Genetics of centenarians and extreme human longevity
The biological study of exceptional human longevity - defined demographically by individuals surviving past 100 years (centenarians), 105 years (semi-supercentenarians), and 110 years (supercentenarians) - provides an invaluable natural model for understanding the molecular mechanisms of healthy aging, physiological resilience, and disease avoidance. Historically, aging research focused predominantly on the gradual accumulation of stochastic cellular damage and the environmental factors that accelerate this decline. However, recent advances in whole-genome sequencing (WGS), multi-omic profiling, ATAC-seq, and the construction of polygenic risk scores have demonstrated that surviving to the extreme limits of the human lifespan requires a highly specific, inherited genetic architecture.
Extreme longevity is not merely characterized by the absence of disease-causing alleles. Rather, it relies heavily on the presence of rare protective genetic variants, highly efficient DNA double-strand break repair mechanisms, enhanced mitochondrial crosstalk, and a significantly reduced burden of rare loss-of-function (LOF) mutations across highly conserved biological pathways. This report synthesizes current molecular and epidemiological research to delineate the genetic variants, structural chromosomal architectures, and biological pathways that enable the oldest old to evade the hallmark diseases of aging.
Heritability and the Demographic Architecture of Exceptional Lifespan
Human lifespan is a complex, multifactorial trait influenced by an intricate interplay of environmental exposures, lifestyle choices, stochastic biological events, and genetic predispositions. Before the age of 80, environmental factors, socio-economic conditions, and lifestyle habits account for the vast majority of lifespan variance, with genetics explaining only an estimated 10% to 25% of the overall survival probability in the general population 12. However, the genetic contribution to human survival undergoes a profound and measurable shift as individuals cross the centenarian threshold.
Longitudinal studies tracking the familial aggregation of longevity, such as the New England Centenarian Study (NECS) and the Long Life Family Study (LLFS), indicate that siblings of centenarians have between 8 and 17 times greater chances of living past 100 compared to individuals from the exact same birth year cohort who lack a familial history of extreme longevity 234. This survival advantage scales exponentially at the extreme ends of human life. For the siblings of supercentenarians (individuals aged 110 and older), relative survival probabilities (RSPs) demonstrate a massive demographic advantage that becomes most pronounced at the oldest ages 4. By age 90, sisters of supercentenarians possess a 2.9 times greater probability of survival, and brothers a 4.3 times greater probability, compared to their respective birth cohorts 45. Furthermore, mothers of supercentenarians demonstrate a 5.8 times greater probability of surviving from age 50 to age 90 45.
Current demographic modeling indicates that while reaching age 90 is approximately 30% genetic and 70% environmental, reaching age 110 is likely the inverse - up to 70% genetic and 30% environmental 6.
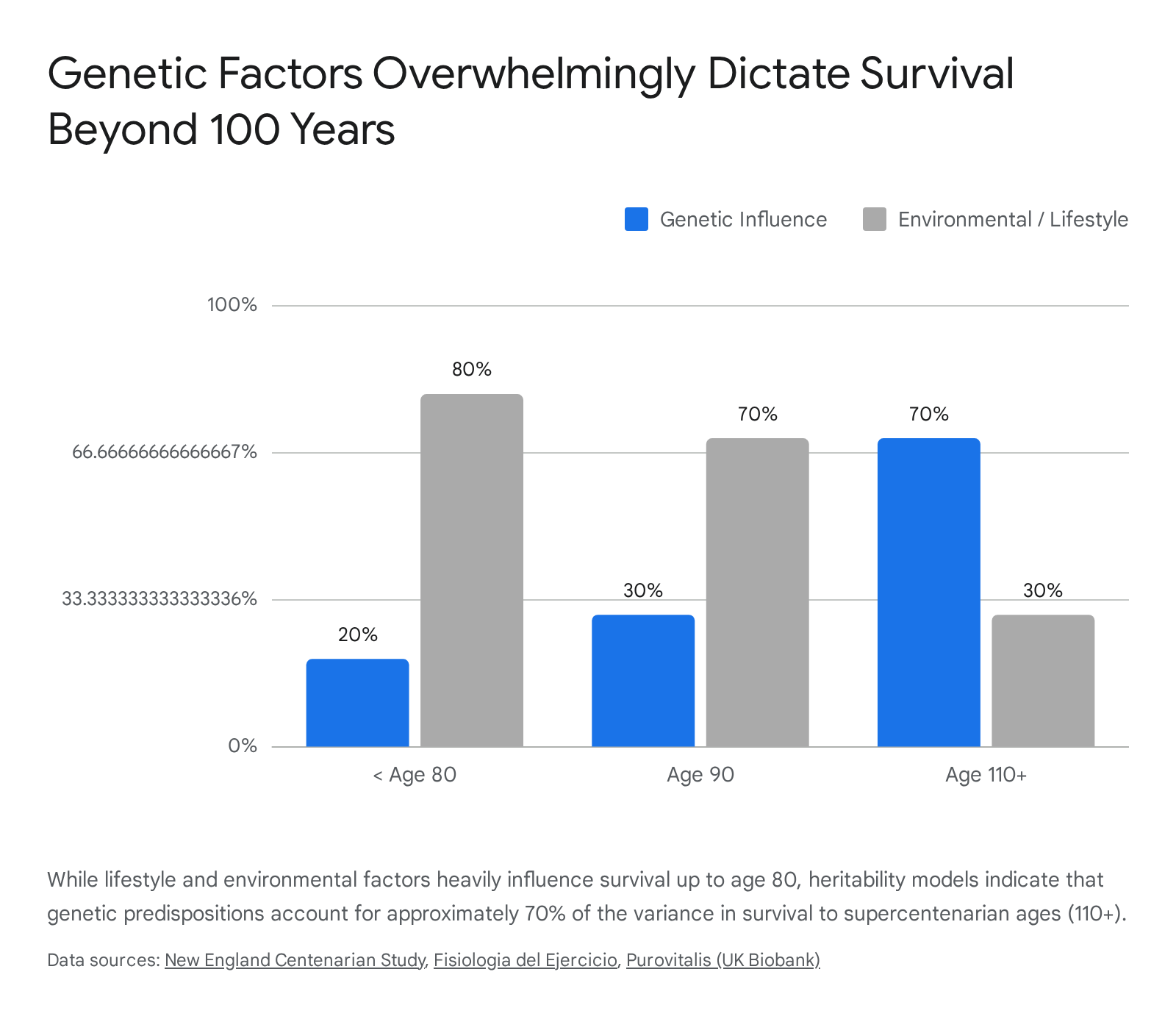
This dramatic transition suggests that pushing past the standard limits of average human lifespan relies on a biological framework largely determined by inherited genetic resilience, rather than cumulative healthy behaviors alone.
Polygenic Risk Scores, Protective Scores, and Intrinsic Capacity
Because the genetic foundation of extreme longevity relies on thousands of interacting alleles, each with relatively small individual effect sizes, researchers utilize Polygenic Risk Scores (PRS) to quantify the collective physiological impact of these variants. A PRS aggregates the weighted effect sizes of independent single nucleotide polymorphisms (SNPs) discovered in large-scale genome-wide association studies (GWAS) 79.
The Alzheimer's Disease Polygenic Protective Score (AD PPS)
A hallmark characteristic of centenarians is their profound resilience to age-related neurodegeneration, specifically Alzheimer's disease (AD). Recent genome-wide association studies have transitioned from simply measuring disease risk to establishing a Polygenic Protective Score specific to AD (AD PPS) 989. Constructed specifically without relying on the highly influential APOE variants in order to isolate non-APOE genetic factors, the AD PPS utilizes loci identified from broad European and multi-ancestry populations 8.
Comparisons across international cohorts - including the NECS, the LLFS, and the 100-plus Study - demonstrate that cognitively healthy centenarians possess a significantly higher AD PPS than population-matched controls without a history of familial longevity 789. Notably, this polygenic genetic protection scales directly with age at the extreme ends of the demographic spectrum. The mean AD PPS exhibits a clear, statistically significant increasing trend from nonagenarians (ages 90-99) to centenarians (100-104), semi-supercentenarians (105-109), and supercentenarians (110+) 89.
The estimated effect size of this protection is an average increase of approximately one additional protective allele for every five years of gained lifetime past the age of 100 89. Consequently, higher AD PPS scores strongly correlate with delayed cognitive decline over time (measured via Telephone Interview for Cognitive Status and logical memory recall tests) and a significantly lower hazard ratio for all-cause mortality 89. This firmly indicates that possessing an accumulation of protective neuro-genetic variants contributes directly to the ability to reach extreme old age.
GWAS of Intrinsic Capacity and Metabolic Resilience
The World Health Organization introduced the concept of "intrinsic capacity" (IC) in 2015 to define the composite of an individual's physical and mental capacities essential for maintaining functional ability and well-being during aging 1011. While the environmental impacts on IC are well documented, a comprehensive 2024-2025 multi-cohort GWAS utilizing data from the UK Biobank (N=44,631) and the Canadian Longitudinal Study on Aging (CLSA; N=13,085) calculated the SNP-based heritability of intrinsic capacity at 19.5% in the CLSA and 25.2% in the UK Biobank 11121314.
The analysis identified 38 independent SNPs across 10 novel genomic loci, which mapped to 197 specific candidate genes 1214. Post-GWAS transcriptomic analyses revealed that these IC-associated genes are highly expressed in critical metabolic and structural tissues, including muscle, heart, adipose, and tibial nerve tissues 1214. The mapped genes govern core cellular processes including cellular proliferation, immune system function, systemic metabolism, and neurodegeneration 1114. Of the 52 distinct traits tested in genetic correlation analyses, 23 showed significant genetic correlations with intrinsic capacity, proving that physical robustness and cognitive healthspan have shared underlying polygenic architectures 1114.
Similarly, an Exceptional Longevity Polygenic Risk Score (ELPRS) has been found to associate inversely with Metabolic Syndrome (MetS). Analyses performed on middle-aged to older adults in the UK Biobank demonstrated that individuals with a high polygenic score for extreme longevity had a significantly lower prevalence of MetS (OR = 0.94) and its individual components 15. This suggests that a healthy metabolic profile, specifically the avoidance of insulin resistance and dyslipidemia, is genetically hardwired into long-lived lineages, operating as a primary mechanism for disease avoidance.
| Polygenic Assessment | Primary Phenotypic Associations | Key Biological Mechanisms & Tissues | Relevance to Extreme Longevity |
|---|---|---|---|
| AD Polygenic Protective Score (AD PPS) | Reduced AD incidence, slower cognitive decline, lower all-cause mortality. | Excludes APOE. Aggregates protective alleles guarding neural integrity. | Score increases incrementally from age 90 to 110+, demonstrating scaling protection. 89 |
| Intrinsic Capacity GWAS Score | Composite physical and mental functional preservation. | 197 genes highly expressed in muscle, heart, adipose, and tibial nerve. | Establishes a 19.5% - 25.2% heritability for maintaining whole-body functionality. 1214 |
| Exceptional Longevity PRS (ELPRS) | Lower prevalence of Metabolic Syndrome (MetS). | Systemic metabolic regulation, glucose homeostasis, lipid processing. | High ELPRS inversely correlates with metabolic dysfunction, postponing cardiometabolic mortality. 15 |
The Apolipoprotein E (APOE) Locus: Isoforms, Ancestry, and Epistasis
Despite the discovery of numerous longevity-associated loci with small effect sizes, the APOE gene on chromosome 19 remains the single most robust and frequently replicated locus associated with extreme human longevity across diverse global populations 1617. The gene primarily encodes a protein essential for the transport of cholesterol and other lipids between the periphery and the central nervous system, directly impacting vascular integrity, cardiovascular disease risk, and amyloid-beta clearance in the brain 18.
Common Isoforms: ε2, ε3, and ε4
The three primary APOE alleles in human populations - ε2, ε3, and ε4 - are defined by varying combinations of genotypes at two single nucleotide polymorphisms: rs7412 and rs429358 17. The ε3 allele is the most prevalent in the general population, functioning as the standard biological baseline. Conversely, the ancestral ε4 allele is a highly established risk factor for both cardiovascular disease and late-onset Alzheimer's disease 171819.
In demographic comparisons between the general population and centenarian cohorts, there is a pronounced, statistically significant depletion of the ε4 allele and a notable enrichment of the protective ε2 allele. Data from a meta-analysis involving 28,297 subjects across seven studies of healthy aging revealed that the ε4 allele frequency drops significantly from approximately 11.2% in younger control populations to roughly 5.2% in centenarians 1720. Conversely, the frequency of the ε2 allele nearly doubles, rising from 6.8% in controls to 12.8% in centenarians 20. The ε2 allele is associated with marked reductions in total and LDL cholesterol, contributing to a lower risk of coronary artery disease and neurodegeneration 18.
The African Ancestry Protective Haplotype (rs10423769)
While the APOE ε4 allele is globally recognized as a severe risk factor for neurodegeneration, modern population genetics have uncovered profound ancestry-specific structural variations that fundamentally alter its penetrance. Specifically, researchers utilizing the Alzheimer's Disease Sequencing Project (ADSP) database have identified a variant - the A allele of rs10423769 - that virtually neutralizes the pathological risk of APOE ε4 in populations of African descent 2122232425.
The rs10423769 variant is situated approximately 2 megabases upstream of the APOE locus, residing within a highly complex region of segmental duplications (SD) on chromosome 19 2123. The protective A allele has a minor allele frequency of approximately 12% in African populations, but is exceedingly rare (roughly 0.03%) in populations of European ancestry 24. In individuals possessing the highly deleterious APOE ε4/ε4 homozygous genotype, the co-occurrence of the rs10423769_A allele reduces the risk of developing AD by up to 75%, effectively lowering their risk profile to approximate that of the neutral ε3/ε3 baseline 21222425.
Advanced PacBio HiFi long-read sequencing has recently resolved the highly complex genomic context of this protection. The protective variant operates within a unique 21-kilobase shared haplotype 2326. This haplotype harbors an expanded variable number tandem repeat (VNTR) containing multiple MEF2D transcription factor binding motifs, alongside distinct allele-specific DNA methylation patterns across 17 differentially methylated positions 2124.
The mechanism by which this variant protects the brain over a distance of two million base pairs relies on the three-dimensional spatial reorganization of the genome. Through enhanced Hi-C capture analysis (eHiCA), researchers established that the rs10423769 locus operates as a distal enhancer. The DNA strand physically folds, creating a long-range chromatin loop that allows the protective haplotype to interact directly with the APOE promoter and a neighboring cluster of pregnancy-specific beta-1 glycoprotein (PSG) genes and zinc finger (ZNF) genes 212627. This structural reorganization overrides the pathogenic transcription of the APOE ε4 allele, indicating that extreme longevity and disease resistance in certain lineages are driven by complex, 3D epigenetic remodeling rather than simple linear sequence variations.
The Burden of Rare Variants and Epistatic Buffering
While common variants (minor allele frequency >1%) identified via GWAS explain a localized portion of human longevity, extreme survival is heavily contingent on the complex interaction of rare coding variants. Recent exome-wide burden analyses of centenarian cohorts indicate that successful aging relies substantially on the depletion of rare, damaging loss-of-function (LOF) mutations 2829.
Depletion of Loss-of-Function Mutations
In whole-exome sequencing of Ashkenazi Jewish individuals from the Longevity Genes Project and the LonGenity study, researchers discovered that centenarians carried an 11% to 22% lower burden of LOF mutations compared to population controls 2829. This depletion phenomenon was highly specific to a subset of 35 longevity-associated genes.
Subsequent Mendelian randomization and multi-omic analyses validated several key genes from this depleted pool - including RGP1, PCNX2, and ANO9 - as having direct causal effects on multiple aging-related traits 2829. Age-related changes in blood gene expression, obtained via transcriptome-wide association studies (TWAS), confirmed that the transcription of genes such as PCNX2 and RGP1 alters significantly as the body ages 29. This implies that a protective genetic background in centenarians is characterized fundamentally by fewer functional disruptions in pathways central to long-term homeostasis, allowing their biology to operate efficiently over a century without system failure 2829.
Epistatic Buffering of Pathogenic Alleles
In addition to structural haplotypes and LOF depletion, extreme longevity is facilitated by epistasis - a phenomenon where rare protective variants actively suppress the adverse phenotypic effects of common pathogenic variants. Whole-exome sequencing of centenarians revealed that these individuals do not necessarily harbor fewer common pathogenic risk alleles for diseases like coronary artery disease compared to the general population 3031. Instead, they possess an abundance of rare coding variants that aggregate in the WNT, insulin/insulin-like growth factor 1 (IIS), and AMP-activating protein kinase (AMPK) signaling pathways 303132.
Specifically, rare coding variants within the WNT signaling pathway were shown to assert a profound pro-longevity effect that actively suppressed the expected mortality risks in individuals harboring the APOE ε4 allele 3032. This epistatic buffering indicates that extreme longevity often arises from a precise combination of biological pathways where robust, highly conserved metabolic signaling overrides localized genetic disease risk.
Genomic Stability: The SIRT6 Mutant and Cellular Senescence
Maintaining the integrity of the genome in the face of continuous oxidative stress and replicative damage is a primary requirement for reaching supercentenarian ages. Sirtuin 6 (SIRT6) is a class III histone deacetylase and mono-ADP ribosyl transferase (mADPr) enzyme that plays a critical role in maintaining genomic stability, regulating glucose metabolism, and delaying cellular senescence 333435. SIRT6 knockout models exhibit premature aging and catastrophic genomic instability, while its overexpression extends lifespan and suppresses tumorigenesis across species 3336.
In targeted sequencing of the SIRT6 locus in Ashkenazi Jewish centenarians, researchers identified a rare allele - termed centSIRT6 - that contains two perfectly linked missense substitutions: N308K and A313S 3337. This unique variant occurs at nearly double the frequency in centenarian populations compared to matched controls (1.0% vs. 0.55%) 3336.
The biophysical alterations caused by the N308K and A313S mutations fundamentally redirect the enzyme's functionality. Computational modeling using tools like SIFT indicates that the N308K substitution significantly impacts protein function by reducing the structural flexibility of the SIRT6 protein 34. This rigidity weakens the enzyme's interaction with acetyl-lysine substrates, resulting in diminished deacetylase activity. However, it concurrently strengthens the enzyme's interaction with ADP-ribose, drastically enhancing its mono-ADP ribosyl transferase (mADPr) activity 333437.
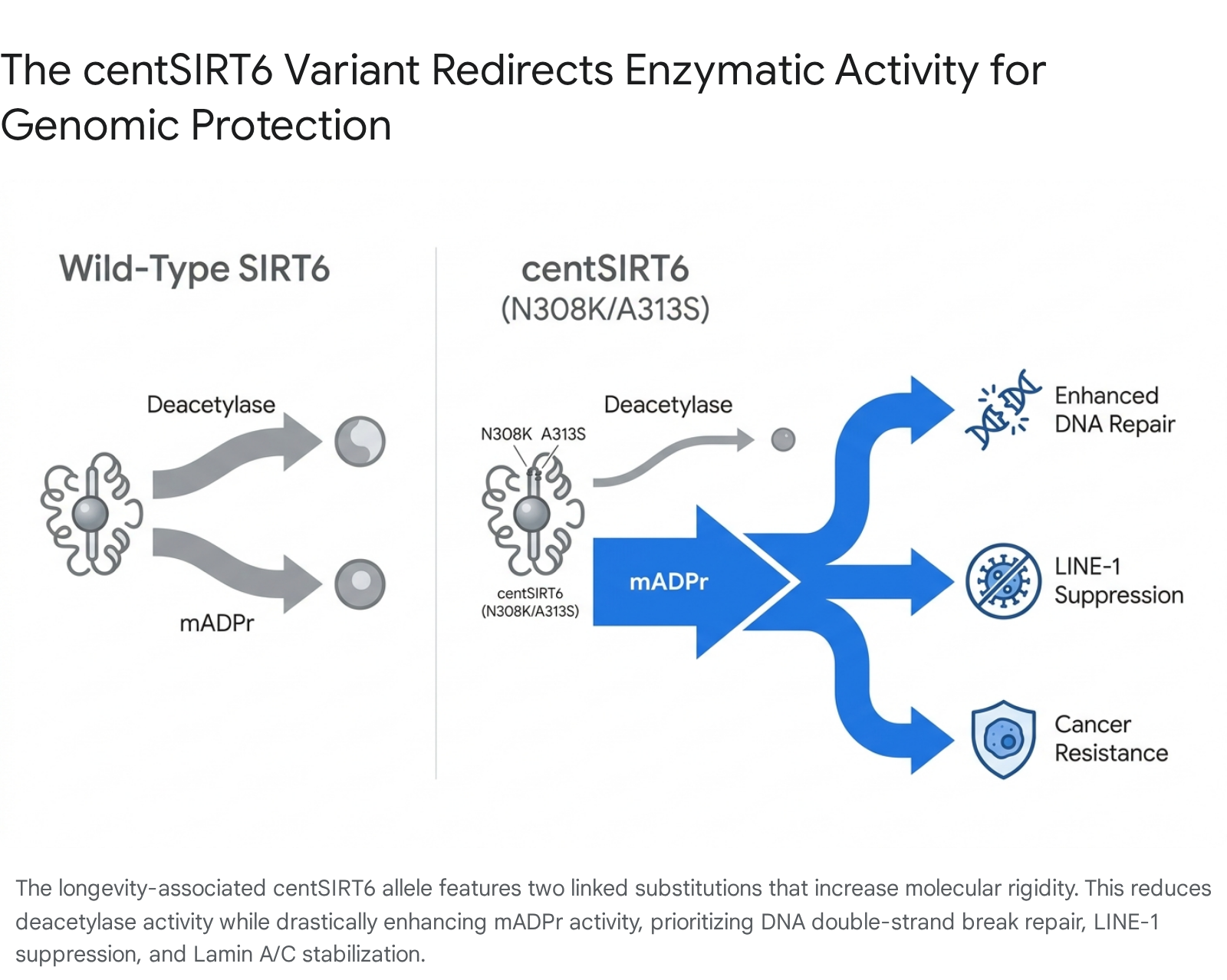
Phenotypically, the centSIRT6 allele provides multiple robust anti-aging benefits that contribute to extended healthspan: 1. Enhanced DNA Double-Strand Break Repair: The heightened mADPr activity rapidly recruits DNA repair machinery to sites of catastrophic double-strand breaks, maintaining chromosomal integrity 3337. 2. Suppression of Retrotransposons: The mutant enzyme is a significantly stronger suppressor of Long Interspersed Nuclear Element-1 (LINE1) retrotransposons, preventing the genomic "jumping" that destabilizes aging genomes 3334. 3. Cancer Resistance: In in-vitro studies utilizing adeno-associated viral (AAV2/8) vectors on hepatocellular carcinoma cell lines (HepG2 and Huh7), centSIRT6 exhibited a robust capacity to induce time-dependent apoptosis in malignant cells without harming healthy tissues, likely by altering glycolytic dependencies away from the Warburg effect 3336. 4. Lamin Interplay: The variant displays an intensified physical interaction with Lamin A/C (LMNA), facilitating enhanced ribosylation of the nuclear envelope. This preserves nuclear morphology and prevents progeroid-like structural collapse 3337.
Supercentenarian WGS: DNA Repair and Mitochondrial Crosstalk
As research parses the genomes of the absolute oldest living humans, extreme resilience against somatic mutations emerges as a defining biological trait. High-coverage (90X) WGS of Italian semi-supercentenarians (105+) and supercentenarians (110+) pinpointed a highly concentrated block of linkage disequilibrium on chromosome 7 383940. Five common variants (including rs7456688 and rs10279856) were profoundly enriched in the 105+ cohort, highlighting an interconnected genetic hub governing DNA repair and mitochondrial function 3839.
These variants act as pleiotropic expression quantitative trait loci (eQTLs), simultaneously modulating the expression of nearby genes STK17A, COA1, and BLVRA across multiple tissue types 414245.
- STK17A (Serine/Threonine Kinase 17a): The supercentenarian variants upregulate the expression of STK17A in critical tissues, including the heart. STK17A is heavily involved in managing the cellular response to DNA damage, regulating reactive oxygen species (ROS) metabolic processes, and triggering apoptosis in irreparably damaged cells 384143. Upregulation of this gene allows centenarian tissues to efficiently eliminate precancerous or senescent cells before dangerous clonal expansion occurs.
- COA1 (Cytochrome C Oxidase Assembly Factor 1): Conversely, these same eQTL variants downregulate the expression of COA1, a gene responsible for coordinating energy production between the cell nucleus and the mitochondria 414344. Because mitochondrial respiration is a primary source of endogenous oxidative stress, researchers hypothesize that the dampening of COA1 expression subtly lowers metabolic turnover, generating fewer ROS and thereby mitigating lifelong intracellular wear-and-tear 4144.
- BLVRA (Biliverdin Reductase A): The locus is also linked to an increased expression of BLVRA in whole blood, a gene important in the direct elimination of reactive oxygen species 414344.
| Supercentenarian Locus | Pleiotropic Effect | Primary Biological Mechanism | Longevity Benefit |
|---|---|---|---|
| STK17A | Upregulated expression | Coordinates cellular response to DNA damage and induces apoptosis in defective cells. 3843 | Prevents malignant transformation and clears damaged cells, maintaining tissue homeostasis. 4142 |
| COA1 | Downregulated expression | Moderates crosstalk between the cell nucleus and mitochondria during energy production. 4344 | Reduces the lifelong generation of reactive oxygen species (ROS) stemming from mitochondrial respiration. 4144 |
| BLVRA | Upregulated in blood | Acts directly in the clearance and elimination of reactive oxygen species. 4344 | Provides enhanced systemic resistance to oxidative stress. 41 |
This dual modulation - heightened genomic surveillance (STK17A) coupled with proactively reduced oxidative output (COA1) and enhanced systemic clearance (BLVRA) - creates a durable cellular environment highly resistant to the somatic mutations that drive cancer and tissue failure. Consequently, 105+ individuals exhibit a strikingly lower somatic mutational load and delayed clonal hematopoiesis of indeterminate potential (CHIP) compared to younger, healthy cohorts 13842.
Epigenomic Remodeling and the Avoidance of Cellular Senescence
The active avoidance of cellular senescence - a state of irreversible cell-cycle arrest accompanied by a toxic Senescence-Associated Secretory Phenotype (SASP) - is a fundamental hallmark of human longevity 454647. Genomic and transcriptomic data indicates that centenarians harbor specific profiles that suppress the accumulation of senescent cells, which largely operate through the p16-RB and p53-p21 pathways 464748.
In human genetic studies, exceptional longevity correlates with reduced expression and altered accessibility of the CDKN2A (p16) and CDKN1A (p21) loci, which otherwise drive tissue dysfunction, fibrosis, and chronic inflammation as humans age 4546. Recent multi-omic studies utilizing ATAC-seq and RNA-seq on peripheral blood mononuclear cells (PBMCs) from centenarians revealed distinct chromatin accessibility landscapes linked to exceptional longevity 49. Integrative analysis highlighted the E-26 transformation-specific (ETS)-related transcription factor ERG as a key longevity-associated regulator. ERG forms nuclear condensates through liquid-liquid phase separation, which structurally represses the transcription of senescence-related genes, notably CDKN2A, thereby attenuating cellular senescence phenotypes in extreme old age 49.
Furthermore, investigations into long-lived multigenerational families have identified rare, reduced-function missense mutations in CGAS (e.g., rs200818241) 53. The canonical cGAS-STING pathway detects cytosolic DNA (a marker of nuclear damage) and triggers SASP-related inflammation; dampening this pathway through inherited, reduced-function variants delays cellular senescence and limits chronic sterile inflammation (inflammaging) 53.
Additionally, the preservation of chromosomal ends is central to preventing the DNA Damage Response (DDR) that triggers senescence. While environmental factors influence telomere attrition, sequence analysis of the TERT and TERC genes (encoding telomerase components) in Ashkenazi centenarians revealed a significant overrepresentation of synonymous and intronic mutations 5051. These specific TERT haplotypes, alongside variants in the DNA repair gene Exonuclease 1 (EXO1), are directly linked to sustained telomere length and highly efficient mismatch repair mechanisms, protecting the genome from chromosomal erosion despite a century of continuous cellular replication 5051.
Conclusion
The genetics of extreme human longevity represents a highly coordinated, multi-system biological architecture. Decades of genomic research have dismantled the hypothesis that living past 100 is a purely random outcome of a healthy lifestyle. Instead, individuals reaching 100, 105, and 110 years rely heavily on inherited genetic resilience that buffers against the inevitable entropy of aging. This resilience is multifaceted: it involves a profound demographic depletion of rare, deleterious loss-of-function mutations in core metabolic genes; the presence of specific protective structural haplotypes, such as the African-origin rs10423769 variant that utilizes 3D chromatin looping to physically neutralize Alzheimer's risk; and finely tuned epistatic buffering across the WNT and AMPK pathways. The unique genomic signatures of supercentenarians reveal that evolutionary adaptations in specific genes - namely SIRT6, STK17A, and COA1 - effectively prioritize genomic stability, rapid DNA double-strand break repair, and controlled mitochondrial respiration, allowing these individuals to systematically delay or entirely evade the hallmark diseases of aging.