Epigenetic reprogramming and aging reversal with Yamanaka factors
The progressive physical and cognitive decline associated with biological aging is increasingly understood not merely as the accumulation of irreversible metabolic damage, but as a fundamental degradation of epigenetic information. Epigenetic reprogramming - the application of exogenous transcription factors or small molecules to revert mature, specialized cells into a more youthful or pluripotent state - has emerged as a central focus of gerontological research. The application of the Yamanaka factors (OCT4, SOX2, KLF4, and c-MYC) to erase age-associated epigenetic signatures has demonstrated profound restorative effects in vitro and in select in vivo models.
However, translating these phenomena into systemic human therapeutics involves navigating immense biological complexity. The field is currently characterized by intense debate regarding the underlying mechanisms of aging, the reliability of in vivo reprogramming models, the safety of systemic cellular dedifferentiation, and the accuracy of the biomarkers used to measure rejuvenation.
Conceptual Framework of Biological Aging
Aging biology encompasses multiple interconnected systemic failures, frequently categorized into established hallmarks. These include genomic instability, telomere attrition, mitochondrial dysfunction, cellular senescence, loss of proteostasis, disabled macroautophagy, deregulated nutrient sensing, stem cell exhaustion, altered intercellular communication, chronic inflammation, dysbiosis, and epigenetic alterations 123. Among these, epigenetic alterations represent a unique paradigm because they signify a loss of regulatory software rather than a permanent destruction of the underlying genetic hardware 4.
The Hallmarks of Aging and Epigenetic Drift
Throughout an organism's lifespan, the epigenome undergoes a continuous accumulation of stochastic perturbations, frequently termed "epigenetic noise" or epigenetic drift. This phenomenon is characterized by a gradual alteration in DNA methylation patterns across the genome, the loss of precise histone post-translational modifications, and the overall dysregulation of chromatin architecture 125. The result is a unidirectional transition from a highly ordered, cell-type-specific transcriptional state to a state of increased informational entropy.
In a youthful cell, the epigenome tightly regulates gene expression, silencing aberrant pathways and maintaining the cell's specialized identity. Over time, environmental stressors, DNA repair events, and metabolic byproducts disrupt these regulatory marks. Histone acetylation tends to decrease with age, leading to condensed chromatin structures that suppress the expression of necessary cellular maintenance genes 23. A recent assessment of postmortem human brain tissue revealed an upregulation of two histone acetyltransferases, H3K27ac and H3K9ac, linking aberrant epigenetic shifts directly with amyloid-beta pathology and neurodegeneration 2. Consequently, cells gradually lose their specialized functions and experience an impaired capacity to execute DNA repair, maintain proteostasis, and resist extrinsic stress 1.
This progressive loss of genome-epigenome coordination has been described as a natural, non-programmed limit on lifespan. Epigenetic factors are proposed to act as molecular "shields," protecting genomic regions from mutations; as these shields degrade with age, an "epigenetic ratchet" accelerates the rate of permanent genetic mutations 6.
Interactions with Mitochondrial Dysfunction and Telomere Attrition
The loss of epigenetic fidelity does not occur in isolation; it functions as both a primary driver and a downstream consequence of other aging hallmarks. A critical and highly reinforcing interaction occurs within the epigenome-mitochondria-telomere axis. Mitochondrial dysfunction - characterized by the accumulation of reactive oxygen species (ROS) and impaired oxidative phosphorylation - triggers retrograde signaling pathways from the mitochondria directly to the nucleus 76. Because mitochondrial DNA lacks the protective histones found in nuclear DNA, it is highly susceptible to the oxidative stress generated by the adjacent electron transport chain 1.
Mitochondrial stress induces the activation of the coactivator heterogeneous ribonucleoprotein A2 (hnRNPA2), a molecule that possesses intrinsic histone acetyltransferase (HAT) activity 76. The hnRNPA2-mediated acetylation of histone H4 at lysine 8 (H4K8) acts as a specific epigenetic signal that accelerates telomere attrition at the ends of chromosomes 76. Therefore, mitochondrial dysfunction can drive telomeric loss independently of classical cellular replication limits (the Hayflick limit), mediated entirely by epigenetic signaling 67.
Conversely, telomere damage activates the p53 signaling pathway and inhibits the PGC-1α pathway (a key regulator of mitochondrial biosynthesis). This creates a self-reinforcing feedback loop that further impairs mitochondrial metabolism, exacerbating the energetic deficit that impairs ATP-dependent processes like DNA repair and protein folding 189.
The Yamanaka Factors and Cellular Rejuvenation
The foundation of modern cellular reprogramming relies on the 2006 discovery by Takahashi and Yamanaka, demonstrating that the ectopic expression of four specific transcription factors (OCT4, SOX2, KLF4, and c-MYC, collectively known as OSKM) can force mature adult somatic cells to revert into induced pluripotent stem cells (iPSCs) 101112.
Complete Dedifferentiation Versus Partial Reprogramming
Prolonged exposure to OSKM completely erases cellular identity. In this state, the epigenetic clock is reset to nearly zero, telomeres are significantly elongated, and the cell regains the embryonic capacity to differentiate into any cell type within the body 1314. While complete reprogramming is an invaluable tool for generating ex vivo cell lines for research and autologous transplantation, it is strictly contraindicated as a systemic in vivo anti-aging therapy. Systemic, continuous expression of OSKM in living organisms results in widespread dedifferentiation, the complete loss of organ function, and the rapid formation of lethal teratomas 121415.
To harness the rejuvenating properties of the Yamanaka factors without triggering oncogenesis or organ failure, researchers developed the concept of "partial reprogramming" (also referred to as transient epigenetic reprogramming). Partial reprogramming involves delivering the factors for a highly calibrated duration - sufficient to erase age-associated epigenetic drift and restore youthful transcriptional profiles, but withdrawn immediately before the cell crosses the threshold of dedifferentiation 101416.
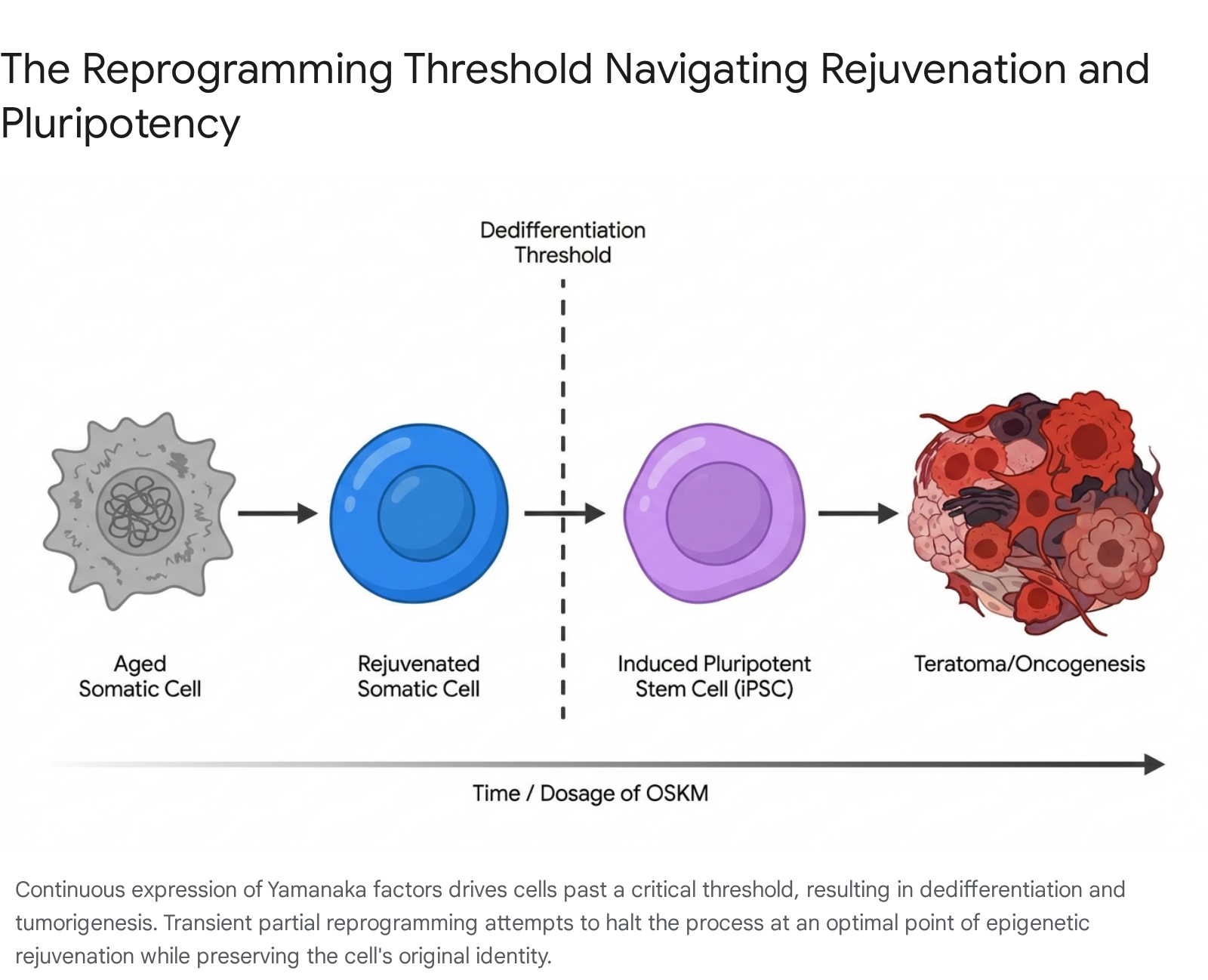
In vitro studies demonstrate that short-term OSKM induction reduces the epigenetic age of human fibroblasts, beginning between days three and seven post-induction, and progressing at a rate of approximately 3.8 years per day 13. In 2016, researchers demonstrated that cyclic, transient expression of OSKM could significantly extend the lifespan of mice suffering from premature aging (progeria) while concurrently improving the regenerative capacity of aged wild-type tissues in the pancreas and skeletal muscle 111617. To further mitigate oncogenic risks, modern therapeutic iterations frequently omit the c-MYC factor entirely. Utilizing an OSK formulation (OCT4, SOX2, KLF4) extends the safe temporal window for transient expression, allowing for deeper epigenetic resetting without triggering malignant transformation 1016.
Epigenetic Clocks and the Measurement of Biological Age
To quantify the efficacy of cellular reprogramming interventions, the field relies heavily on epigenetic aging clocks. These machine learning models analyze predictable changes in DNA methylation (DNAm) across specific loci in the genome to estimate an organism's or a tissue's "biological age" relative to chronological age 181920. First-generation multi-tissue clocks, such as the original Horvath clock, were designed to capture a universal aging signature shared across virtually all tissue types 13. Second-generation "phenotypic" clocks attempt to incorporate physiological phenotypes and mortality risk factors to enhance predictions of overall healthspan 19.
Despite their ubiquitous application in longevity research, epigenetic clocks present significant limitations when evaluating novel reprogramming interventions. A 2024 analysis published in Aging Cell demonstrated that current clock models suffer from profound epistemic uncertainty 18. The primary flaw lies in the machine learning training datasets: the specific DNA methylation profiles generated during artificial cellular reprogramming are fundamentally poorly represented in the standard, natural aging datasets used to train the models 18.
Consequently, the algorithms are forced to extrapolate outside their learned biological parameters. Predictions generated by different published clocks are frequently highly inconsistent when applied to the exact same reprogrammed cells, with some models suggesting massive age reversal while others indicate zero or even negative rejuvenation 18. Furthermore, a reduction in measured epigenetic age does not necessarily guarantee true physiological improvement. Clocks capture specific dimensions of aging but frequently reflect confounding factors, such as shifts in local blood cell populations or transient metabolic stress responses, rather than a permanent reversal of intrinsic aging mechanisms 13. Therefore, while epigenetic clocks serve as qualitative or binary indicators of molecular shifts, their precise quantitative predictions regarding OSK-mediated rejuvenation remain inherently unverifiable until correlated with long-term survival and functional healthspan metrics in vivo 1819.
The Information Theory of Aging Debate
The theoretical framework explaining exactly why partial reprogramming succeeds is the subject of intense academic debate. David Sinclair and colleagues have proposed the "Information Theory of Aging," which posits that biological aging is driven primarily by the loss of epigenetic information rather than the accumulation of permanent genetic mutations 4212223.
The Dual-Layer Inheritance Hypothesis
According to this theory, the mammalian cellular state functions as a dual-layer information system. While the DNA sequence is digital and highly stable (preserving the hardware blueprint), the epigenome relies on a more fragile analog system to dictate gene expression (the software) 410. Over time, DNA repair events distract epigenetic modifiers from their standard regulatory roles; as they rush to repair double-strand breaks, they fail to return to their original loci, gradually corrupting the epigenome 4.
The Information Theory hypothesizes the existence of a cellular "biological observer" - a latent backup copy of youthful epigenetic information that survives the aging process intact 23. By utilizing the OSK factors, cells can theoretically access this backup, executing a process akin to "reinstalling software" to correct the accumulated epigenetic noise without altering cellular identity 424.
Methodological Critiques and the DNA Damage Confound
The Information Theory of Aging, and the highly publicized claims surrounding age reversal, have faced severe scrutiny from the broader gerontological community. In early 2024, researchers James Timmons and Charles Brenner published a formal, peer-reviewed critique in Cell titled "The information theory of aging has not been tested" 242526.
Critics argue that the experimental designs utilized to support the Information Theory fail to isolate epigenetic information loss from other confounding variables, primarily genotoxic stress and cell-selection effects 2224. For example, the Inducible Changes to the Epigenome (ICE) mouse model used by the Sinclair laboratory introduces non-mutagenic double-strand breaks to deliberately accelerate epigenetic aging 4. Critics assert that this method inherently activates p53-mediated apoptotic (cell death) pathways. Consequently, the observed tissue changes may result from the clearance of damaged cells and subsequent compensatory proliferation of neighboring healthy cells, rather than a pure epigenetic "reboot" within individual surviving cells 24.
Furthermore, critics argue that characterizing OSK transcription factors as a safe "software reinstall" drastically minimizes their established oncogenic nature 2427. The hyperfunction theory of aging presents an alternative view, suggesting that aging is not a software corruption, but rather the quasi-programmed continuation of developmental growth pathways (such as mTOR) that become hyperactive and detrimental later in life 23.
| Aspect of the Debate | Information Theory of Aging (Proponents) | Mainstream and Damage Theories (Critics) |
|---|---|---|
| Primary Driver of Aging | Loss of epigenetic regulatory information (software corruption and entropy) 1023. | Accumulation of macromolecular damage, mutations, and hyperfunction (hardware failure) 4623. |
| Role of DNA Damage | DNA repair distracts epigenetic modifiers, causing them to drift from target loci 4. | DNA damage is a direct, toxic driver of dysfunction, apoptosis, and cellular senescence 124. |
| Mechanism of OSK Action | OSK accesses a "backup copy" of youthful epigenetic data, restoring original states 2324. | OSK triggers general dedifferentiation; apparent rejuvenation may be an artifact of cell selection or trophic responses 2428. |
| Safety Profile | Partial reprogramming via OSK cleanly circumvents pluripotency and avoids cancer 1029. | Yamanaka factors are inherent oncogenes; safety margins in vivo are narrow and untested in large human cohorts 1227. |
In Vivo Reprogramming and the Reproducibility Crisis
The translation of cellular reprogramming from isolated in vitro cell cultures to the complex three-dimensional architecture of a living organism introduces extreme variables. The reliability of in vivo reprogramming research is currently hindered by the broader biomedical reproducibility crisis.
General Replicability Challenges in Biomedicine
Current estimates suggest that up to 50% to 65% of preclinical biomedical research faces significant replication challenges, wasting an estimated $28 billion annually in the United States alone 303431. A late 2024 global survey published in PLOS Biology indicated that 72% of biomedical researchers agree the field faces a reproducibility crisis, largely driven by the "publish or perish" culture, poorly planned studies, and a lack of rigorous standardization 32. However, others argue that biological diversity inherently resists exact replication; living organisms possess degenerate pathways, meaning multiple molecular mechanisms can produce the same physiological behavior 33. A failure to replicate a finding does not universally indicate flawed original research, but often exposes the intrinsic variability of the biological system or an overstatement of the initial conclusions 3033.
The Glia-to-Neuron Transdifferentiation Controversy
Within the specific field of in vivo cellular reprogramming, this reproducibility crisis has crystallized around the highly controversial attempts to transdifferentiate glial cells directly into neurons within the living adult brain.
Neurological diseases like Parkinson's are characterized by the irreversible loss of specific neuronal populations, notably dopaminergic neurons in the substantia nigra. Early in vivo reprogramming studies suggested that delivering proneuronal transcription factors (like NeuroD1) or knocking down the RNA-binding protein PTBP1 could successfully convert endogenous, non-neuronal astrocytes directly into functional neurons in situ, entirely bypassing the intermediate pluripotent state 343536. These initial breakthrough studies relied heavily on Adeno-Associated Viruses (AAVs) carrying reporter genes (like mCherry, RFP, or GFP) driven by supposedly astrocyte-specific promoters, such as the Glial Fibrillary Acidic Protein (GFAP) promoter 3536. The sudden appearance of reporter-positive mature neurons in the brains of mice was widely celebrated as definitive proof of successful in vivo cell conversion 3537.
Resolving Artifacts with Genetic Lineage Tracing
However, subsequent independent laboratories applying highly rigorous genetic lineage-tracing models completely failed to reproduce these neurogenesis findings 363839. When researchers traced the origin of cells using transgenic mouse lines (such as Aldh1l1-CreERT2) rather than relying solely on viral reporters, they discovered that the newly generated neurons did not originate from astrocytes 2836.
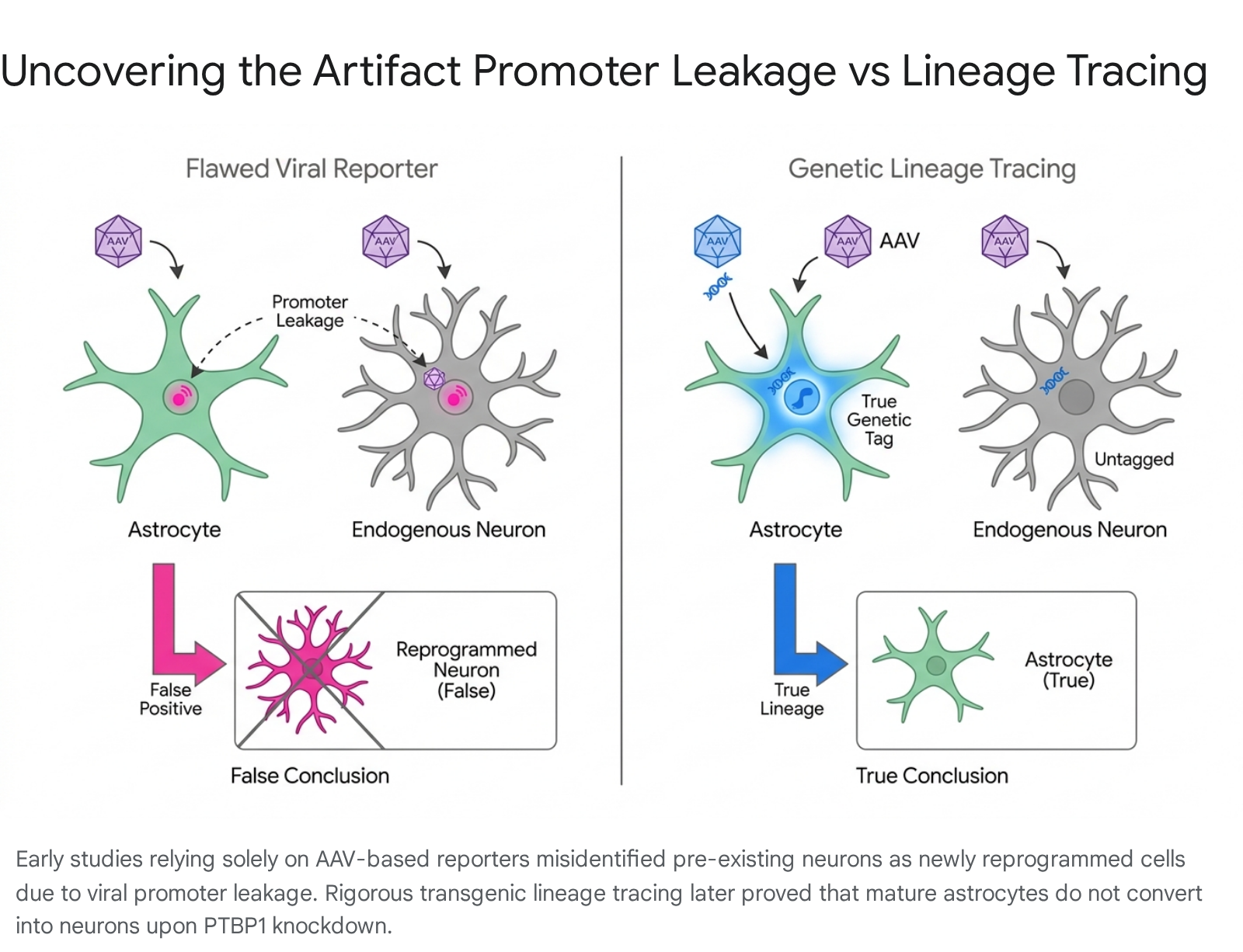
Instead, the phenomenon was driven entirely by "promoter leakage." Under certain conditions - especially when AAVs are administered at high titers or when they express highly potent neurogenic factors like NeuroD1 - the viruses aberrantly express their reporter genes directly in endogenous, pre-existing neurons 283637. The original studies had observed endogenously labeled neurons and misidentified them as newly converted astrocytes.
Extensive RNA splicing analysis and strict lineage tracing confirmed that depleting PTBP1 in mature astrocytes is entirely insufficient to induce neuronal conversion 283839. Apparent neurological improvements observed in earlier animal models were likely due to trophic responses - whereby the viral intervention incidentally reduced local inflammation or altered the cellular environment, improving tissue health without achieving actual transdifferentiation 28. This ongoing controversy underscores the severe epistemic risks of in vivo reprogramming, demanding extreme methodological stringency to separate genuine epigenetic shifts from viral artifacts.
Delivery Modalities and Manufacturing Bottlenecks
Moving partial reprogramming from the laboratory into widespread clinical practice necessitates the development of delivery systems that are safe, tissue-specific, and economically scalable. The choice of modality dictates the therapeutic limits of the intervention.
Adeno-Associated Viral Vectors
Adeno-Associated Virus (AAV) vectors remain the industry workhorse for in vivo gene delivery due to their minimal immunogenicity and their ability to provide highly durable expression in static, post-mitotic tissues 4041. However, unmodified AAV relies heavily on its native tropism, frequently requiring massive systemic doses to achieve adequate transduction in target organs. These high vector doses frequently result in severe hepatotoxicity and neutralizing immune responses that preclude future re-dosing 41.
Furthermore, AAV manufacturing faces critical commercial bottlenecks. The standard upstream production process yields a significant percentage of "empty capsids" - viral protein shells that lack the therapeutic DNA payload 4243. Empty capsids act as product-related impurities that increase the required overall viral load, exacerbating host immune responses 4243. Separating empty capsids from full capsids requires difficult-to-scale downstream methods like ultracentrifugation or advanced affinity and ion-exchange chromatography, drastically inflating the cost of goods sold (COGS) 404243. To overcome native tropism limitations, researchers are employing click chemistry techniques, such as strain-promoted azide-alkyne cycloaddition (SPAAC), to incorporate unnatural amino acids into the AAV capsid, allowing for the covalent conjugation of specific targeting nanobodies 41.
Non-Viral Lipid Nanoparticles
Non-viral vectors, specifically Lipid Nanoparticles (LNPs), offer an alternative capable of delivering mRNA encoding the Yamanaka factors. LNPs avoid the long-term genomic integration risks of AAVs, allowing for transient, temporary expression of reprogramming factors. This transient nature aligns perfectly with the safety requirements of partial reprogramming, and LNPs are generally re-dosable 40.
However, systemically administered LNPs inherently accumulate in the liver. Upon intravenous injection, LNPs absorb circulating apolipoprotein E (ApoE) and undergo rapid endocytosis mediated by low-density lipoprotein receptors (LDLR) on the surface of hepatocytes 44. Achieving extrahepatic targeting - delivering epigenetic reprogramming factors specifically to the brain, heart, or skeletal muscle - remains a profound biochemical hurdle, requiring intricate functionalization of the LNP surface with highly specific targeting ligands or peptides 44.
Chemical Reprogramming Cocktails
To completely circumvent the oncogenic risks, immunogenicity, and manufacturing complexities of genetic delivery, researchers have developed entirely chemical reprogramming cocktails. These cocktails utilize combinations of small molecule drugs to manipulate cellular signaling pathways and inhibit specific chromatin-modifying enzymes 4546.
Recent developments include a seven-compound (7c) cocktail - comprising CHIR99021, DZNep, Forskolin, TTNPB, Valproic acid (VPA), Repsox, and Tranylcypromine (TCP) - as well as an optimized two-compound (2c) variant 454647. While short-term treatment with the 7c cocktail successfully reduces DNA damage markers (like γH2AX) and ameliorates specific epigenetic dysregulation in aged human fibroblasts, it also induces significant cellular stress 4546. The 7c cocktail frequently impairs overall cellular proliferation and heavily upregulates the production of reactive oxygen species (ROS) and inflammatory cytokines, notably IL-6 454647. Consequently, while chemical reprogramming successfully avoids viral integration, its current lack of safe, multiparametric rejuvenation efficiency limits its immediate systemic application 45.
| Modality | Primary Advantages | Key Bottlenecks and Challenges | Optimal Application |
|---|---|---|---|
| AAV Vectors | Durable expression; extensive clinical validation; highly effective for localized delivery. 4041 | Suboptimal tissue specificity; hepatotoxicity at high systemic doses; high rate of empty capsids inflating manufacturing costs. 414243 | Single-dose interventions; targeted local therapies (e.g., intraocular injections). 40 |
| Lipid Nanoparticles (LNPs) | Non-integrating (transient mRNA expression); scalable chemical synthesis; re-dosable. 4044 | Overwhelming accumulation in the liver via ApoE/LDLR pathways; requires complex engineering for extrahepatic targeting. 44 | Therapies requiring highly controlled, repeated, and transient dosing. 4044 |
| Chemical Cocktails (Small Molecules) | Eliminates viral integration and teratoma risks; standard pharmacological regulatory and manufacturing pathways. 4546 | Low efficiency; frequently induces oxidative stress, upregulates inflammation (IL-6), and impairs cell proliferation. 4547 | Ex vivo cellular rejuvenation; early-stage localized tissue repair. 4647 |
Clinical Translation and Ex Vivo Therapies
Despite the theoretical debates and technical delivery bottlenecks, the clinical translation of epigenetic reprogramming is accelerating, driven primarily by localized interventions and ex vivo cellular therapies where safety thresholds are easier to manage.
Ocular Reprogramming and Phase 1 Trials
In early 2026, Life Biosciences received Investigational New Drug (IND) clearance from the U.S. FDA for ER-100, marking the first human clinical trial for an epigenetic cellular rejuvenation therapy 294849. ER-100 utilizes an AAV2 vector to deliver a proprietary partial reprogramming payload consisting of three Yamanaka factors (OCT4, SOX2, and KLF4, omitting c-MYC to eliminate oncogenic risk) directly into the eye via intravitreal injection 294850.
The Phase 1 first-in-human study targets severe optic neuropathies, specifically open-angle glaucoma (OAG) and non-arteritic anterior ischemic optic neuropathy (NAION), diseases characterized by the irreversible degeneration of retinal ganglion cells 4950. Preclinical data in non-human primates demonstrated that ER-100 successfully restored cellular methylation patterns and generated measurable improvements in visual function, as quantified by pattern electroretinogram (pERG) analyses, following induced optic nerve injury 294950. By restricting the gene therapy to the enclosed, immune-privileged environment of the eye, developers bypass the systemic delivery bottlenecks and hepatotoxicity risks while directly testing the Information Theory of Aging in living human tissue 48.
Pluripotent Stem Cell Therapies in Spinal Cord Injury
Parallel advancements are occurring in ex vivo stem cell therapies, where reprogramming is taken to completion in a laboratory setting before therapeutic administration. In 2025, researchers at Keio University in Japan, led by Hideyuki Okano, achieved a landmark breakthrough in treating severe spinal cord injuries 515253. The clinical trial safely implanted over two million neural stem cells - derived from fully reprogrammed induced pluripotent stem cells (iPSCs) - into four paralyzed adult patients 5152.
To prevent rejection, the patients were maintained on immunosuppressive medication for six months. Following a year of monitoring, no serious adverse events or teratomas were observed 52. Remarkably, two of the four recipients exhibited measurable improvements in motor function, with one elderly patient regaining the ability to stand without support 5152. While larger cohort trials are required to determine long-term cellular survival and efficacy, the study provides vital proof-of-concept for the safety of iPSC-derived cellular replacement in the central nervous system.
Primate Models of Systemic Rejuvenation
Research into the systemic administration of reprogrammed cells is also yielding significant preclinical results. In 2025, researchers at the Chinese Academy of Sciences (CAS) published landmark findings in Cell detailing the systemic rejuvenation of elderly non-human primates 5455. The team engineered a novel cell line known as Senescence-Resistant mesenchymal progenitor Cells (SRCs) by reprogramming specific longevity-associated genetic pathways, notably the FOXO3 gene 5455.
Biweekly intravenous infusions of SRCs over 44 weeks into elderly cynomolgus monkeys resulted in multi-system rejuvenation across 10 major physiological systems and 61 distinct tissue types 5455. The intervention reduced brain atrophy, reversed osteoporosis, and restored reproductive function 54. The restorative effects were driven primarily by exosomes - microscopic extracellular vesicles released by the SRCs - that systematically suppressed chronic inflammation and restored epigenomic integrity throughout the organism 5455. Machine learning-based aging clocks estimated that the biological age of immature neurons and oocytes was reversed by five to seven years. Crucially, the SRCs navigated the safety threshold successfully, engrafting without inducing tumors or structural tissue damage 5455. Furthermore, Chinese researchers at Tsinghua University have recently identified a specific chemical cocktail (TAW, comprising TTNPB, 1-Azakenpaullone, and WS6) capable of elevating pluripotent stem cells to a totipotent state, highlighting rapid advances in non-genetic cellular manipulation 56.
Global Institutional Research Networks
The pursuit of epigenetic reprogramming and aging biology is highly distributed, supported by massive institutional investments globally.
In Europe, the European Research Council (ERC) heavily funds extensive scientific networks bridging the disciplines of epigenetics and systems biology. Initiatives such as the EpiGeneSys network and the NutriSTEM project focus on illuminating the epigenetic transitions that dictate cellular memory and hematopoietic stem cell aging 57. Institutions like the Max Planck Institute of Immunobiology and Epigenetics, and the Helmholtz Munich research center, serve as primary international hubs 575859. European researchers, including teams at the Babraham Institute, have extensively mapped how dietary restriction triggers physiological epigenetic changes that reprogram lipid metabolism and protect against age-induced methylation drift 60.
In Asia, Japan continues to aggressively leverage the Nobel-winning discovery of the Yamanaka factors, leading the world in the clinical translation of iPSC-derived cellular therapeutics for neurological applications 51. Concurrently, Chinese institutions, spearheaded by the Chinese Academy of Sciences, Capital Medical University, and Tsinghua University, have rapidly advanced the development of both chemical reprogramming cocktails and large-scale non-human primate longevity models 545556. Additional research from Northwest A&F University has identified critical genetic switches, such as the Aldh1a2 retinoic acid pathway, that govern mammalian tissue regeneration, broadening the scope of regenerative medicine beyond pure epigenetic reprogramming 61.
The proposition that the physiological decay of aging can be reversed by resetting the epigenome represents one of the most profound paradigm shifts in modern biology. The transition from viewing aging as irreversible hardware degradation to a reversible software corruption opens unprecedented avenues for regenerative therapies. However, the path to systemic human rejuvenation remains fraught with intense epistemic and mechanical challenges. As partial epigenetic reprogramming enters phase 1 clinical trials, the field will soon acquire the empirical human data necessary to separate theoretical optimism from medical reality.