Genetic and environmental interplay in personality
The architecture of human personality is governed by an extraordinarily complex web of interactions between inherited genetic variants and dynamic environmental exposures. Historically, behavioral genetics relied on a dichotomous paradigm that attempted to partition the variance of human traits into broad genetic and environmental components using twin and family studies 12. Modern statistical genomics has fundamentally transcended this dichotomy. Advanced methodologies demonstrate that personality traits are highly polygenic, environmentally contingent, and biologically embedded through complex mechanisms of gene-environment interplay.
Recent breakthroughs in genome-wide association studies (GWAS), whole-genome sequencing (WGS), and multivariate statistical modeling have successfully mapped the precise molecular architecture of the Five-Factor Model (FFM) of personality: extraversion, neuroticism, agreeableness, conscientiousness, and openness to experience 23. Simultaneously, developmental psychology and behavioral epigenetics have refined the mathematical models used to measure how these polygenic predispositions react to, and are shaped by, socio-environmental contexts 456. This analysis examines the molecular genetic foundation of personality, the recent resolution of the "missing heritability" problem via whole-genome sequencing, the theoretical frameworks of gene-environment correlation (rGE) and interaction (GxE), and the profound pleiotropic overlap between normal personality variation and psychiatric disorders.
Molecular Architecture of Personality Traits
For decades, classical twin and adoption studies consistently demonstrated that human personality traits possess a broad-sense heritability ($H^2$) ranging from 30% to 60% 789. These studies indicated that additive genetic variance is a primary driver of phenotypic individual differences, with shared family environments contributing relatively little to long-term personality development compared to non-shared, idiosyncratic environmental factors 81011. However, traditional quantitative genetics could not identify the specific alleles responsible for this variation.
The advent of massive-scale Genome-Wide Association Studies (GWAS) fundamentally redefined the field by shifting the focus to single nucleotide polymorphism (SNP) heritability ($h^2_{SNP}$). Current evidence confirms that personality is a highly polygenic construct, influenced by thousands of common genetic variants, each conferring a minuscule individual effect size 912.
Genome-Wide Discoveries in the Five-Factor Model
A landmark 2024 meta-analysis by Gupta et al., leveraging data from the Million Veteran Program (MVP), the UK Biobank (UKB), and the Genetics of Personality Consortium (GPC), represents the most highly powered genomic investigation of the Big Five personality traits to date 131415. Encompassing sample sizes ranging from 237,390 to 682,688 individuals, this multi-ancestry GWAS successfully identified hundreds of independent genomic loci associated with personality traits, establishing definitive molecular correlates for the entire FFM taxonomy 314.
The variance explained by additive common SNPs ($h^2_{SNP}$) for these traits remains modest compared to classical twin estimates, ranging from 4% to 10% depending on the specific trait and the granularity of the measurement instrument 1317. Following initial locus discovery, researchers applied advanced variant fine-mapping algorithms, such as PolyFun, to identify the most probable causal variants among the thousands of highly correlated SNPs residing in linkage disequilibrium (LD) blocks 13.
| Personality Trait | Independent GWAS Loci | Novel Loci Discovered (2024) | Fine-Mapped Causal Variants | Estimated SNP Heritability ($h^2_{SNP}$) |
|---|---|---|---|---|
| Neuroticism | 208 | 62 | 155 | 6.0% - 10.0% |
| Extraversion | 14 | 14 | 8 | 7.0% |
| Openness | 7 | 7 | 3 | 5.0% |
| Conscientiousness | 2 | 2 | 7 | 5.0% |
| Agreeableness | 3 | 3 | 4 | 4.0% |
In total, 254 distinct genes were implicated across the five traits 23. To determine the functional impact of these loci, researchers utilized transcriptome-wide association studies (TWAS) and proteome-wide association studies (PWAS). These analyses indicated that personality-associated variants heavily regulate the expression of genes such as CRHR1, MAPT, MAD1L1, and SLC12A5 2313.
Tissue and Cellular Enrichment Profiles
The functional genomics of personality traits extend beyond single gene associations to specific neurobiological networks. Stratified linkage disequilibrium score regression (S-LDSC) applied to personality GWAS summary statistics reveals extreme functional enrichment in specific central nervous system cell types 1819.
Personality-associated genetic variants are predominantly enriched in cortical and subcortical neurons, particularly within GABAergic and glutamatergic transmission pathways 181916. Conversely, support cells such as glia show negligible heritability enrichment for personality phenotypes, confirming that the genetic architecture of the FFM is intrinsically tied to neuronal signaling and synaptic plasticity 19. Neurobiological theories aligning with these genomic findings posit a largely dopaminergic etiology driving variation in extraversion and openness to experience, juxtaposed against a serotonergic etiology underlying variation in conscientiousness, neuroticism, and agreeableness 1718.
The Missing Heritability Resolution
A central paradox in psychiatric and behavioral genetics over the past two decades has been the "missing heritability" problem. While twin and pedigree studies suggested that the broad-sense heritability of complex traits ranges from 40% to 80%, the cumulative variance explained by early GWAS - which exclusively measured common single-nucleotide polymorphisms via microarray genotyping - rarely exceeded 10% to 20% 192025. Various hypotheses were proposed to explain this massive gap, ranging from unaccounted gene-gene interactions (epistasis or "phantom heritability"), inflated classical twin estimates due to shared family environmental confounding, and the inability of GWAS microarrays to capture ultra-rare and structural variants 20212728.
In 2025, large-scale whole-genome sequencing (WGS) provided a definitive resolution to this debate. A highly powered Nature study analyzing the complete genomes of nearly half a million UK Biobank participants demonstrated that modern sequencing can capture the vast majority of pedigree-based heritability, directly observing the genetic variance that microarray platforms missed 252230.
The Contribution of Rare and Structural Variants
By measuring the full spectrum of genetic variation rather than relying on imputed common variants, researchers found that WGS data captured an average of 88% of the narrow-sense heritability previously established by family studies across dozens of complex biomedical and behavioral phenotypes 3023.
The sequencing data successfully decomposed this genetic variance, revealing that the "missing" gap was heavily populated by ultra-rare variants and non-coding changes. Across 34 densely mapped traits, the recovered heritability ratio highlighted severe functional enrichment within specific variant classes 2230.
| Variant Classification | Minor Allele Frequency (MAF) | Contribution to Recovered WGS Heritability | Genomic Representation |
|---|---|---|---|
| Common Variants | $\ge 1\%$ | 68% | Majority of the sequenced genome |
| Rare Coding Variants | $< 1\%$ | 21% (of the rare variant total) | $< 1\%$ of all analyzed variants |
| Rare Non-Coding Variants | $< 1\%$ | 79% (of the rare variant total) | Distributed across regulatory regions |
Coding regions, despite representing less than 1% of all analyzed variants, contributed roughly 20% of the rare variant heritability 2230. This implies a 26- to 36-fold functional enrichment for coding changes in shaping complex traits 22. Non-coding variants, although individually possessing much weaker effect sizes, collectively accounted for the majority of the rare variant contribution due to their massive quantity across regulatory regions 2230.
While WGS has largely solved the mystery of missing heritability by identifying the sum total of additive genetic variance, approximately 12% of the variance remains unaccounted for 22. Geneticists theorize that this residual fraction is driven by complex structural haplotypes currently missed by standard genome builds, extreme ultra-rare variants requiring sample sizes in the millions to reach statistical significance, and potential non-additive epistatic effects 202822.
Cross-Ancestry Transferability of Polygenic Scores
A critical limitation in modern statistical genomics is the overwhelming reliance on cohorts of European descent. According to demographic analyses of GWAS repositories, approximately 95% of participants are of European origin, which severely restricts the global transferability of genetic findings 24. Polygenic Risk Scores (PRS) constructed from European GWAS data suffer substantial performance degradation when applied to diverse ancestral populations. This phenomenon is driven by population-specific differences in linkage disequilibrium (LD) patterns, divergent minor allele frequencies, and unique socio-cultural environmental interactions 242526.
In cross-ancestry validation analyses, predictive accuracy precipitously drops as genetic distance from the training population increases. The median effect size of polygenic scores in African ancestry (AFR) samples is frequently estimated at only 42% of the performance observed in matched European (EUR) samples 25. East Asian (EAS) samples retain roughly 95% of the predictive accuracy, while South Asian (SAS) samples retain approximately 60% 25.
Recent collaborative efforts have begun to address this disparity through multi-ancestry meta-analyses. The 2024 MVP personality study included an African ancestry cohort of roughly 37,000 individuals, leading to the discovery of two novel loci specifically associated with agreeableness in the AFR population (ARGAP24 and CCDC6) that exhibited no significant signal in the EUR cohort 21417.
To optimize predictive utility across diverse clinical populations, researchers are deploying advanced algorithmic frameworks such as empirical Bayes estimation of cross-ancestry effects (e.g., SPLENDID) and multi-ancestry continuous shrinkage priors (PRS-CSx) 172627. Ensemble models utilizing algorithms like S4-Multi have demonstrated substantial improvements in predicting traits in non-European target samples; however, parity in absolute predictive power remains mathematically unachievable without massively expanding primary GWAS participation among underrepresented global populations 2426.
Genetic Pleiotropy and Personality Meta-Traits
The interplay of genetics and environment extends beyond normal personality variation, heavily overlapping with the etiology of clinical psychopathology. Large-scale GWAS have revealed profound genetic correlations - or pleiotropy - between the Big Five personality traits and a wide spectrum of psychiatric disorders, suggesting that personality structure and mental illness share foundational neurobiological architectures 132829.
The Meta-Traits of Stability and Plasticity
Recent applications of Genomic Structural Equation Modeling (Genomic SEM) have demonstrated that the genetic variance of the FFM can be statistically distilled into two higher-order genetic meta-traits: Stability and Plasticity 181930.
The Stability meta-trait captures the shared genetic variance across neuroticism (inversely), agreeableness, and conscientiousness. It encapsulates the intrinsic genetic propensity to regulate emotional, social, and behavioral impulses. The multivariate GWAS for the Stability meta-trait successfully identified 81 independent genome-wide significant loci 181930.
The Plasticity meta-trait isolates the shared genetic variance between extraversion and openness to experience. It models the genetic propensity for exploration, cognitive flexibility, and engagement with novel stimuli. A parallel GWAS for Plasticity mapped 13 independent genome-wide significant loci 1830.
Pleiotropy with Psychiatric Outcomes
The genetic architecture of these meta-traits is inextricably linked to the genetic risk for psychiatric conditions 30. Genomic analyses reveal a massive genetic correlation between trait neuroticism and major depressive disorder (MDD) ($r_g \approx 0.68$), as well as generalized anxiety disorder ($r_g \approx 0.80$) 13. Similarly, the Plasticity meta-trait and trait openness share significant genetic loci with schizophrenia risk, reflecting shared neurodevelopmental pathways related to cognitive salience attribution and dopaminergic activity 1330.
Studies identifying a "General Psychopathology Factor" (the p-factor, representing a generalized genetic liability to mental illness) demonstrate that the p-factor is heavily inversely correlated with the Stability personality meta-trait 3031. Genetic propensity for high Stability functions as an inherent protective factor against both internalizing and externalizing psychopathologies. This indicates that personality traits operate as the sub-clinical temperamental foundation upon which environmental stressors enact psychiatric disease 3031. When researchers condition GWAS models of MDD risk specifically on trait neuroticism, the variance explained is substantially attenuated, confirming that high trait neuroticism acts as a primary, early-life manifestation of the underlying genetic vulnerability that later erupts into clinical depression under environmental strain 314.
Frameworks of Gene-Environment Interplay
The high heritability of personality does not indicate genetic determinism. Contemporary behavioral genetics conceptualizes human development through the lens of gene-environment interplay, which is formally divided into two distinct, quantifiable biological mechanisms: Gene-Environment Correlation (rGE) and Gene-Environment Interaction (GxE) 432.
Gene-Environment Correlation Dynamics
Gene-environment correlation (rGE) describes the non-random distribution of environments among individuals of different genotypes. It posits that an individual's genetic makeup actively influences the environments they experience, effectively blurring the line between external stimuli and internal biology 433. rGE is operationalized via three mechanisms: 1. Passive rGE: Occurs predominantly in infancy and early childhood when biological parents provide both the child's genetic predispositions and the rearing environment. Because the child's genetics inherently correlate with the environment provided by their genetically similar parents, separating environmental causality from genetic transmission is highly difficult without adoption study designs 4. 2. Evocative rGE: Manifests when an individual's genetically influenced temperament elicits specific, predictable responses from the environment. For instance, a child with a high genetic propensity for agreeableness may consistently evoke warmer, more supportive parenting and peer interactions compared to a highly reactive or neurotic child, thereby actively shaping their own social environment 433. 3. Active rGE (Niche-Picking): Occurs when individuals actively select, modify, or construct environments that align with their genetic predispositions. As individuals age and gain executive autonomy, active rGE becomes the dominant developmental force. This mechanism explains the counter-intuitive phenomenon wherein the heritability of many cognitive and personality traits actually increases from early childhood into late adulthood 4.
Interaction Models and Environmental Sensitivity
Unlike rGE, which models how genes dictate exposure to environments, Gene-Environment Interaction (GxE) describes a reciprocal conditioning process: it occurs when the phenotypic effect of a genotype depends on the environmental context, or conversely, when the effect of an environment depends on the underlying genotype 432. In this paradigm, genetic factors are viewed as differential susceptibilities rather than rigid determinants.
When analyzing how distinct personality profiles react to environmental adversity and enrichment, researchers utilize three primary theoretical frameworks to interpret GxE interaction effects: the Diathesis-Stress model, the Vantage Sensitivity model, and the Differential Susceptibility model 634.
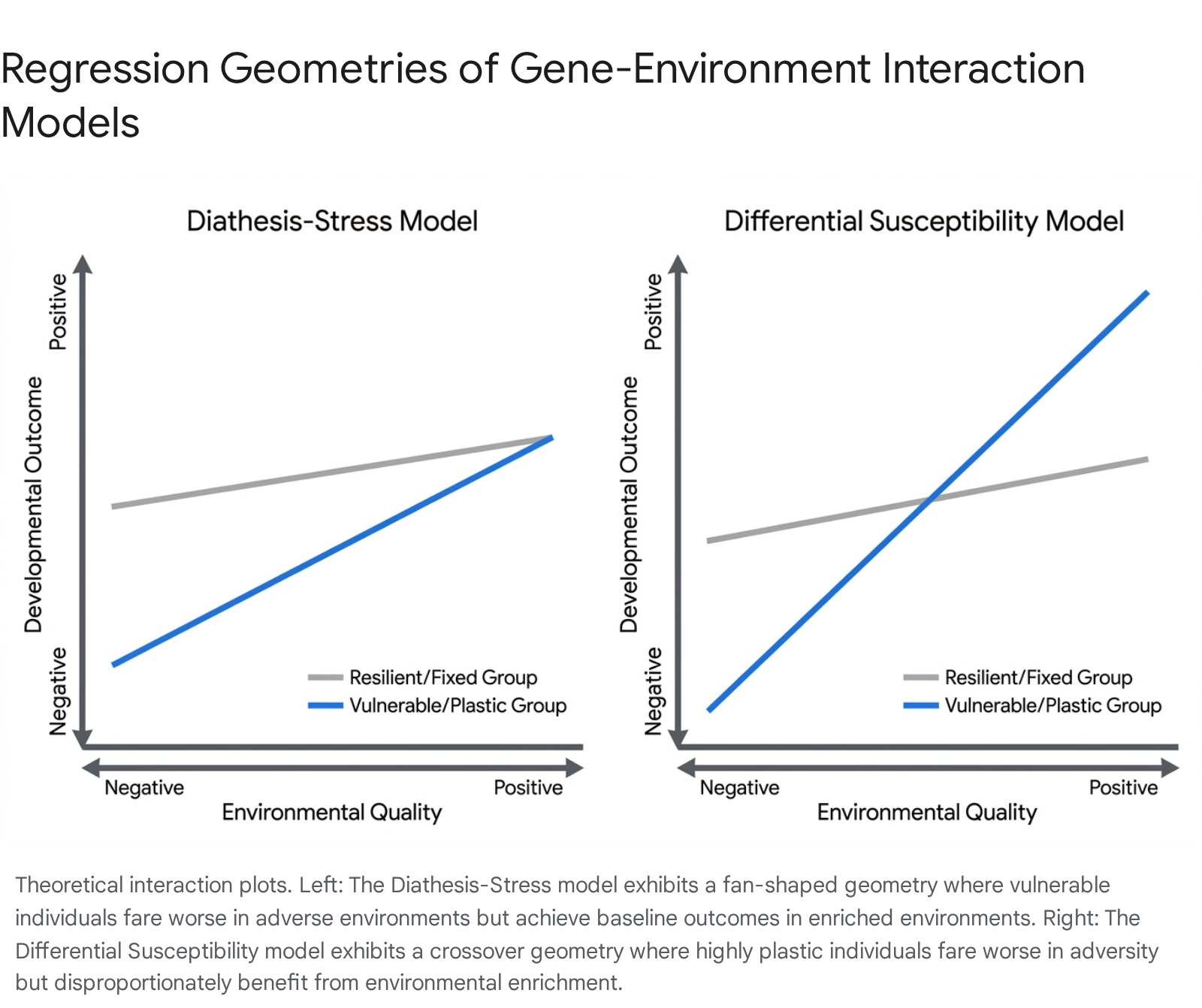
The historically dominant framework, the Diathesis-Stress model, posits that individuals possess varying levels of underlying vulnerability (diathesis) to adverse environments. In this model, an individual bearing a "risk" genotype or a difficult temperament will exhibit severe maladaptive outcomes when exposed to negative environments, such as poor parenting or severe socio-economic stress. However, in the presence of positive or neutral environments, these vulnerable individuals display outcomes virtually indistinguishable from their non-vulnerable peers 323536. Statistically, this translates to an ordinal or "fan-shaped" regression interaction, where the impact of environmental quality solely registers on the vulnerable group in the negative domain 3537.
Challenging the premise of unilateral vulnerability, the Differential Susceptibility hypothesis suggests that evolutionary pressures favored varying degrees of overall developmental plasticity. Rather than carrying strictly "vulnerability" genes, certain individuals carry "plasticity" genes. These individuals are exquisitely sensitive to their environmental contexts - for better and for worse 343637. Under differential susceptibility, a highly plastic child will indeed suffer worse outcomes in an abusive environment compared to a resilient peer. However, if that same highly plastic child is placed in an enriched, supportive environment, they will achieve significantly better outcomes than the resilient peer, who remains relatively unaffected by environmental extremes 343538. This generates a disordinal or "crossover" interaction in statistical models 3539.
Quantitative Metrics for Interaction Geometry
To rigorously differentiate between diathesis-stress and differential susceptibility in empirical data, statistical geneticists rely on formal quantitative criteria based on the precise geometry of interaction regression lines 638. Subjective visual interpretation of interaction plots is highly discouraged in favor of the following standardized indices 53437.
| Metric | Definition | Indicator for Diathesis-Stress | Indicator for Differential Susceptibility |
|---|---|---|---|
| Region of Significance (RoS) | The specific environmental bounds where genetic slopes significantly diverge. | Significant divergence is isolated solely to the negative environmental extreme. | Significant divergence occurs at both the negative and positive environmental extremes. |
| Proportion of Interaction (PoI) | The area ratio of the interaction bounded by the crossover point. | $\sim 0.00$ to $0.20$ (ordinal/fan-shape). | $0.20$ to $0.80$ (ideal is $\sim 0.50$; disordinal/crossover). |
| Proportion Affected (PA) | Percentage of the empirical sample benefiting from the "for better" crossover. | $0\%$ to $<2\%$ (No substantial benefit observed in the sample). | $\ge 16\%$ of the empirical sample population exists above the crossover threshold. |
The Proportion of Interaction (PoI) defines the total area between the regression lines on either side of the crossover point. A PoI value near 0.00 indicates a fan-shaped interaction, whereas a PoI approaching 0.50 indicates a perfectly symmetrical crossover effect characteristic of pure differential susceptibility 5363738. Furthermore, the Proportion Affected (PA) index calculates the proportion of the actual sample data that falls on the "for better" side of the crossover point. For an interaction to be confidently classified as differential susceptibility, the PA index must confirm that a nontrivial portion of the population - specifically at least 16% - actively benefits from the interaction in a positive environment 53839.
Epigenetic Mechanisms of Environmental Embedding
The statistical models of gene-environment interaction dictate a theoretical need for a physical mechanism through which transient environmental exposures can exert long-lasting, highly stable effects on personality and behavior. Epigenetics provides this molecular bridge 440.
While the sequence of the genome remains fixed throughout the lifespan, the epigenome acts as a dynamic regulatory layer that dictates gene expression and suppression. Environmental stimuli - ranging from early childhood trauma to chronic psychosocial stress - can trigger profound biochemical modifications to this regulatory layer. The most heavily researched epigenetic mechanisms are DNA methylation, which involves the addition of methyl groups to CpG dinucleotides to silence gene transcription, and histone modification, involving the acetylation or methylation of the proteins around which DNA is spooled to alter chromatin accessibility 440.
These epigenetic marks frequently target and regulate genes involved in neuroendocrine signaling, particularly those governing the hypothalamic-pituitary-adrenal (HPA) axis, which manages the body's physiological stress response 440. Consequently, the abstract concept of gene-environment interplay is physically written onto the genome. A genetic predisposition toward high trait neuroticism may remain entirely latent unless adverse environmental stress triggers the specific epigenetic methylation required to alter HPA axis reactivity 440. The extreme persistence of these epigenetic marks across cell divisions explains the enduring stability of certain personality traits and stress reactivities long after the initial environmental trigger has resolved.
As statistical genomics continues to evolve - moving beyond narrow European microarray data to embrace diverse, multi-ancestry whole-genome sequencing - the field edges closer to a truly precise understanding of personality. By integrating massive multi-omic datasets with rigorous environmental interaction models, researchers will eventually untangle the specific, localized mechanisms through which individual biology and external experience converge to shape human character.