Current status of in vivo partial cellular reprogramming
The paradigm of organismal aging has historically been understood as a unidirectional and irreversible accumulation of molecular damage, cellular senescence, and epigenetic erosion. This classical understanding was fundamentally disrupted by the advent of cellular reprogramming. Originating with the discovery by Shinya Yamanaka in 2006 that differentiated adult somatic cells could be reverted to embryonic-like induced pluripotent stem cells (iPSCs) via the transient expression of four transcription factors - Oct4, Sox2, Klf4, and c-Myc (collectively known as OSKM or the Yamanaka factors) - the field established that cellular identity and biological age are highly plastic 123. However, full in vivo reprogramming results in the total loss of cellular identity and the formation of lethal teratomas across multiple organ systems 154.
To harness the rejuvenating potential of this discovery without triggering oncogenesis, researchers developed "partial reprogramming." This process exposes cells to reprogramming factors just long enough to erase epigenetic markers of aging, withdrawing the factors before the cell crosses the threshold of dedifferentiation 567. Spearheaded by foundational in vivo research in 2016 by Juan Carlos Izpisua Belmonte and Alejandro Ocampo at the Salk Institute, and now heavily capitalized by biotechnology entities such as Altos Labs, partial reprogramming represents the frontier of geroscience and regenerative medicine 101112. As the field transitions from preclinical mammalian models to early-phase human clinical trials in 2025 and 2026, research has expanded into optimized factor configurations, artificial intelligence-driven target discovery, and chemical small-molecule alternatives designed to circumvent the risks of transgenic integration 131415.
Fundamental Biology of Cellular Reprogramming
The core mechanism of partial reprogramming relies on the temporal uncoupling of two distinct biological processes: age reprogramming, which involves the erasure of age-associated epigenetic and transcriptomic signatures, and developmental reprogramming, which entails the permanent loss of somatic identity 16.
Epigenetic Remodeling and Chromatin Reconstitution
During the physiological aging process, cells experience a degradation of the epigenetic landscape. This degradation is characterized by the progressive loss of heterochromatin, alterations in DNA methylation fidelity, and global genomic instability 28. Short-term, cyclic induction of OSKM initiates a profound chromatin remodeling process that directly counteracts these age-associated changes 29.
In vitro time-course experiments analyzing single-cell transcriptomics reveal that the trajectory of reprogramming follows a highly specific timeline. When adult somatic cells, such as human dermal fibroblasts, are exposed to OSKM, they enter a "zone of epigenetic instability" where age-associated signatures begin to rapidly reverse 6. Rejuvenation of the epigenome is marked by the restoration of Heterochromatin Protein 1 beta (HP1β) mobility, the reconstitution of H3K9me3, H4K20me3, and H3K36me2 histone marks, and a significant reduction in senescence-associated gene expression 1710. If the reprogramming factors are withdrawn between days 10 and 13 of this process, the cells safely revert to their original somatic identity, yet they retain a younger epigenetic and transcriptomic profile 67. Single-cell RNA sequencing data has shown that such transiently reprogrammed cells maintain down-regulated age-associated gene expression and upregulated expression of youthful markers, such as collagens, for weeks after the withdrawal of OSKM 7.
The Threshold of Dedifferentiation
A critical "point of no return" exists within the reprogramming timeline.
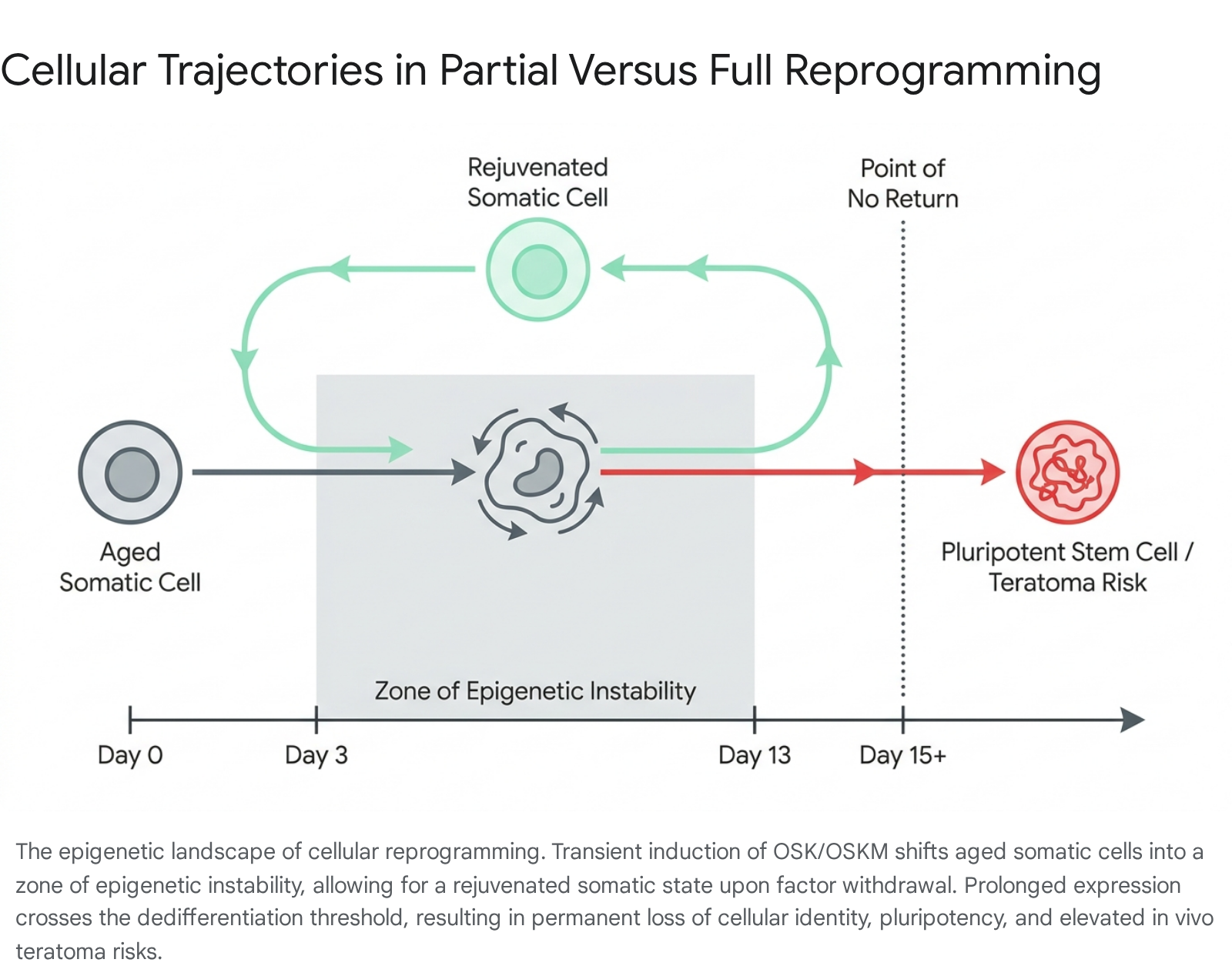
If OSKM expression is sustained beyond 15 days, the cells traverse a maturation phase where the molecular memory of their original identity is permanently erased, resulting in the acquisition of pluripotency 11. In vivo, cells that successfully dedifferentiate without proper control mechanisms lead to dysplastic cell proliferation and the formation of teratomas 112.
Studies utilizing single-cell RNA sequencing in pancreatic tissue undergoing OSKM reprogramming have mapped this trajectory precisely. Upon OSKM activation, acinar cells upregulate markers such as Reg3a, entering a pre-reprogramming state 13. While some cells remain refractory, others lose their acinar identity and acquire distinct gene expression profiles associated with intermediate dedifferentiation, marked by proteins like LY6A 13. Ultimately, a fraction of these cells reaches a fully pluripotent state, characterized by the endogenous expression of Nanog and Oct4 13. Identifying these intermediate markers allows researchers to visualize the precise state of dedifferentiation in situ, ensuring that therapeutic applications do not push tissues past the threshold of safety 13.
Mesenchymal Drift and Tissue Fibrosis
Recent transcriptomic analyses across human and murine tissues have identified "mesenchymal drift" as a unifying mechanism of biological aging 2214. Mesenchymal drift describes the progressive acquisition of mesenchymal traits - such as increased cellular motility, extracellular matrix (ECM) deposition, and a loss of cellular adhesion - by non-mesenchymal epithelial and endothelial cells 2214. This process erodes strict lineage identity, promotes chronic inflammation, stiffens the tissue microenvironment, and drives systemic fibrosis 2214.
In vivo partial reprogramming via OSKM has demonstrated a profound ability to inhibit and reverse mesenchymal drift 2214. Through a partial mesenchymal-epithelial transition (MET), transient OSKM induction returns cells to a more youthful epithelial state, reducing fibrotic signatures and restoring organ integrity 2214. In aged mouse models, long-term cyclic OSKM induction (over seven months) resulted in a highly significant decrease in mesenchymal drift gene expression in the kidney and liver, and a broad trend toward restoration across most other solid organs 14.
Modulation of Cellular Senescence
Cellular senescence is defined by an irreversible arrest of the cell cycle accompanied by the senescence-associated secretory phenotype (SASP), a dynamic secretome that drives chronic, low-grade tissue inflammation (inflammaging) 2415. Senescent cells present a cell-intrinsic barrier to tissue regeneration and accumulate progressively with age 24.
Partial reprogramming exerts significant senomorphic effects. Unlike senolytics, which seek to induce apoptosis and physically clear senescent cells from the body, senomorphic interventions modulate the behavior of these cells 1416. Evidence indicates that both genetic and chemical partial reprogramming directly target senescent cells, markedly attenuating their pro-inflammatory secretory activity, restoring mitochondrial homeostasis, and reducing apoptotic priming without restoring their proliferative capacity 1216. For example, the targeted expression of OSK driven by the Cdkn2a promoter (a marker for senescence) in aged wild-type mice preferentially rejuvenated senescent cells, reduced pro-inflammatory cytokine expression, improved wound healing, and extended lifespan without altering tumor incidence 16.
However, the interaction between reprogramming and senescence is highly complex and heavily dependent on the tissue microenvironment. Certain SASP components, such as Amphiregulin (AREG), have been found to facilitate cellular plasticity and actively enhance in vitro and in vivo reprogramming efficiency in a paracrine manner via epidermal growth factor receptor (EGFR) signaling 24. This suggests that while chronic senescence is detrimental, acute, stress-induced senescence and specific secretory factors may serve as essential components of normal tissue repair and cellular plasticity 2417.
Yamanaka Factors and Genetic Delivery Mechanisms
The transition of partial reprogramming from theoretical biology to an actionable therapeutic strategy is heavily constrained by the specific transcription factors utilized and the vectors engineered to deliver them. The classical Yamanaka cocktail includes the oncogene c-Myc, which introduces severe translational risks.
The OSKM Paradigm and Oncogenic Risk
The original OSKM configuration (Oct4, Sox2, Klf4, and c-Myc) is highly efficient at rapidly remodeling chromatin and erasing epigenetic aging marks 312. However, c-Myc is a potent driver of cellular proliferation and is strongly linked to cellular dysplasia and tumor formation 51218. Research indicates that c-Myc overexpression induces dysplasia similar to that found in squamous cell carcinomas and is overexpressed in up to 70% of breast cancer specimens 18.
In models utilizing full OSKM expression in vivo, the therapeutic window is extremely narrow. Continuous expression rapidly results in weight loss, gastrointestinal abnormalities, liver failure, and the widespread development of teratomas, ultimately leading to the premature death of the animal 549. Even brief or low-level induction can inadvertently push highly plastic cells across the dedifferentiation threshold, making OSKM largely unsuitable for broad systemic human therapeutics without sophisticated genetic safeguards 19.
The OSK Configuration
To mitigate oncogenic risk, the majority of translational efforts have dropped c-Myc entirely, relying on the three-factor OSK (Oct4, Sox2, Klf4) configuration 55. OSK aims to achieve epigenetic resetting while avoiding full dedifferentiation dynamics and uncontrolled proliferation 5.
While OSK significantly reduces the probability of tumor formation, it is inherently less efficient than OSKM, generally requiring longer exposure durations to achieve a comparable degree of epigenetic rejuvenation 3. Despite this reduced efficiency, the safety profile of OSK makes it the preferred candidate for clinical development. Long-term, continuous expression of OSK in specific isolated tissues, such as the retinal ganglion cells of mice, has been maintained for over 20 months without causing detectable tumor formation, retinal structural abnormalities, or loss of cellular identity 2.
Viral and Nucleic Acid Delivery Modalities
Initial in vivo proofs-of-concept utilized transgenic mouse models (e.g., LAKI progeroid mice carrying a doxycycline-inducible polycistronic OSKM cassette) 48. Because this relies on organism-wide germline genetic modification, clinical translation requires the development of exogenously delivered vectors capable of penetrating adult somatic tissues.
Adeno-associated viruses (AAVs) have been the primary vehicle for systemic gene therapy delivery. Research conducted by Rejuvenate Bio demonstrated that systemically delivered AAVs encoding an inducible OSK system in 124-week-old wild-type mice (equivalent to approximately 77 human years) extended the median remaining lifespan of the subjects by 109% and significantly enhanced overall health parameters 142021. However, systemic AAV delivery yields highly heterogeneous expression; some tissues receive excessive induction while others receive too little 19. Organs with naturally high plasticity and rapid cell turnover, such as the liver and intestines, are uniquely vulnerable to fatal dysplasia if the factors are overexpressed, complicating the systemic administration of AAVs 519.
To bypass the risks of permanent genomic integration and improve temporal control, transient delivery systems utilizing mRNA, lipid nanoparticles, and non-integrating viral vectors are under active investigation 322. For example, the European Horizon-funded REJUVIMMUNE project utilizes mRNA therapeutics to target hematopoietic stem cells (HSCs), aiming to mitigate age-associated immune decline and rejuvenate the blood system without altering the host genome 23. Similarly, Sendai virus vectors - which replicate in the cytoplasm without integrating into the host DNA - have been successfully utilized by researchers at the Chinese Academy of Sciences to reverse replicative senescence in cultured human endometrial mesenchymal stem cells, restoring youthful morphology and activating DNA damage response mechanisms 2435.
| Characteristic | OSKM (Oct4, Sox2, Klf4, c-Myc) | OSK (Oct4, Sox2, Klf4) | Single-Gene Targets (e.g., SB000) |
|---|---|---|---|
| Reprogramming Efficiency | High; rapidly remodels chromatin structure 3. | Moderate; requires extended exposure for full epigenetic reset 3. | High; claimed efficacy on par with full OSKM in vitro 515. |
| Pluripotency Induction | Rapidly induces pluripotency and permanent loss of somatic identity 5. | Capable of forming pluripotent colonies in vitro, but at a slower rate 5. | No evidence of pluripotency activation or loss of cellular identity 2537. |
| Oncogenic & Teratoma Risk | High; c-Myc strongly drives cellular dysplasia and tumor formation 51218. | Significantly reduced; bypasses c-Myc driven hyperproliferation 538. | Theoretically negligible; bypasses standard Yamanaka stem-cell pathways 2526. |
| Clinical Translation Status | Heavily restricted due to systemic toxicity and strict safety constraints 5. | Advancing to Phase 1 human trials for isolated organ targets 1327. | Preclinical; requires robust in vivo validation of healthspan and safety 2628. |
Chemical and Alternative Reprogramming Modalities
The technical challenges of viral delivery, coupled with the inherent stochasticity and inefficiency of transcription factor-mediated reprogramming, have driven the development of alternative modalities. Chemical reprogramming replaces transgenic elements with small-molecule cocktails that modulate specific epigenetic modifiers and cellular signaling pathways 192930.
Small Molecule Cocktails
Chemical reprogramming circumvents the difficulties of viral vector delivery and permanent genomic alteration, offering scalable manufacturing, straightforward dosing adjustments, and easier withdrawal protocols to prevent hyperproliferation 192930. Human chemical reprogramming was definitively achieved in 2022, building upon a decade of research into murine chemical induction 29.
Significant preclinical breakthroughs in 2024 and 2025 identified defined chemical regimens capable of inducing multiparametric rejuvenation in mammalian models 143132. A seven-compound cocktail (7c) was shown to lower the transcriptomic and epigenetic age of both human and murine fibroblasts without inducing dedifferentiation 3133. Treatment with 7c significantly altered mitochondrial interaction networks, increased the size of mitochondria, sped up movement velocity, and enhanced oxidative phosphorylation while safely preserving fibroblast somatic identity 3133.
Further refinement of this approach yielded a reduced two-compound (2c) cocktail - comprising the small molecules tranylcypromine and RepSox - that demonstrated comparable epigenetic rejuvenation efficacy in vitro 1632. The 2c cocktail successfully ameliorated cellular senescence, restored heterochromatin, and reduced genomic instability and oxidative stress 32. Crucially, the in vivo application of this 2c combination significantly extended the median lifespan and healthspan of C. elegans by over 42%, serving as a robust proof-of-concept for non-genetic organismal rejuvenation 1632.
Efficacy and Mechanism of Chemical Induction
The molecular mechanisms underlying chemical reprogramming differ in subtle but critical ways from traditional OSKM induction. While OSKM suppresses the p53 tumor-suppressor pathway during the transition toward dedifferentiation, partial chemical reprogramming using the 7c cocktail actually upregulates p53-related networks 12. This upregulation closely mimics normal physiological states and provides a crucial safety mechanism, potentially offering a vastly superior oncogenic profile compared to transcription factors 12.
However, chemical reprogramming is not without translational challenges. The broad biological activity of small molecules introduces concerns regarding off-target effects, as many compounds simultaneously affect multiple interconnected pathways 19. Achieving precise spatiotemporal control remains technically demanding, and further in vivo validation across diverse mammalian tissues is required to confirm that systemic chemical administration does not induce unforeseen toxicities 19.
Biomarkers and Epigenetic Clocks
The clinical viability of any rejuvenation therapy relies absolutely on the ability to quantify "biological age" accurately and differentiate it from simple chronological time. If a therapy purports to reverse aging, researchers must possess highly sensitive, universally accepted metrics to prove it.
DNA Methylation Clocks
Historically, biological age assessment has been dominated by DNA methylation (DNAm) "epigenetic clocks" 103435. First-generation models, such as the Horvath pan-tissue clock and the Hannum clock (both published in 2013), utilized machine learning algorithms to correlate the methylation status of specific CpG dinucleotides across the genome with chronological age 104936. These first-generation clocks proved highly accurate at predicting chronological age across virtually all tissues and cell types 10.
Subsequent second-generation and third-generation clocks, including PhenoAge, GrimAge, and DunedinPACE, improved upon this foundation by training their algorithms not just on chronological time, but on health-related variables, physiological functional measures, and mortality risk 4937. These advanced models effectively measure the "pace of aging," demonstrating that individuals with accelerated epigenetic clocks face significantly higher risks of cancer, non-AIDS severe events, and all-cause mortality 4937.
Limitations of First-Generation Epigenetic Clocks
However, the rapid advancement of partial reprogramming therapeutics has exposed severe conceptual and technical limitations in standard DNAm clocks. Recent critical reviews from leading institutions, including the Max Planck Institute and Harvard Medical School (e.g., Vadim Gladyshev), emphasize that first-generation clocks inherently conflate the accumulation of stochastic molecular damage with beneficial, adaptive physiological changes 3738.
Because DNAm clocks are trained on correlative data, an intervention (such as a specific reprogramming factor) may superficially "turn back the clock" by altering the methylation status of a handful of heavily weighted CpG sites without actually generating true physiological rejuvenation 1539. This phenomenon, termed the decoupling of epigenetic age from biological function, raises concerns that researchers may mistake localized methylation changes for wholesale cellular rejuvenation 39. Furthermore, standard DNAm assays based on bisulfite conversion are hampered by high costs, low clinical scalability, and an inability to reliably distinguish between 5-methylcytosine (5mC) and 5-hydroxymethylcytosine (5hmC) without the use of advanced, low-throughput TAB-array platforms 3549.
Transcriptomic and AI-Driven Temporal Models
To resolve the limitations of DNA methylation, the field is rapidly shifting toward single-cell transcriptomic clocks and high-dimensional artificial intelligence models. Transcriptomic clocks assess the actual RNA expression profiles of cells, providing a direct measurement of cellular function and identity rather than relying on proxy methylation markers 515.
For instance, UK-based Shift Bioscience developed "AC3," a proprietary aging clock based entirely on single-cell RNA sequencing and trained on a wide donor pool aged from 1 to 87 515. This platform allows researchers to measure age reversal across distinct cellular lineages while simultaneously tracking genes that govern somatic identity, ensuring that rejuvenation does not mask silent, dangerous dedifferentiation 515.
Similarly, researchers at the RIKEN Center for Integrative Medical Sciences in Japan recently introduced "MaxToki," a temporal AI model trained on nearly one trillion gene tokens across human lifespans 35. MaxToki utilizes advanced in-context learning to predict non-monotonic gene activation during partial reprogramming trajectories 35. By recognizing complex, non-linear transcriptomic changes that traditional regression models miss, MaxToki successfully identified novel pro-aging and rejuvenating perturbations, which were subsequently validated in vivo to influence age-related cardiovascular decline 35.
| Epigenetic Clock Generation | Primary Data Source | Training Variable | Key Characteristics and Limitations |
|---|---|---|---|
| First-Generation | DNA Methylation (CpG) | Chronological Age | High predictive accuracy for time alive (Horvath, Hannum). Limited physiological insight; conflates damage with adaptation 4937. |
| Second/Third-Generation | DNA Methylation + Biomarkers | Mortality / Morbidity | Measures pace of aging (PhenoAge, GrimAge, DunedinPACE). High throughput sequencing required; clinically expensive 4937. |
| Transcriptomic (AI-Driven) | Single-Cell RNA Sequencing | Multi-omic Gene Expression | Direct functional measurement (e.g., AC3, MaxToki). Ensures cell identity preservation; highly computationally intensive 1535. |
Institutional Research and Academic Milestones
The current wave of commercial longevity biotechnology is anchored by decades of academic research across global institutions. As the biological mechanisms of reprogramming are elucidated, a collaborative yet highly competitive international landscape has emerged.
The Salk Institute and Altos Labs Framework
The foundational architecture for modern partial reprogramming was constructed at the Salk Institute by Juan Carlos Izpisua Belmonte and colleagues. Their landmark 2016 study demonstrated that cyclic OSKM induction could safely extend the lifespan of LAKI progeroid mice and improve tissue regeneration 8.
This academic foundation was rapidly commercialized with the launch of Altos Labs, which secured an unprecedented $3 billion in initial funding 1254. Under Belmonte's guidance, Altos Labs operationalizes a highly rigorous paradigm focused strictly on reversing cellular disease states and restoring tissue resilience through epigenetic resetting, rather than explicitly pursuing life extension as an endpoint 1112. In recent preclinical models, the Altos framework has demonstrated that Topoisomerase-2-mediated partial reprogramming can induce profound cellular plasticity, counteract liver failure, and significantly enhance the regenerative capacity of skeletal muscle, the optic nerve, and the cardiovascular system 11. However, recognizing the severe risks of cellular dysplasia, the company maintains a highly cautious translational timeline, prioritizing the deep mapping of epigenetic clocks over rapid clinical entry 51234.
European and Asian Academic Initiatives
Beyond the United States, significant public and academic resources are dedicated to uncoupling the complex biology of aging. * Max Planck Institute for Biology of Ageing (Germany): Researchers at the Max Planck Institute, including Linda Partridge, focus heavily on the intersection of genetics, multi-target interventions, and environmental factors using model organisms like C. elegans and Drosophila 3840. Recent studies have demonstrated that combinatorial approaches, such as pairing rapamycin with trametinib, extend lifespan by 30% while reducing tumorigenesis, highlighting the institute's emphasis on multi-pathway modulation 2238. * Karolinska Institutet (Sweden): Supported by the StratRegen initiative, researchers at Karolinska are advancing the spatial profiling of tissue regeneration. Projects include Marco Gerling's work on modulating liver regeneration to restrain aggressive tumor growth via CRISPR-based approaches, and Emma Andersson's development of single-cell barcode-based lineage tracing for mesenchymal cells, which directly impacts the study of age-related fibrosis 4142. * Chinese Academy of Sciences & Tsinghua University (China): Chinese researchers are pushing the boundaries of in vivo tracking and targeted delivery. Bin Zhou's team at the Chinese Academy of Sciences developed the p16-tdT reporter mouse line, utilizing a dual homologous recombinase system to precisely trace cellular senescence in vivo and directly observe how senescent cells drive liver fibrosis 43. Additionally, researchers at Tsinghua University have developed novel optogenetic approaches using LOV2-inserted Acr proteins to achieve precise light-controlled in vivo cellular reprogramming 44. * RIKEN (Japan): The RIKEN Center for Integrative Medical Sciences has pioneered novel methods for generating functional neurons without passing through an iPSC intermediate state. By combining the neuronal transcription factor NEUROD1 with the Yamanaka factors (OCT3/4, SOX2, KLF4, and c-MYC), researchers successfully converted peripheral blood cells into glutamatergic neurons within three weeks, providing a highly efficient, minimally invasive model for studying neurodegenerative decline 6045.
Corporate Translation and Clinical Pipelines
The transition from academic theory to clinical reality is accelerating, driven by biotech startups focusing on specific, compartmentalized diseases of aging to satisfy regulatory requirements.
Ophthalmic Indications and Life Biosciences
The eye represents the most viable initial target for reprogramming therapeutics. It is an isolated, immune-privileged compartment, allowing for localized injection with minimal risk of systemic distribution to highly plastic organs like the liver 538. Furthermore, ophthalmic trials offer clear, non-invasive functional endpoints, such as electroretinogram responses.
In 2026, the U.S. Food and Drug Administration (FDA) approved Life Biosciences' Investigational New Drug (IND) application for ER-100, marking the first human clinical trial of a targeted cellular rejuvenation approach using partial reprogramming 1338. ER-100 is an AAV-delivered gene therapy utilizing the safer three-factor OSK configuration 3827. In rigorous preclinical non-human primate (NHP) models of non-arteritic anterior ischemic optic neuropathy (NAION), a single intravitreal injection of ER-100 combined with systemic doxycycline successfully restored youthful methylation patterns in retinal ganglion cells 1327. The therapy rescued axon density and significantly improved visually evoked functional responses without triggering any systemic toxicities or tumor formation 3827. The ensuing Phase 1 human trial, targeting NAION and glaucoma, represents a watershed moment for the translation of longevity science into mainstream medicine 1738.
Neurological and Cognitive Interventions
Age-related cognitive decline and neurodegeneration represent the next frontier. Preclinical investigations have repeatedly demonstrated that transient OSK/OSKM expression in the brains of aged rodents mitigates memory deficits, increases dendritic spine density, favors the migration of neural stem cells, and noticeably reduces amyloid-beta plaques in Alzheimer's disease models 10.
Building on these robust preclinical findings, Seattle-based YouthBio Therapeutics received positive FDA INTERACT feedback in late 2025 for YB002, a first-in-class, brain-specific Alzheimer's gene therapy utilizing partial reprogramming 10. The company is actively preparing to enter the clinic, positioning YB002 as a potential disease-modifying treatment that targets the root epigenetic degradation underlying neurodegeneration, rather than merely managing amyloid symptoms 10.
Hematopoietic and Autophagy Interventions
Age-associated immune decline is primarily driven by the deterioration of hematopoietic stem cells (HSCs), which undergo a myeloid bias and lose their regenerative capacity, leading to increased susceptibility to infections and a failure to clear senescent cells 23.
Retro Biosciences, backed by an initial $180 million investment from Sam Altman, has expanded its approach beyond transcription factor reprogramming to encompass cellular replacement and autophagy enhancement 4663. Retro achieved clinical-stage status in late 2025 by dosing the first participant in a Phase 1 trial in Australia for RTR242, a small-molecule therapy designed to restore lysosomal function and reactivate cellular autophagy 4664. By clearing toxic protein aggregates, RTR242 is targeted primarily at Alzheimer's disease 64. Concurrently, Retro is advancing preclinical programs (RTR890) to extract, reprogram, and replace old blood stem cells and brain microglia with younger counterparts to directly combat systemic immune dysfunction and leukemic disorders 4663.
Artificial Intelligence in Target Discovery
To bypass the inherent risks of standard Yamanaka factors, several companies are deploying advanced artificial intelligence to discover novel, safer reprogramming targets.
- Shift Bioscience: UK-based Shift Bioscience utilized its proprietary AI-powered single-cell transcriptomic aging clock (AC3) to screen a massive library of 1,500 candidate genes 515. In 2025, the company identified "SB000," a novel single-gene target capable of driving multi-omic rejuvenation in human fibroblasts and keratinocytes 515. Preclinical data indicates that SB000 matches or exceeds the epigenetic clock reversal achieved by OSKM (reducing transcriptomic age by up to 13.6 years in skin cells) but does not activate pluripotency pathways 252628. This allows the cells to preserve their original somatic identity and functions, such as collagen secretion, presenting a theoretically safer alternative to OSK 3726.
- Clock.bio: Based at the University of Cambridge, Clock.bio utilized human induced pluripotent stem cells (hiPSCs) in a proprietary force-aged "aging in a dish" model 6566. Conducting a genome-wide CRISPR screen on 3.4 million cells, the company generated a massive 20-terabyte single-cell RNA-sequencing dataset 666747. From this, Clock.bio successfully mapped an "Atlas of Rejuvenation Factors" comprising 151 genetic regulators linked to autophagy, epigenetic remodeling, and telomere restoration 4870. Backed by $5.3 million in seed funding, the company is leveraging its AI-powered "imAgeScore" platform to advance these targets toward clinical translation for skin rejuvenation and neurodegeneration 6548.
| Company | Core Technology / Target | Funding / Backing | Preclinical/Clinical Status (2026) |
|---|---|---|---|
| Altos Labs | Cyclic OSK/OSKM Reprogramming | $3 Billion | Extensive preclinical (in vivo mouse models, organ plasticity) 1254 |
| Retro Biosciences | RTR242 (Autophagy), GPT-4b micro | $180 Million+ (Sam Altman) | Phase 1 Clinical Trial (Alzheimer's / Autophagy) 4664 |
| Life Biosciences | ER-100 (AAV-delivered OSK) | Venture Capital | FDA IND Approved; Phase 1 Trial for Optic Neuropathies 1327 |
| Shift Bioscience | SB000 (Single-Gene Target), AC3 Clock | Innovate UK / VC | Preclinical (In vitro multi-omic rejuvenation without pluripotency) 525 |
| Clock.bio | Atlas of Rejuvenation Factors (151 genes) | $5.3 Million (Seed) | Preclinical / Target Validation (imAgeScore platform) 6548 |
| YouthBio Therapeutics | YB002 (Brain-specific gene therapy) | Venture Capital | FDA INTERACT clearance; Preclinical preparation for Alzheimer's 10 |
Safety Constraints and Translational Challenges
Despite the immense therapeutic promise and the initiation of early-stage human trials, in vivo partial reprogramming remains constrained by severe physiological safety risks.
Tissue Heterogeneity and Dysplasia
The primary obstacle to organism-wide systemic reprogramming is the uneven distribution and variable response of different tissues to viral gene therapies. Systemic delivery of doxycycline-inducible OSKM yields unequal induction; certain tissues receive sub-therapeutic doses while others receive toxic levels 19. Organs with naturally high endogenous plasticity and rapid cellular turnover, such as the liver and the intestines, are uniquely vulnerable 519. Even brief over-induction in these organs can cause cells to cross the threshold into dedifferentiation, leading directly to fatal dysplasia and the loss of organ integrity 54.
Furthermore, the tissue microenvironment severely complicates this delicate balance. Chronic inflammatory signals - often present in aged or diseased tissues - can abnormally sensitize local cells to the reprogramming factors 19. This stochastic response makes it exceedingly difficult to predict exactly how a specific organ will react to a standardized systemic dose of OSK or chemical modulators, raising significant concerns for human clinical translation where inter-patient variability is high 19.
The Narrow Therapeutic Window
Ultimately, in vivo reprogramming reflects an intrinsic biological trade-off between regenerative plasticity and pathological risk 19. The transient relaxation of cell identity and proliferative constraints successfully enhances tissue repair, reverses the epigenetic clock, and clears senescent phenotypes in permissive contexts 1619. Yet, this exact same plasticity drives tumorigenesis and teratoma formation when genetic safeguards fail or the exposure duration slightly exceeds the targeted window 1219.
The success of the field over the coming decade relies on the ability to uncouple these dual outcomes. Whether through the targeted, localized delivery of AAVs to immune-privileged sites like the eye, the identification of safer single-gene targets via artificial intelligence, or the perfection of non-transgenic chemical cocktails, mastering the spatiotemporal control of the epigenome remains the ultimate goal of longevity biotechnology.