility of genetic codes 19.
Mutual information represents the reduction in uncertainty about one variable given knowledge of another. In biological signaling, it serves as the primary metric for signal fidelity, quantifying the degree of correlation between an environmental input and a cellular output 35. Estimating mutual information in high-dimensional biological data is notoriously difficult due to finite sample sizes, which can introduce significant variance depending on the binning strategies used for probability distribution estimation 5. To circumvent these computational bottlenecks, researchers have increasingly turned to analytical approximations using Fisher Information, which provides closed-form formulas to benchmark the theoretical limits of sensing without requiring computationally expensive mutual information estimations 12.
| Information Metric | Core Definition | Biological Application | Limitations |
|---|---|---|---|
| Shannon Entropy | The measure of average uncertainty or randomness in a given probability distribution. | Quantifying ecological diversity, genetic sequence capacity, and overall channel limits. | Fails to account for local structural patterns; ignores the functional "meaning" or biological utility of the data 613. |
| Kolmogorov Complexity | The length of the shortest computer program capable of generating a specific dataset or sequence. | Estimating the compressibility of genomes; identifying non-random algorithmic structures in DNA. | Mathematically non-computable in its pure form; requires approximation via compression algorithms 19. |
| Mutual Information | The quantification of the amount of information obtained about one random variable through observing another. | Measuring the fidelity of signal transduction; determining how well a cell predicts external ligand concentrations. | Computationally expensive to estimate from limited biological sample sizes, particularly in high dimensions 5. |
| Functional Information | The probability that an arbitrary configuration of a system will achieve a specific biological function. | Evaluating the emergence of complexity in origin-of-life models and ribozyme evolution. | Requires the explicit definition and measurement of a specific degree of function prior to calculation 1015. |
Neural Coding and Sensory Channel Capacity
Sensory systems are the primary interfaces through which organisms acquire information about the external world. Because the physical environment is infinitely complex and biological bandwidth is finite, nervous systems face a severe data compression problem. Information theory provides the exact mathematical language required to understand how neural circuits solve this optimization challenge.
The Efficient Coding Hypothesis
The efficient coding hypothesis, first proposed by Horace Barlow in 1961, posits that the evolutionary pressure to maximize information transfer while minimizing metabolic cost has heavily shaped the receptive fields and firing patterns of sensory neurons. Under this framework, the sensory pathway is treated as a communication channel where neuronal spiking serves as a code designed to represent sensory signals. To maximize the available channel capacity, this code must minimize the redundancy between representational units, adapting specifically to the statistical regularities of the natural environment 11.
Experimental validations of this hypothesis have been particularly successful in the early visual system. Receptive fields in the mammalian retina and the visual cortex exhibit spatial and temporal filtering properties that closely match the optimal theoretical filters for processing natural images. Retinal ganglion cells, the output neurons of the retina, group into dozens of functionally distinct mosaics that tile the visual field. Recent analyses demonstrate that information transmission in these retinal ganglion cells operates at an overall efficiency of approximately 80%, indicating that the functional connectivity between photoreceptors and retinal ganglion cells closely approaches the theoretical Shannon limit given the noise conditions inherent in the biological hardware 1112.
The organization of these neural mosaics is highly dependent on both channel capacity limitations and the inherent noise of the biological hardware. When channel capacity is constrained, theoretical models predict that sensory populations should split into parallel pathways, such as ON and OFF cells that signal light increments and decrements, respectively. Depending on the asymmetries in the natural stimulus distribution and the baseline noise level, the optimal thresholds for these ON and OFF populations shift to minimize decoding error and maximize mutual information. This theoretical prediction elegantly accounts for the diverse biases and threshold distributions observed experimentally across different sensory modalities 1314. Furthermore, empirical studies on primate retinas demonstrate that the detection of complex environmental features, such as approaching motion, begins surprisingly early in the visual pathway, driven by specific retinal ganglion cell types whose synaptic currents optimize the extraction of these ethologically relevant signals 15.
Noise Correlations and Information Optimization
Because biological sensors rely on stochastic processes, such as the random binding of photons to rhodopsin or the probabilistic opening of voltage-gated ion channels, individual sensory neurons are inherently noisy. A fundamental strategy to improve the signal-to-noise ratio is population coding, where downstream circuits pool the responses of multiple noisy receptors 1617. However, this pooling mechanism is complicated by noise correlations, wherein the noise across different neurons is not independent but shared, often due to overlapping inputs or dense interconnectivity.
If a neural decoder assumes that neuronal noise is uncorrelated when it is actually correlated, the resulting information loss can be catastrophic. Studies in the retina have shown that failing to account for noise correlations can degrade decoding performance by a massive margin, particularly at low light levels where the signal is weakest and the correlation structure carries the most relative information. To decode these signals effectively, the brain relies on adaptive noise filters that change with environmental conditions, allowing neural populations to extract significantly more accurate representations of the stimulus 18.
Neural circuits also actively counter these correlations through specific anatomical architectures. For instance, inhibitory feedback in recurrent neural networks serves to actively decorrelate the population activity, thereby increasing the net mutual information transmitted by the network 1925.
This active decorrelation is an energy-intensive process. Inhibitory activity decreases the overall gain of the neuronal population, meaning that depolarization requires stronger excitatory synaptic input. This heightened excitation is directly associated with higher adenosine triphosphate consumption. The brain must therefore constantly balance the increased reliability of the decorrelated signal against the heightened metabolic expense. Optimization models maximizing mutual information under stringent metabolic cost constraints confirm that there is an optimal strength of recurrent connections that maximizes the bits-per-cost ratio.
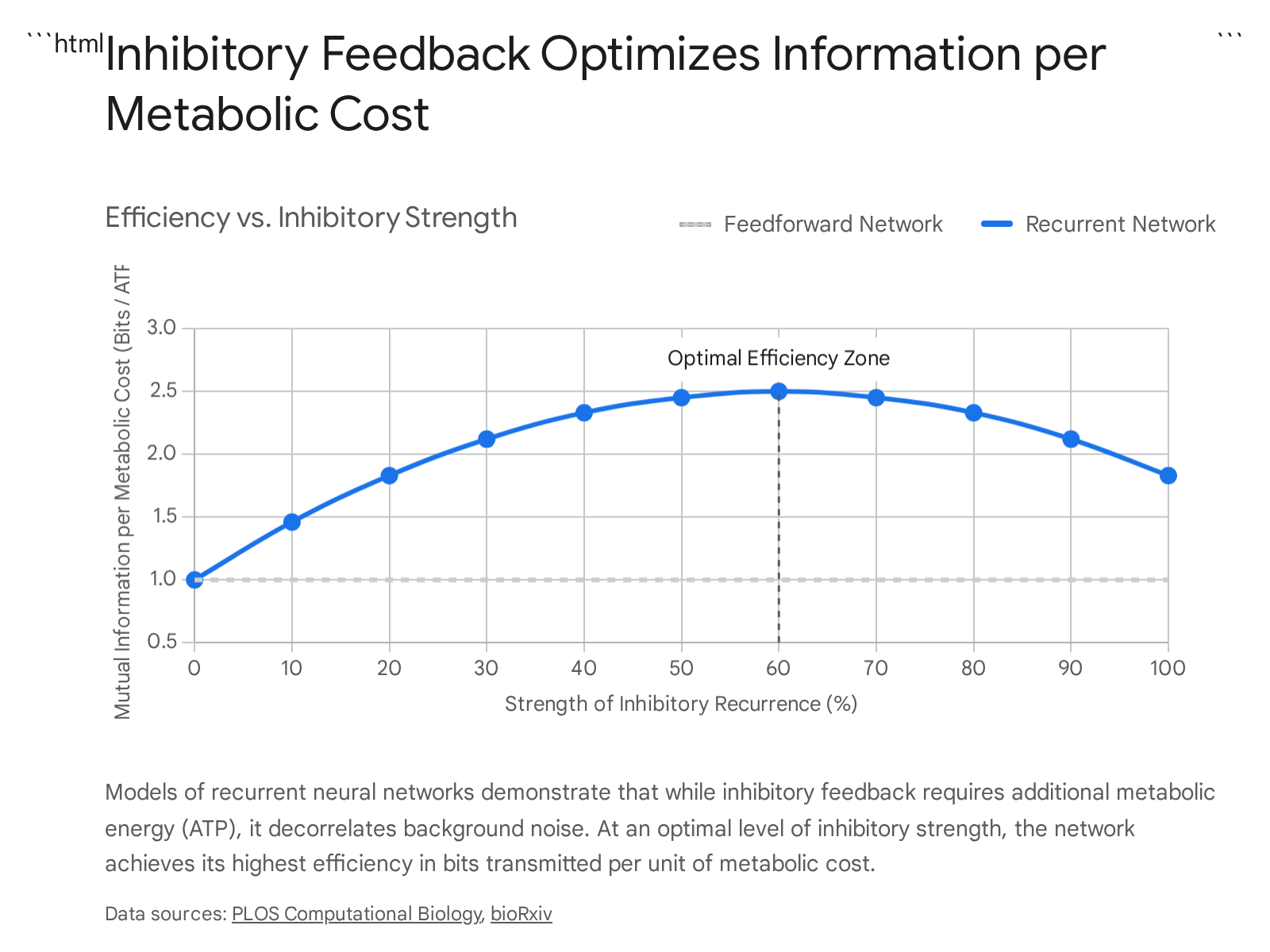
This demonstrates that the optimal synaptic strength of a recurrent network can be inferred purely from metabolically efficient coding arguments 1925.
The Strict Metabolic Cost of Biological Information
While classical information theory treats the bit as an abstract mathematical entity, in biological systems, every bit represents a physical change in state. Transmitting information requires pushing a cellular system out of thermodynamic equilibrium. In neural systems, this requires the continuous action of ion pumps to maintain steep electrochemical gradients across cellular membranes.
The human neocortex utilizes approximately 20% of the body's total resting energy consumption, metabolizing glucose at a rate that yields roughly 3.4 * 1021 molecules of adenosine triphosphate per minute. A highly significant fraction of this energy, estimated to be up to 80% of the total cortical budget, is devoted exclusively to signaling processes, namely the maintenance of resting potentials and the generation of discrete action potentials 202721. Cortical neurons are estimated to spike at exceptionally low average rates, approximately 0.16 times per second, strictly due to the massive metabolic burden associated with each spike 27.
Biophysical measurements provide staggering estimates of the true thermodynamic cost of biological information. Groundbreaking intracellular recordings from the blowfly retina have allowed researchers to map the exact number of ATP molecules hydrolyzed to transmit a specific quantity of mutual information.
| Biological Signaling Mechanism | Estimated Metabolic Cost | Reference System |
|---|---|---|
| Chemical Synapse Transmission | ~104 ATP per bit | Blowfly photoreceptor synapses 2022 |
| Graded Analog Signals | 106 to 107 ATP per bit | Invertebrate interneurons 2022 |
| Spike Coding (Action Potentials) | 106 to 107 ATP per bit | Mammalian cortical neurons 2022 |
| Action Potential Generation | ~2.4 * 109 ATP per spike | Human neocortex estimates 27 |
| Memory Formation (Long-Term) | ~10 mJ per bit | Drosophila associative conditioning 23 |
These empirical measurements reveal a profound inefficiency when compared to the absolute thermodynamic minimum required for computation, known as the Landauer limit. Energy consumption in neural signaling is several orders of magnitude greater than the theoretical physical floor 2022. Furthermore, neural systems are subject to a strict law of diminishing returns: pushing a neuron to transmit at higher bit rates requires an exponentially greater expenditure of metabolic energy.
Because high-capacity pathways are metabolically exorbitant, noise-limited biological systems strongly favor weak pathways of low capacity. It is vastly more economical to distribute information across multiple parallel, low-capacity neurons than to force a single neuron to operate near its absolute maximum channel capacity 2022. This basic thermodynamic reality explains the massive parallelization universally observed in animal brains.
Sensory versus Kinematic Trade-offs
The metabolic cost of information does not solely dictate internal neurological wiring; it actively shapes organismal behavior and physical morphology. For an organism to acquire visual, auditory, or electrical information about its environment, it must either evolve highly acute, metabolically expensive stationary sensors, or it must expend mechanical energy to move its sensors through space.
Studies on aquatic models, such as electric fish, beautifully illustrate this dynamic. When searching for prey in environments beyond the range of their immediate sensory fields, these fish will adopt energetically inefficient swimming patterns that significantly increase hydrodynamic drag. This mechanical inefficiency is actively chosen because moving the body in this specific, exaggerated manner sweeps the fish's sensorium across a larger volume of space per unit of time, dramatically increasing the prey encounter rate. The energetic yield obtained from the increased information acquisition outweighs the mechanical penalty of inefficient swimming 24. This behavior underscores how the pursuit of information mathematically governs macro-ecological survival strategies, forcing a strict trade-off between the metabolic cost of sensing and the metabolic cost of locomotion.
Intracellular Signaling and Molecular Computation
Beyond the central nervous system, single cells face identical information-theoretic challenges. Cells exist in highly variable, noisy environments and must reliably measure the concentrations of hormones, cytokines, nutrients, and pathogens. They process these inputs through intricate biochemical cascades composed of kinases, transcription factors, and secondary messengers. The consistently small channel capacities observed in most cellular signaling pathways indicate that cells often operate with a fairly coarse representation of their surroundings. Measuring mutual information at single time points typically yields transmission capacities of merely 1 to 3 bits, meaning the pathway can only reliably distinguish between a handful of distinct concentration levels 3.
Channel Capacity in MAPK and NF-κB Pathways
In mammalian immune cells, such as macrophages, the detection of pathogens relies heavily on Toll-like receptors which feed into two primary downstream signaling pathways: NF-κB and Mitogen-Activated Protein Kinases (MAPK). Information theory has been instrumental in deciphering how cells use these two distinct pathways simultaneously to discriminate between mild environmental noise, such as the presence of commensal bacteria, and severe pathogenic threats 2533.
Mutual information calculations on these pathways reveal distinct processing strategies tailored to channel capacity limits. The MAPK channel capacity is estimated to be highly robust, capable of transmitting between 6.0 and 8.5 bits per hour, depending on the temporal encoding protocol utilized by the cell 26. In response to low concentrations of stimulatory ligands, the NF-κB pathway activates in a graded manner, initiating a limited, non-inflammatory gene expression profile that primes the macrophage without triggering immediate, tissue-damaging inflammation. However, when the ligand concentration exceeds a critical threshold, the MAPK pathway activates in a sharp, switch-like manner, facilitating the massive production of inflammatory mediators 25.
By partitioning the input concentration space, the cell utilizes NF-κB for low-threshold, low-risk information processing, and reserves the higher-capacity, ultrasensitive MAPK pathway as a rigid inflammatory threshold.
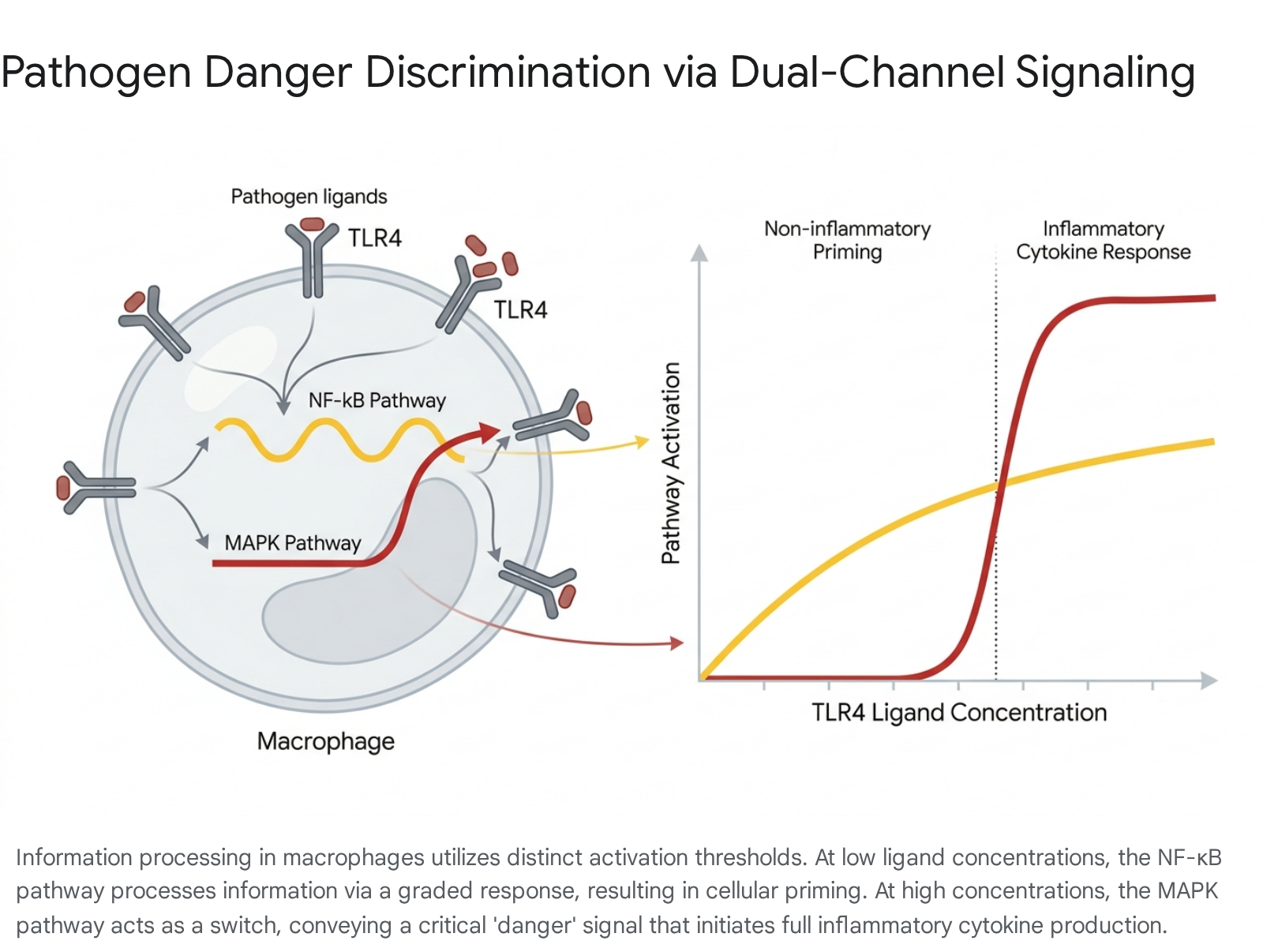
This architecture ensures that the mutual information between the pathogen load and the cellular response is optimized to prevent autoimmune damage while preserving a robust defensive response when genuine threats are detected 25.
Bacterial Signaling and the Cost of Transduction
Information theory applies equally to prokaryotic life. Bacteria commonly utilize Two-Component Systems to detect physical and chemical stimuli, relying on a sensor kinase and an internal response regulator. Interestingly, bacteria frequently deploy signaling architectures that are highly energetically expensive to operate, despite facing severe evolutionary pressure for metabolic efficiency.
Bond graph modeling of energy flow through these bacterial systems reveals that the availability of adenosine triphosphate directly governs the network's information processing parameters. When internal energy concentrations drop, the Hill coefficient of the system shifts dramatically, increasing the network's sensitivity to weak stimuli but severely lowering the maximum possible signal response 27. Thus, energetically expensive signaling variants are maintained by bacterial evolution because they afford superior noise-filtering capabilities and broader dynamic ranges in energy-rich environments. This proves that cellular networks can dynamically scale their information capacity based directly on available metabolic fuel 27.
Collectivity in bacteria also exhibits advanced signaling that mirrors neurological information processing. Dense bacterial communities known as biofilms experience profound metabolic stress when internal nutrients are depleted. To coordinate a colony-wide response, biofilms of Bacillus subtilis generate synchronized electrical signaling waves. Starving cells in the biofilm interior release intracellular potassium ions, which diffuse to neighboring cells and trigger local membrane depolarization. This triggers further potassium release, creating a continuous bucket-brigade wave of electrical information that propagates from the core to the extreme periphery of the colony. Because potassium and glutamate are central to this bacterial signaling, and are identical to the primary excitatory mediators in mammalian brains, the data suggests a profound evolutionary conservation of ion-based information processing spanning the entire tree of life 28.
Thermodynamic Constraints on Biological Computation
If information transmission costs energy, what are the thermodynamic implications for information storage and erasure? In 1961, IBM physicist Rolf Landauer established a strict physical bridge between information theory and thermodynamics. Landauer's principle states that any logically irreversible operation, such as the erasure of a single bit of classical memory or the merging of two computational paths, must inherently dissipate a minimum amount of heat into the surrounding environment 29383031.
The Landauer bound is mathematically defined as $E \geq k_B T \ln 2$, where $k_B$ is the Boltzmann constant and $T$ is the absolute temperature of the thermal bath 2938303233. This principle has been rigorously validated in experimental settings, most notably in 2012 when researchers used a single colloidal particle trapped in a modulated double-well optical potential to demonstrate that the mean dissipated heat during an erasure cycle precisely saturates at the Landauer bound in the limit of long erasure times 3435. While modern silicon computers operate billions of times above this absolute limit, biological systems operate remarkably closer to the thermodynamic floor. Furthermore, recent experiments employing quantum field simulators of ultracold Bose gases have successfully tracked the temporal evolution of quantum fields to confirm the validity of Landauer's principle in complex quantum many-body regimes, reinforcing its status as a universal physical law 36.
The Cost of Erasing Cellular Memory
A classic problem in biological physics is the Berg-Purcell limit, which defines the theoretical maximum accuracy with which a cell, such as a chemotactic bacterium, can measure the concentration of a chemical ligand in its surrounding environment via surface receptors.
For the cell to continuously track temporal changes in a highly dynamic environment, it cannot simply accumulate measurements infinitely; it must continuously integrate new data and completely "forget" old data. According to Landauer's principle, this erasure of past molecular observations is a logically irreversible process that strictly mandates the breaking of detailed thermodynamic balance. The biochemical network executing the computation must consume continuous energy to reset its internal sensors. Calculations of simple two-component cellular networks confirm that greater learning, defined as achieving higher mutual information between the internal cellular state and the external chemical concentration, requires proportionally greater energy consumption strictly to erase the outdated molecular states 33738.
In resource-poor biological environments, the thermodynamic cost of cellular memory erasure acts as a severe evolutionary constraint. Synthetic biologists attempting to engineer novel signaling networks are heavily limited by these fundamental physical laws. By understanding that energy must be spent to reset post-translational modifications, such as utilizing adenosine triphosphate to dephosphorylate a protein and clear the "memory" of a previous signal, genetic engineers can design synthetic biological circuits that manage information flow without triggering total metabolic exhaustion in the host organism 39.
Chromatin Fatigue and the Cost of DNA Repair
The physical consequences of information erasure extend deeply into the genome. The DNA molecule is a vast information storage medium folded into an immensely complex three-dimensional chromatin architecture, allowing meters of genetic sequence to be packed into a microscopic nucleus. When DNA suffers a double-strand break due to environmental radiation or metabolic stress, the cell initiates powerful repair mechanisms to restore the precise genetic sequence.
However, recent research leveraging high-content microscopy and chromosome conformation capture reveals a hidden thermodynamic and informational cost to this repair process. While the linear DNA sequence may be successfully restored, the surrounding three-dimensional chromatin structure often fails to refold exactly to its intricate pre-damage topological state. This phenomenon, newly termed "chromatin fatigue," results in persistent changes to gene expression and cell physiology that are permanently passed on to daughter cells 40. The failure of the cellular machinery to completely "erase" the damage event and perfectly restore the original physical state highlights the sheer thermodynamic difficulty of perfectly reversing structural biological information at the macroscopic level.
Restriction Costs in Metabolic Networks
Furthermore, Landauer's principle and broader non-equilibrium thermodynamics shed critical light on the structural architecture of biological metabolic pathways. A given cell possesses thousands of chemically viable pathways to convert a specific nutrient into a metabolic product. Why does nature consistently favor certain specific cycles, such as the Calvin cycle in photosynthesis?
Recent thermodynamic models have introduced the concept of a "restriction cost." This metric quantifies the immense entropic and energetic cost the cell must continuously pay to suppress alternative, physically possible chemical pathways while keeping only the desired pathway active. From a purely classical mechanics perspective, compartmentalizing reactions should incur no thermodynamic cost. However, from an information-theoretic viewpoint, preventing chemical flows from dispersing into all physically possible alternatives requires continuous constraint. The metabolic pathways universally observed in modern biology represent the optimal configurations where the energetic cost of maintaining functional boundaries and erasing unwanted chemical states is absolutely minimized 50.
Evolutionary Dynamics and Functional Information
Despite its immense utility in calculating channel capacities and energetic trade-offs, classical Shannon information theory is frequently criticized by evolutionary biologists for a critical omission: it deals exclusively with the statistical properties and probability distributions of a signal, completely ignoring the signal's meaning, utility, or value to the host organism 1341. In Shannon's strict mathematical framework, a string of randomized, non-coding DNA possesses a higher entropy, and thus contains more theoretical information, than a highly conserved, repetitive gene that codes for a vital survival protein. This dissonance highlights the limitations of using raw entropy to assess biological evolution.
The Metric of Functional Information
To address this profound conceptual gap, evolutionary biologists and origin-of-life researchers have proposed alternative metrics to strictly quantify biological complexity. Researchers Jack Szostak and Robert Hazen advanced the concept of "Functional Information," which argues that complexity in a biological context only has measurable meaning relative to a specific physiological function 1015.
If a biological system has a combinatorially vast number of possible configurations, such as all possible sequence permutations of a 300-amino-acid protein, functional information $I(E_x)$ measures the probability that an arbitrary configuration will achieve a specific degree of function $E_x$, such as the ability to fold and catalyze a specific reaction. It is defined mathematically as:
$$I(E_x) = -\log_2 [F(E_x)]$$
where $F(E_x)$ is the fraction of all possible configurations that possess a degree of function greater than or equal to the designated threshold $E_x$ 101542.
Unlike Shannon entropy, which naturally maximizes in a state of pure structural randomness, functional information relies entirely on the rarity of a functional state. In advanced computational models of artificial life, such as the Avida platform, and in physical studies of RNA aptamers, the vast majority of random mutations result in completely non-functional sequences. Only rare "islands" of high function exist within the massive sequence space. Natural selection acts to isolate and preserve these exceedingly rare configurations. Thus, functional information bridges the vital gap between raw data capacity and biological utility, providing a rigorously quantifiable metric for tracking the emergence of genuine complexity in evolutionary biology 1043.
Critiques of the Free Energy Principle
In recent years, highly ambitious attempts to merge thermodynamics, Bayesian inference, and information theory into grand unifying biological theories have sparked significant academic debate. The most prominent of these frameworks is the Free Energy Principle, championed extensively by neuroscientist Karl Friston. The principle argues that all sentient and biological systems, from single cells to human brains, operate fundamentally to minimize "surprise" by constantly updating internal predictive models of the environment to conform to expected sensory inputs 444556.
While highly influential in theoretical cognitive science, the Free Energy Principle faces severe criticism from both evolutionary biologists and thermodynamic physicists. Critics argue that the framework suffers from a fundamental conceptual flaw: it categorically conflates information-theoretic "free energy", which is merely a measure of statistical surprise or Kullback-Leibler divergence, with actual thermodynamic free energy, which is the physical capacity of a system to perform thermodynamic work 455657.
Furthermore, critics point out a hidden teleological category error inherent in the math. The framework imposes advanced cognitive concepts like "goals," "preferences," and "predictions" onto low-level mechanistic processes, such as primitive bacterial responses or single-cell adaptations, where such anthropomorphic concepts lack any genuine physiological relevance 4445. Philosophers of science and neurobiologists have warned that while predictive processing is a highly valid, testable hypothesis for specific higher-order brain functions, elevating it to the status of a universal physical principle via impenetrable mathematics borders on obfuscation rather than scientific clarification 56. For information theory to remain a rigorous tool in biology, the distinct boundaries between mathematical abstractions of probability and measurable thermodynamic reality must be strictly maintained.
Synthesis of Biological Information Theory
The integration of information theory into the biological sciences has fundamentally transformed the understanding of how life operates at the physical level. By treating the organism not merely as a complex chemical engine, but as an advanced information-processing entity subject to the strict laws of physics, researchers have uncovered the absolute thermodynamic and informational constraints guiding evolution.
From the macro-scale, where the nervous system actively sacrifices kinematic efficiency to maximize sensory data acquisition, down to the deepest molecular scales, where the Landauer bound extracts a relentless thermodynamic tax for the erasure of chemical memory, biological life is rigidly governed by the physical cost of information. Nervous systems and cellular signaling networks uniformly exhibit sophisticated structural architectures - such as inhibitory recurrence in the cortex and dual-channel thresholding in the immune system - that act to maximize mutual information while avoiding catastrophic metabolic exhaustion. While classical Shannon theory continues to unravel the absolute physical limits of neural and cellular transmission capacity, emerging frameworks like functional information are finally allowing researchers to mathematically quantify the raw physiological utility and genuine evolutionary meaning of the biological code.