Genomic surveillance and diagnostics in rural hantavirus hotspots 2026
Epidemiological Landscape of Orthohantaviruses
Hantavirus infections constitute a persistent, severe, and globally distributed zoonotic threat, resulting in tens of thousands to over one hundred thousand estimated human infections annually 1. The causative agents are enveloped, negative-sense, single-stranded RNA viruses belonging to the genus Orthohantavirus within the family Hantaviridae 2345. The clinical manifestations of these infections are strictly delineated by the viral species and their geographically specific rodent reservoirs. In the Eastern Hemisphere, specifically across Europe and Asia, viral species such as Hantaan, Seoul, Puumala, and Dobrava-Belgrade are endemic 267. These Old World strains precipitate hemorrhagic fever with renal syndrome (HFRS), an illness characterized by renal failure, vascular leakage, and hemorrhagic manifestations, which carries a case fatality rate (CFR) varying from less than 1% to 15% depending on the specific viral strain 25678.
Conversely, the Western Hemisphere is dominated by New World hantaviruses, including the Sin Nombre virus in North America and the Andes virus in South America 691011. These pathogens precipitate hantavirus pulmonary syndrome (HPS), occasionally referred to as hantavirus cardiopulmonary syndrome (HCPS). HPS is marked by a rapid onset of non-cardiogenic pulmonary edema, acute respiratory distress, and cardiogenic shock 291213. While the absolute number of HPS cases is lower than HFRS cases globally, HPS is substantially more lethal, with historical mortality rates averaging between 35% and 40% 81214. The stark contrast in clinical outcomes between Old World and New World strains necessitates highly specialized diagnostic and surveillance frameworks tailored to the localized viral ecology of the affected region.
The 2025 and 2026 South American Outbreak Surge
The epidemiological profile of New World hantaviruses shifted dramatically between late 2025 and early 2026, driven by complex environmental and ecological factors. Climatic variations, including anomalous precipitation patterns and localized warming, catalyzed the proliferation of rodent populations and expanded their geographical overlap with human agricultural and residential zones 1415. Data compiled by the Pan American Health Organization (PAHO) at the conclusion of 2025 recorded a total of 229 confirmed HPS cases and 59 associated deaths across eight countries in the Americas, yielding a regional CFR of 25.7% 1618.
The burden of disease was heavily concentrated in the Southern Cone. Argentina reported the highest volume of infections, documenting 66 cases and 21 deaths, equating to a 32% CFR 18. Brazil recorded 20 cases with 11 deaths, resulting in a disproportionately high CFR of 55%, while Bolivia and Chile reported 48 and 35 cases, respectively, with CFRs hovering around 20% to 23% 1618. Beyond the raw incidence rates, the geographical distribution of outbreaks within these countries exhibited alarming alterations. In Argentina, viral transmission shifted from historically endemic southern regions in Patagonia toward the densely populated Central Region. By early 2026, Buenos Aires Province and the Greater La Plata area accounted for 57% of national cases, doubling the regional incidence observed during the previous epidemiological season 17. The national CFR in Argentina temporarily surged to approximately 34% during the 2025 - 2026 summer peak, underscoring the severity of the expanding ecological risk zones and the urgent need for decentralized genomic surveillance 17.
The MV Hondius Transmission Cluster
The latent vulnerabilities in global hantavirus preparedness were abruptly exposed in May 2026 following a concentrated outbreak aboard the expedition cruise ship MV Hondius 49181920. The vessel, carrying 147 passengers and crew from 23 distinct nations, departed from Ushuaia, Argentina, on April 1, 2026, navigating a transatlantic route via remote South Atlantic outposts including Antarctica, South Georgia Island, and Saint Helena 3921. The index case fell ill on April 6 with non-specific symptoms including fever, headache, and gastrointestinal distress, and subsequently died onboard on April 11 before definitive diagnostic testing could be administered 22.
The pathogen remained undetected as the vessel continued its voyage. The spouse of the index case disembarked at Saint Helena on April 24, traveled via commercial aircraft to South Africa, and collapsed fatally at the Johannesburg airport, precipitating an international public health crisis 2022. The subsequent epidemiological investigation and international contact tracing operation identified a total of eight cases linked to the vessel, comprising six laboratory-confirmed infections and two probable cases, culminating in three deaths and an overall cluster CFR of 38% 222326.
Laboratory analysis confirmed the etiologic agent as the Andes virus (ANDV) 2923. ANDV occupies a unique position within the Orthohantavirus genus as the only strain proven to sustain human-to-human transmission 292425. Such transmission is generally confined to settings involving close, prolonged contact with an infected individual during the prodromal phase or the early acute stages of the illness 21624. Genomic sequencing of clinical isolates from the cruise ship patients - conducted by the National Institute for Communicable Diseases (NICD) in South Africa in collaboration with European reference laboratories - revealed exceptionally low viral diversity among the cluster 22. This genetic uniformity strongly supports a scenario of initial zoonotic exposure to an infected rodent followed by sequential human-to-human transmission within the confined environment of the ship's cabins 22.
Pathophysiology and Clinical Progression
The effective deployment of diagnostics is entirely dependent upon an accurate understanding of the temporal dynamics of the viral infection. The pathogenesis of HPS involves a profound vascular assault 410. The virus specifically targets the vascular endothelium - the cellular lining of blood vessels - and macrophages, triggering a severe, localized inflammatory response 410. This immune hyperactivation produces a cytokine storm that drastically increases microvascular permeability, forcing blood plasma to leak from capillaries into the pulmonary alveoli 410.
The Prodromal Window
Clinical progression follows a predictable temporal sequence, preceded by a highly variable incubation period ranging from 9 to 33 days, and occasionally extending beyond six weeks 21223. The illness initiates with a prodromal phase lasting 3 to 5 days, characterized entirely by non-specific, flu-like symptoms 61213. Patients typically present with fever, chills, severe myalgia predominantly affecting large muscle groups such as the thighs and lower back, headaches, and significant gastrointestinal symptoms including nausea, vomiting, and abdominal pain 691213.
During this initial 72-hour window, clinical diagnosis is notoriously difficult 921. Viral shedding in bodily secretions is minimal, and the clinical presentation is frequently indistinguishable from endemic arboviral infections such as dengue, chikungunya, or Zika, as well as bacterial infections like leptospirosis 49212627. Following the prodrome, the disease abruptly transitions into the cardiopulmonary phase, wherein patients experience a sudden onset of coughing, severe dyspnea, and hypoxia due to bilateral pulmonary infiltrates 9121321. In the absence of targeted antiviral therapeutics, patient survival is entirely dependent upon the rapid initiation of aggressive supportive care, such as extracorporeal membrane oxygenation (ECMO) or mechanical ventilation 9132128. The rapid deterioration characteristic of HPS means that any delay in diagnosis during the prodromal phase directly correlates with elevated mortality 41321.
Hematological Indicators
Because direct virological detection is challenging in the earliest stages of the disease, hematological markers serve as critical diagnostic proxies. Progressive thrombocytopenia - a rapid and severe decline in platelet counts - is an early clinical hallmark of hantavirus infection 4612. The drop in platelets is not merely a consequence of cellular destruction; rather, platelets are actively sequestered within the pulmonary microvasculature as part of the endothelial inflammatory response 4.
By 2026, modern pathology laboratories in endemic regions have increasingly adopted systems of participatory diagnostics to monitor these markers. Cloud-connected health informatics platforms are utilized to analyze anonymized complete blood count (CBC) data across regional populations 4. By establishing automated algorithms to detect localized clusters of unexplained thrombocytopenia, public health networks can identify potential hantavirus or severe arboviral outbreaks days before specialized molecular serology confirms the specific pathogen 4.
Deployable Point-of-Care Diagnostic Technologies
Historically, the diagnostic framework for hantavirus has relied heavily on centralized reference laboratories 1929. Serum samples collected from rural clinics were transported across long distances to facilities capable of executing complex molecular assays. However, the geographic expansion of hantavirus risk zones and the stringent time constraints of the cardiopulmonary phase have necessitated the decentralization of testing infrastructure and the rapid deployment of point-of-care (POC) diagnostics.
Serological Methodologies
The foundational method for diagnosing acute hantavirus infection remains serology, specifically the detection of hantavirus-specific immunoglobulin M (IgM) and immunoglobulin G (IgG) antibodies 68121930. In centralized facilities, enzyme-linked immunosorbent assays (ELISA) and immunofluorescence assays are heavily utilized due to their high sensitivity and cost-effectiveness at scale, often costing less than US$1.00 per test run 31. However, traditional ELISAs require stable electrical infrastructure, specialized spectrophotometers, and stringent cold chains to preserve reagents, rendering them unsuitable for remote deployment.
To bridge this gap, rapid immunochromatographic lateral flow assays have been engineered for field use 832. These tests operate on the same mechanical principles as rapid antigen tests, providing a visual readout of IgM antibodies within 15 to 30 minutes 83233. While highly portable and requiring minimal training, rapid serology suffers from reduced sensitivity - ranging from 60% to 85% - particularly during the early prodromal phase before the host immune system has mounted a robust, detectable antibody response 3233. A negative rapid test in the first 72 hours of fever cannot definitively rule out hantavirus infection.
Isothermal Amplification and Portable PCR
To achieve definitive virological confirmation in the field, molecular diagnostics have been aggressively miniaturized. Reverse transcription-polymerase chain reaction (RT-PCR) remains the gold standard for detecting viral RNA in blood or serum 102930. Platforms such as the FilmArray or the POCKIT system are increasingly deployed in rural healthcare settings 293234. These "lab-in-a-pouch" and cartridge-based systems integrate automated nucleic acid extraction, amplification, and data analysis into ruggedized, portable units 34. They are capable of executing multiplexed panels that test simultaneously for a variety of biothreats and endemic zoonoses within a single hour 3334.
Furthermore, significant advancements have been made in colorimetric reverse transcription loop-mediated isothermal amplification (RT-LAMP) 2932. Unlike traditional PCR, which requires rapid thermal cycling to amplify DNA, RT-LAMP utilizes specialized enzymes that operate at a constant temperature (typically around 65°C) 32. This eliminates the need for expensive and power-hungry thermocyclers; the reaction can be conducted using a simple heating block or water bath. Field-deployable RT-LAMP assays provide a visual color change in the presence of viral RNA within 30 to 60 minutes, achieving diagnostic sensitivities exceeding 90% while significantly reducing the hardware and financial burdens on rural clinics 2932.
Comparison of Rural Diagnostic Modalities
The selection of a diagnostic approach in a rural setting requires a careful calculation balancing sensitivity against logistical feasibility. The following table summarizes the comparative metrics of the primary modalities utilized in 2026.
| Diagnostic Modality | Analytical Target | Average Turnaround Time | Estimated Cost per Test (USD) | Sensitivity (Acute Phase) | Infrastructure and Hardware Requirements |
|---|---|---|---|---|---|
| Rapid Serology (Lateral Flow) | IgM / IgG antibodies | 15 - 30 minutes | $5 - $15 | 60% - 85% | Minimal; visual readout, room temperature storage possible 3233 |
| Traditional ELISA | IgM / IgG antibodies | 1 - 4 hours | < $5 (at scale) | 70% - 95% | Centralized laboratory, spectrophotometer, cold chain reagents 3133 |
| Portable RT-PCR (e.g., FilmArray) | Viral RNA | ~1 hour | $25 - $75 | 85% - 99% | Portable device, reliable field AC power, proprietary test cartridges 293334 |
| Colorimetric RT-LAMP | Viral RNA | 30 - 60 minutes | $10 - $25 | > 90% | Constant heating block (65°C), standard pipettes, visual readout 2932 |
| Nanopore Metagenomics | Whole genome RNA/DNA | 3 - 8 hours | > $100 | 50% - 83% | Portable sequencer, computing device, nucleic acid extraction kit, stable power 35363738 |
Miniaturized Genomic Sequencing Platforms
Beyond acute patient diagnostics, genomic epidemiology is an indispensable tool for managing hantavirus outbreaks. Rapid sequencing is required to differentiate between circulating endemic strains, pinpoint the precise geographic origins of zoonotic spillover events, and detect localized viral mutations that may confer the ability for human-to-human transmission 103940. In the past, next-generation sequencing (NGS) was heavily centralized, requiring samples to be transported to well-equipped reference centers - a process that typically delayed actionable results by two to five days 41.
Oxford Nanopore Technologies
By 2026, the paradigm of genomic surveillance has been fundamentally decentralized through the proliferation of third-generation long-read sequencing technologies, most notably the platforms developed by Oxford Nanopore Technologies (ONT) 3945424344. Nanopore sequencing operates on a distinct physical principle compared to traditional short-read platforms; it identifies nucleotide sequences by measuring minute fluctuations in ionic current as a single DNA or RNA molecule is ratcheted through a nanoscale protein pore embedded in a synthetic membrane 424344.
The primary advantage of this technology is its portability. The MinION sequencer is a compact, USB-powered device capable of real-time data streaming 394546. For highly localized, cost-sensitive rural surveillance, epidemiologists frequently utilize the Flongle, an adapter for the MinION designed to run lower-throughput, single-use flow cells 3545. A Flongle flow cell can generate up to 2.6 Gb of sequence data, which is more than sufficient for characterizing small viral genomes at a consumable cost of approximately US$100 per run 3545.
When investigating a localized outbreak, field researchers utilize targeted amplicon-based sequencing protocols. By utilizing specific primer sets, researchers can rapidly amplify the tripartite RNA genome of the hantavirus (comprising the Small, Medium, and Large segments) directly from human clinical samples or captured rodent tissues 12353944. This targeted approach routinely achieves over 95% genome coverage with high sequence fidelity within three hours of sample preparation 35394447. When a human clinical isolate is sequenced in the field and phylogenetically matched to the genome of a virus extracted from a local rodent reservoir (e.g., Oligoryzomys longicaudatus for the Andes virus), health authorities can precisely map the environmental site of exposure and implement targeted rodent control measures 10223544.
Clinical Metagenomics and Sensitivity Constraints
While targeted sequencing is highly effective, infectious disease specialists also utilize clinical metagenomics (CMg) for broad, untargeted pathogen discovery. Metagenomics involves sequencing all nucleic acids present in a clinical sample without prior knowledge of the infectious agent, making it a powerful tool for investigating undifferentiated febrile illnesses 37464849. Techniques such as sequence-independent single primer amplification (SISPA) allow for the non-selective reverse transcription and amplification of all RNA in a sample 48.
However, deploying clinical metagenomics in rural clinics for acute diagnostics presents severe analytical challenges. Because clinical samples (such as blood or nasopharyngeal swabs) contain an overwhelming volume of host genetic material compared to viral RNA, sequencing depth is often consumed by analyzing human DNA rather than the pathogen 3654. Independent clinical validation studies conducted in 2026 revealed that the diagnostic sensitivity of untargeted nanopore metagenomics for respiratory and systemic RNA viruses ranged between 51% and 61% when compared to gold-standard multiplex PCR 3654. While sensitivity can be mathematically improved to 83% by restricting analysis to samples with high viral loads (e.g., PCR cycle threshold values below 35), the fundamental limit remains 3654. Furthermore, read crossover errors during the multiplexing of multiple patient samples on a single flow cell frequently lead to false-positive identifications 363754. Consequently, while metagenomics is unparalleled for discovering novel reassortments and establishing baseline viromes, targeted PCR or amplicon sequencing remains the definitive standard for acute patient triage in resource-limited settings 3754.
Field Hardware and Logistical Constraints
The theoretical capability to sequence viral genomes at the site of an outbreak does not negate the severe physical and infrastructural realities of rural deployment. Epidemiologists operating in the agricultural zones of Argentina, the isolated valleys of Chile, or the remote settlements of West Africa must engineer self-sustaining hardware ecosystems capable of functioning independently of the electrical grid.
Micro-Laboratories and Power Consumption
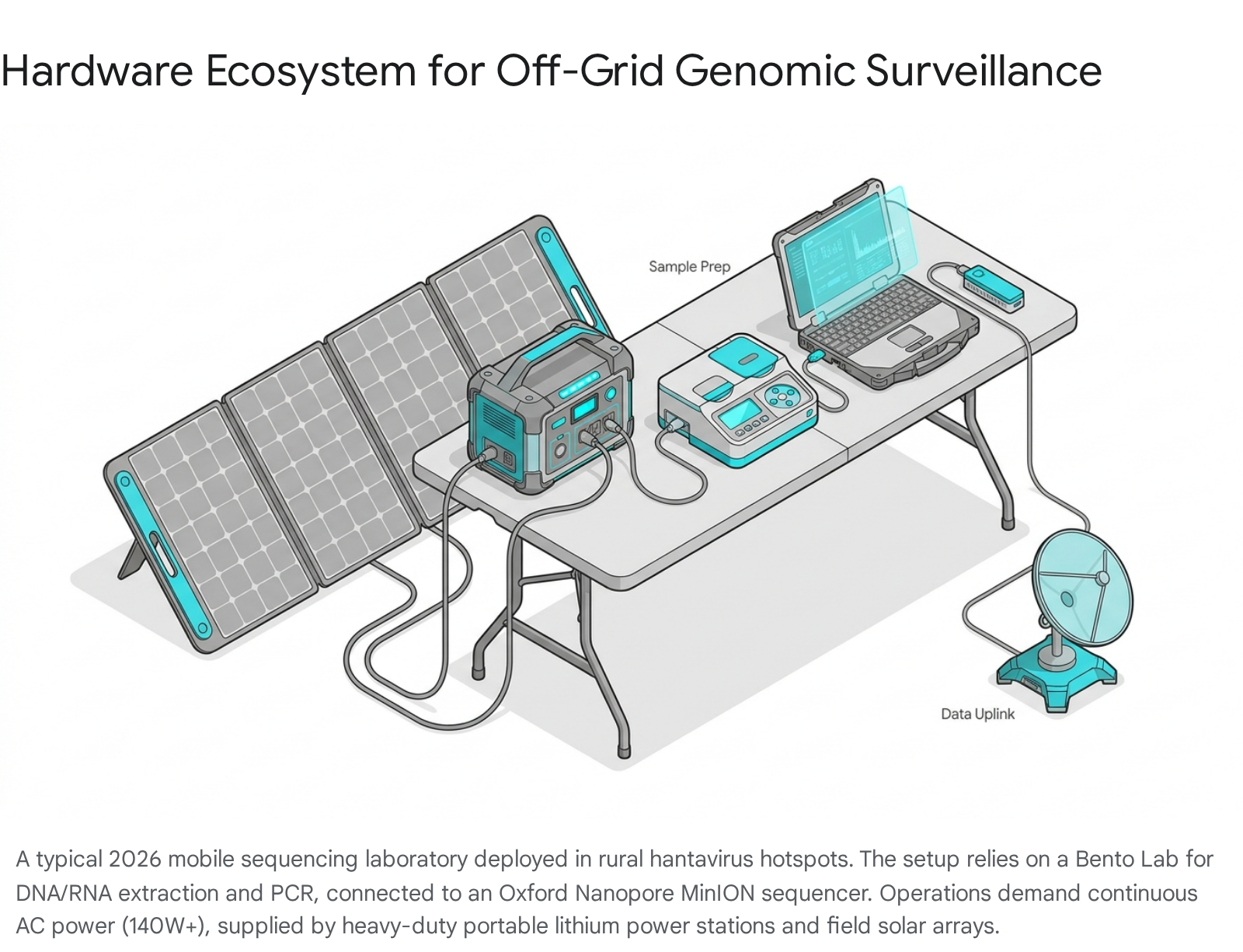
Pre-sequencing procedures, including nucleic acid extraction, purification, and library preparation, traditionally require bulky laboratory equipment 3845. To consolidate this footprint, field teams frequently employ the Bento Lab, a mobile, laptop-sized device that integrates a microcentrifuge, a PCR thermal cycler block, and an illuminated gel electrophoresis unit into a single ruggedized chassis 384555.
However, thermal cycling and centrifugation are highly energy-intensive processes. The Bento Lab draws a maximum power of 140 watts and requires a stable alternating current (AC) mains output to function correctly 56. To sustain continuous operations during an extended field investigation, researchers rely on high-capacity portable lithium power stations, ranging from 146 watt-hours (Wh) to over 400Wh 5650. These power stations are continually recharged throughout the day via 200-watt folding solar panel arrays 50. Once the sequencing library is prepared and loaded onto the MinION flow cell, the raw ionic current data must be basecalled into nucleotide sequences. Because local computing hardware is often insufficient for rapid neural-network-based basecalling, field vehicles (such as specialized Class B recreational vehicles) are outfitted with satellite internet terminals, such as Starlink, to transmit raw data to centralized cloud computing clusters for real-time phylogenetic alignment 38.
Reagent Stability and Cold Chain Mitigation
The logistical fragility of cold chains presents a massive barrier to rural molecular diagnostics. Traditionally, polymerase enzymes and nucleotide buffers degrade rapidly when exposed to ambient tropical or summer temperatures, rendering field deployments highly vulnerable to equipment failure. To bypass these thermal limitations, diagnostic manufacturers and genomic suppliers have pivoted toward lyophilized (freeze-dried) chemistries 3450.
Protocols such as Oxford Nanopore's Field Sequencing Kit (SQK-LRK001) utilize lyophilized reagents that do not require continuous refrigeration, allowing them to be transported in standard field kits and rehydrated with specialized buffers immediately prior to use 4550. Furthermore, organizations like PATH have established collaborative projects to generate, characterize, and bank standardized antibody reagents with extended real-time stability profiles extending through 2026 51. Ensuring that both extraction chemistries and standardized testing antibodies remain stable at ambient temperatures is crucial for maintaining the analytical integrity of diagnostics executed far outside of a controlled laboratory environment.
Institutional Funding and One Health Integration
The rapid advancement of miniaturized diagnostic technologies is persistently juxtaposed against the extreme volatility of global health financing. The macroeconomic landscape in 2026 has witnessed a severe contraction in official development assistance (ODA) directed toward health initiatives in low- and middle-income countries, forcing reliance on targeted philanthropic coalitions and strategic global mandates 52.
Vaccine Development Stagnation
The urgency of funding robust diagnostic frameworks is magnified by the systemic failure to deliver a universal prophylactic vaccine against the Orthohantavirus genus. The development of a hantavirus vaccine has stalled repeatedly due to the immense genetic diversity among viral strains and the lack of a reliable immunological correlate of protection 53. The only commercial vaccine currently in use is Hantavax, an inactivated whole-virus formulation licensed in the Republic of Korea in 1990 53545556. While Hantavax provides variable efficacy against the Old World Hantaan and Seoul viruses (responsible for HFRS), it confers zero cross-protection against the New World Andes and Sin Nombre viruses driving the outbreaks in the Americas 28535557.
Alternative vaccine platforms, including a DNA vaccine targeting the Andes virus developed by the U.S. Army Medical Research Institute of Infectious Diseases (USAMRIID) and mRNA candidates heavily supported by Korean academic institutions and Moderna, remain stalled in early Phase I or preclinical stages 535556. Without an approved vaccine for HPS, public health strategies are entirely reliant on early diagnostics, vector surveillance, and aggressive supportive care to mitigate mortality 2853.
CEPI and Geopolitical Pressures
The Coalition for Epidemic Preparedness Innovations (CEPI) has integrated hantavirus countermeasures into its overarching "100 Days Mission," a strategic framework designed to compress the timeline for developing diagnostics, therapeutics, and vaccines against novel pathogens to just over three months 51525859. Through continuous calls for proposals and targeted "Tech Talks," CEPI actively funds regional manufacturing hubs and technology transfer programs, prioritizing researchers within the Global South to ensure that countermeasures can be produced locally during an outbreak 6061.
However, international collaborative networks remain highly susceptible to geopolitical disruption. In a significant setback to zoonotic research, the U.S. administration abruptly terminated funding in 2025 for the National Institutes of Health's Centers for Research in Emerging Infectious Diseases (CREID) network 18. This sudden shutdown dismantled ten global centers, including the West African Center for Emerging Infectious Diseases (WAC-EID), and directly halted pilot projects specifically designed to study the mechanical pathways of hantavirus transmission from rodents to humans 18. The loss of this funding eliminated critical field data streams exactly a year before the MV Hondius outbreak underscored the danger of human-to-human hantavirus transmission 1820.
The Imperative of the One Health Framework
To counter the fragmentation of global funding, regional bodies such as the Pan American Health Organization (PAHO) and the World Health Organization (WHO) have heavily mandated a "One Health" approach to epidemiological surveillance 1662636465. The One Health paradigm acknowledges the inseparable link between human epidemiology, veterinary pathology, and environmental ecology 27626566.
For a zoonotic pathogen like hantavirus, human incidence rates are a lagging indicator. The true driver of an outbreak is the environmental fluctuation that causes an explosion in the local rodent population 14. By embedding genomic surveillance teams within rural ministries of health, governments in Argentina, Chile, and Bolivia can proactively monitor the viral load and genetic drift of local rodents long before agricultural workers or rural residents are exposed 16636467. Deploying portable sequencing hardware directly to the fields allows for the immediate identification of high-risk viral reassortments, enabling localized vector control, targeted community education, and the pre-positioning of critical care equipment (such as ECMO machines) at regional hospitals before the first human patient transitions into the cardiopulmonary phase.
In conclusion, the mitigation of hantavirus in 2026 demands a highly decentralized, technologically advanced, and ecologically aware public health strategy. The extreme lethality of New World strains, coupled with the proven transmission capabilities of the Andes virus, dictates that waiting for centralized laboratory confirmation is no longer viable. The deployment of field-ready point-of-care diagnostics and miniaturized genomic sequencers, sustained by off-grid power ecosystems and integrated into a holistic One Health framework, represents the only effective defense against the escalating threat of hantavirus spillover.