AI in Drug Discovery and Clinical Translation 2023 - 2026
1. Executive Summary: The Maturation of Computational Therapeutics
The integration of artificial intelligence (AI) and machine learning (ML) within the global biopharmaceutical sector has crossed a critical threshold. Between late 2023 and mid-2026, the industry definitively transitioned from an era of speculative, isolated algorithmic piloting into a mature, production-driven operating model 1. Propelled by the looming acceleration of patent cliffs - which threaten over $180 billion in United States branded drug revenues by 2030 - pharmaceutical conglomerates and agile biotechnology startups alike have positioned AI as foundational research and development (R&D) infrastructure rather than an experimental adjunct 23. The global AI drug discovery market, valued at approximately $6.93 billion in 2025, is projected to expand significantly as computational methods reliably compress early-stage development timelines. Historically, the journey from target identification to Phase I clinical trials required four to five years; contemporary AI-native pipelines are achieving these milestones in roughly 18 to 30 months 345.
However, the empirical reality of the 2025 - 2026 landscape is defined by a sharp dichotomy: unprecedented upstream acceleration juxtaposed against stubborn downstream clinical attrition. While computational candidate generation has drastically improved Phase I safety success rates - pushing them from a historical average of 52% to between 80% and 90% for AI-derived molecules - Phase II and Phase III efficacy trials remain a formidable barrier 56. As of May 2026, no drug entirely discovered and designed by an end-to-end AI platform has achieved full regulatory approval from the U.S. Food and Drug Administration (FDA) or the European Medicines Agency (EMA) 357.
This comprehensive report evaluates the empirical advancements in AI-driven drug discovery, synthesizing recent evidence published in high-tier sources such as Nature Biotechnology, Cell, Bioinformatics, and Nature Reviews Drug Discovery, alongside industry reporting from Fierce Biotech and Endpoints News. It explicitly delineates the epistemological distinction between AI-identified targets and AI-designed molecules, maps the evolution of specific ML architectures (including Graph Neural Networks, Diffusion models, and Large Language Models), and catalogs the clinical progression and strategic delays of high-profile assets. Furthermore, this analysis addresses the critical "garbage in, garbage out" data bottlenecks through the lens of federated learning solutions, evaluates the impact of newly harmonized FDA and EMA regulatory frameworks, and highlights the ascendance of geographically diverse ecosystems, particularly the shifting paradigms within Asian biotechnology hubs.
2. Epistemological Corrections: AI-Identified Targets vs. AI-Designed Molecules
A pervasive and persistent misconception in both investor discourse and broader scientific reporting is the conflation of "AI-identified targets" with "AI-designed molecules." Rigorous empirical evaluation of the biopharmaceutical landscape requires a strict epistemological separation of these two distinct computational applications, as they carry fundamentally different biological risks, require different mathematical models, and address separate phases of the scientific method 429.
2.1 The Target Identification Challenge: Establishing Biological Causality
Target identification represents the most complex, high-risk stage of the drug discovery pipeline. It requires determining precisely which biological molecule - typically a specific protein, enzyme, or nucleic acid sequence - is the causal driver of a disease state, and whether modulating that target will yield a therapeutic benefit without inducing unacceptable systemic toxicity. The scale of this biological challenge is immense. The human genome contains approximately 20,000 protein-coding genes, yet scientists estimate that only about 4,500 possess the structural attributes necessary to be considered "druggable." More notably, all historically approved pharmaceutical drugs act on a mere 716 distinct biological targets 311.
Utilizing AI to discover a truly novel target means navigating chaotic, highly dimensional human biology. As detailed in comprehensive reviews within Nature Reviews Drug Discovery, modern target identification algorithms must parse vast, multi-omics datasets encompassing genomics, transcriptomics, proteomics, and metabolomics, while simultaneously cross-referencing electronic health records, clinical trial databases, and high-content cellular imaging (such as Cell Painting) 9312. This is fundamentally a problem of inferring biological causality, not merely structural chemistry. A machine learning model must identify hidden disease-associated nodes within complex, incompletely mapped biological networks and formulate a testable therapeutic hypothesis 312. Discovering a novel target inherently carries massive translational risk, as the underlying biological mechanism has never been validated in human clinical practice.
2.2 The Molecular Design Challenge: Navigating Chemical Space
Conversely, using AI to design a molecule is a bounded exercise in chemical engineering and physics. In this scenario, the biological target and its disease mechanism are already well-understood and clinically validated. The algorithm's objective is to explore theoretical chemical space - estimated at an unfathomable $10^{60}$ possible small molecules - to engineer a specific chemical structure that binds optimally to the known target's active pocket 41415.
Generative AI models in this space focus on optimizing binding affinity, managing off-target selectivity, predicting pharmacokinetics, and ensuring the compound is synthetically accessible in a laboratory environment 415. While computationally intensive, designing a key for a known lock is inherently more solvable than discovering the lock itself.
2.3 Empirical Distinctions and the Non-Negotiable Requirement of the Wet Lab
The profound difference between these two modalities is best illustrated by the historical progression of the industry's flagship assets. For example, Exscientia's DSP-1181 was widely celebrated in 2020 as the industry's "first AI-designed molecule" to enter human clinical trials 4. However, the biological target of DSP-1181 was a well-established serotonin receptor (5-HT1A) for the treatment of obsessive-compulsive disorder 5. The AI was utilized solely to rapidly optimize the chemical structure, meaning the program bypassed the severe risks associated with novel target discovery 4.
In sharp contrast, Insilico Medicine's INS018_055 (Rentosertib) represents a much rarer, "end-to-end" dual-AI achievement. In this program, the company's biological AI engine, PandaOmics, first discovered a completely novel biological target for idiopathic pulmonary fibrosis (IPF) - specifically, Traf2- and Nck-interacting kinase (TNIK), which had not previously been implicated in fibrosis 43. Subsequently, a separate generative chemistry engine, Chemistry42, was deployed to design the specific small molecule to inhibit it 33. The success of INS018_055 serves as the current benchmark for validating holistic AI platforms capable of managing both biological causality and chemical design 47.
Despite the unprecedented acceleration provided by algorithmic hypothesis generation, the absolute requirement for physical wet-lab testing remains a cornerstone of the industry. Predictive algorithms frequently encounter the "activity cliff" phenomenon, wherein minor structural modifications to a chemical scaffold result in disproportionately large, unpredictable changes in biological activity that 2D computational descriptors fail to anticipate 2. AI platforms do not bypass the scientific method; they accelerate the formulation of hypotheses that must still undergo rigorous robotic synthesis, in vitro bioassays, and in vivo animal model validation. Recognizing this, leading AI firms are heavily investing in automated robotic facilities. Companies like XtalPi and Excelsior Sciences have integrated "machine-native chemistry" and automated synthesis loops directly with their generative models, reportedly raising chemical synthesis success rates from a traditional 20-30% up to 90%, thereby ensuring that dry-lab predictions remain strictly tethered to wet-lab physical realities 678.
3. Mapping Machine Learning Architectures to the Drug Discovery Pipeline
The underlying mathematical architectures powering computational drug discovery have evolved dramatically. The field has moved beyond the early, simplified deep neural networks of the past decade into an era dominated by highly sophisticated, multimodal foundation models. As highlighted in literature spanning Cell, Bioinformatics, and Nature Biotechnology, the current ecosystem relies predominantly on three primary architectural families: Graph Neural Networks (GNNs), Diffusion Models, and Large Language Models (LLMs) or Transformer-based architectures 41492110.
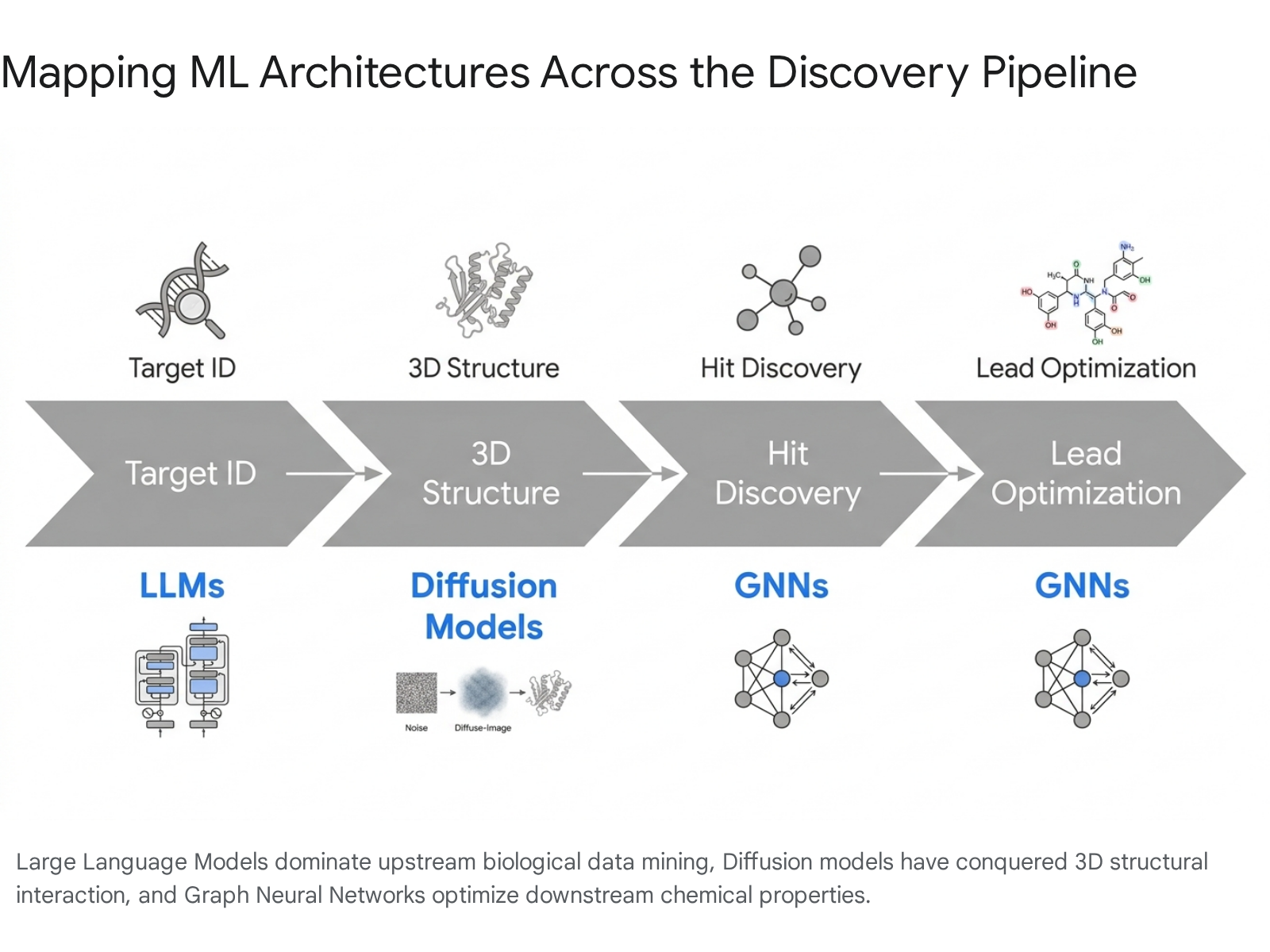
3.1 The Algorithmic Arsenal (2025 - 2026)
Graph Neural Networks (GNNs): GNNs natively operate on graph-structured data, making them mathematically optimized for representing molecular chemistry, where atoms function as nodes and chemical bonds function as edges 39. GNNs have established themselves as the gold standard for downstream chemical applications, particularly in predicting molecular properties (ADMET: Absorption, Distribution, Metabolism, Excretion, and Toxicity) 911. By accurately modeling molecular interactions and capturing hidden relationships between atomic structures, generative GNNs are instrumental in virtual screening, lead optimization, and predicting synthetic accessibility, thereby drastically reducing late-stage chemical failures 159.
Diffusion Models: Originally popularized in the domain of text-to-image generation, diffusion models have fundamentally revolutionized 3D structural biology. The watershed moment occurred with the release of AlphaFold 3 in 2024, co-developed by Google DeepMind and Isomorphic Labs. Moving away from the Evoformer architecture utilized in AlphaFold 2, AlphaFold 3 integrated a diffusion-based generative model capable of directly generating atomic coordinates 1012. This architectural pivot enabled the model to predict not merely static, single-chain proteins, but highly complex, dynamic multi-molecular assemblies. AlphaFold 3 can now model protein-ligand interactions, protein-DNA complexes, and protein-RNA assemblies with near-experimental accuracy 101225. In benchmark datasets, AlphaFold 3 achieved approximately 76% success in ligand binding predictions (RMSD <2 Å), significantly outperforming traditional physics-based docking methodologies and bringing the rational design of transcription factor modulators and CRISPR complexes into reach 1025.
Large Language Models (LLMs) and Transformers: Architectures that treat biological data as linguistic text have achieved massive scale. Following the broader convergence of LLM architectures between 2023 and 2025 - which standardized on pre-normalization (RMSNorm), Rotary Positional Embeddings (RoPE), SwiGLU activations, and Key-Value sharing attention mechanisms - biomolecular language models have proliferated 21. Models such as ESM-3 (Evolutionary Scale Modeling, developed by Meta AI) utilize transformer architectures to parse massive datasets of single-cell transcriptomes and evolutionary sequences 326. While ESM-3 may have a slightly lower structural accuracy ceiling compared to AlphaFold 3, it operates at a 60x faster prediction speed, making it the premier tool for ultra-large-scale proteome screening, high-throughput hypothesis generation, and capturing the evolutionary semantics of proteins 2526.
Table 1: Mapping Core ML Architectures to Drug Discovery Pipeline Stages
| Pipeline Stage | Primary ML Architecture | Leading Models / Frameworks | Application & Utility | Empirical Impact (2025 - 2026) |
|---|---|---|---|---|
| Target Identification & Validation | LLMs, Multi-modal Transformers, Knowledge Graphs | PandaOmics, ESM-3, EpistemicGPT | Mining literature, multi-omics data, and clinical records to find causal disease networks and novel biological nodes. | 58% of surveyed biopharma firms use AI for target ID. Reduced target nomination timelines from years to months 13. |
| 3D Structure Prediction & Complex Assembly | Diffusion Models | AlphaFold 3, Boltz-2, Chai-1 | Predicting the 3D structures of proteins and complex multi-molecular assemblies (protein-ligand, protein-nucleic acid). | AF3 achieves ~76% success in ligand binding predictions (RMSD <2 Å), significantly outperforming physics-based docking 1025. |
| Hit Discovery & Virtual Screening | GNNs, Equivariant Neural Networks | OpenBind, GATv2 | Ultra-large-scale virtual screening of billions of compounds against target pockets. | Enables computational screening of massive chemical spaces in days rather than deploying physical HTS campaigns 2427. |
| De Novo Molecular Design | Generative AI, VAEs, Biomolecular LLMs | Chemistry42, AIDDISON | Designing entirely novel chemical entities or peptide sequences optimized for specific pockets. | Accelerated generation of highly specific small molecules, though synthetic feasibility remains a bottleneck requiring automated synthesis loops 3628. |
| Lead Optimization & ADMET Prediction | GNNs, Active Learning | GNN-based property predictors | Predicting pharmacokinetics, toxicity, and synthetic accessibility to reduce late-stage failure. | Reduces the number of synthesis-test cycles. AI-optimized molecules show up to an 80-90% success rate clearing Phase I safety 259. |
4. Clinical Progression, Setbacks, and Strategic Reprioritizations (2024 - 2026)
The true measure of AI in drug discovery relies not on computational benchmarking, but on the ruthless evaluation of human clinical trials. By mid-2026, the biopharmaceutical industry began parsing the pivotal Phase II and Phase III readouts from the first historical cohort of AI-designed molecules 35. The data reveals a complex, bifurcated narrative. On one hand, generative chemistry has proven extraordinarily adept at engineering out toxicity, leading to Phase I safety success rates that dwarf historical norms 56. On the other hand, the leap from Phase II (proof-of-concept) to Phase III (pivotal efficacy) remains fraught with biological uncertainty.
Furthermore, comprehensive analyses of clinical trial landscapes highlight a shift in failure modalities. A landmark 2026 study published in Nature Reviews Drug Discovery, utilizing the EpistemicGPT knowledge graph to analyze 3,180 terminated Phase II and III trials over the past decade, revealed that late-stage terminations have doubled, rising from roughly 11% to 22% 29. Critically, this spike was driven predominantly by strategic and business factors - such as corporate portfolio reprioritization and mergers - rather than pure scientific or clinical efficacy failures 29. Parsing the difference between a technical biological failure and a strategic pipeline discontinuation is essential to accurately audit the performance of AI biotechs.
4.1 Flagship Successes and End-to-End Validation
Insilico Medicine (INS018_055): Insilico Medicine's INS018_055 (Rentosertib) remains the undisputed frontrunner and primary validation point for the AI drug discovery sector 4. As the first therapeutic to feature both an AI-discovered target and an AI-designed compound, its progression is closely monitored. In mid-2025, Insilico published highly anticipated Phase IIa clinical results in Nature Medicine. The multicenter, double-blind trial demonstrated that the TNIK inhibitor was not only safe and well-tolerated but also yielded a statistically significant, dose-dependent improvement in forced vital capacity (FVC) over 12 weeks in patients suffering from idiopathic pulmonary fibrosis (IPF) 3457. The economics of this achievement are equally notable: preclinical nomination was completed in approximately 18 months at a total discovery cost of roughly $6 million, an inversion of the traditional $100 - $200 million, multi-year preclinical paradigm 37. As of 2026, the asset is preparing for expansive Phase IIb/III evaluations 45.
4.2 Biological Disappointments and Complexity Limits
Recursion Pharmaceuticals (REC-994 and REC-4881): Recursion, a pioneer in applying AI to high-throughput phenotypic screening, has experienced the full spectrum of clinical outcomes. In May 2025, the company announced the discontinuation of REC-994, an asset advancing through Phase II trials for symptomatic cerebral cavernous malformation (CCM) 1331. While preliminary 12-month data demonstrated promising trends in MRI imaging - specifically a reduction in total lesion volume at the highest 400mg dose - the subsequent long-term extension phase failed to confirm these signals. Most critically, the anatomical imaging changes did not correspond to any meaningful improvements in patient-reported functional outcomes 1331. This failure underscores a persistent boundary condition in AI discovery: phenotypic or biomarker improvements modeled in silico or in vitro do not automatically translate to systemic, functional clinical efficacy in complex human populations.
Conversely, Recursion's oncology and rare disease pipelines have shown immense promise. In early 2026, the company reported strong Phase II efficacy for REC-4881 (a MEK1/2 inhibitor) in patients with familial adenomatous polyposis (FAP), a disease lacking approved pharmacotherapies. The trial achieved rapid clinical activity, with 75% of evaluable patients showing reductions in total polyp burden, deepening to a 53% median reduction at 25 weeks 14. Recursion subsequently initiated FDA engagement to define a registrational pathway for the asset 14. Supported by disciplined capital execution, Recursion reported $665.2 million in cash reserves in Q1 2026, extending its financial runway into 2028 14.
4.3 Strategic Terminations and the Realities of Big Pharma
Exscientia (EXS4318 and EXS21546): The pipeline of Exscientia clearly illustrates the intersection of clinical mechanics and macroeconomic reality. EXS4318, a potential first-in-class selective PKC-theta inhibitor designed via generative AI, demonstrated highly positive early Phase I safety results and was successfully in-licensed by Bristol Myers Squibb (BMS) 1535. However, in late 2024, BMS abruptly discontinued the program and returned the asset 35. This termination was not the result of clinical toxicity or a failure of the AI's design; rather, it was a casualty of a broader $1.5 billion internal corporate portfolio streamlining effort by BMS, classifying it as a strategic, rather than scientific, failure 35.
Separately, Exscientia internally wound down its own Phase I/II trial for EXS21546 (an A2A receptor antagonist for oncology) in late 2023. In this instance, AI-driven clinical modeling indicated that it would be inherently challenging for the compound to reach a suitable therapeutic index, constituting a genuine scientific termination that allowed the company to reallocate resources to higher-value oncology targets like their CDK7 and LSD1 programs 36.
Table 2: Clinical Progress, Indications, and Delays of High-Profile AI-Derived Assets (As of Mid-2026)
| Company | Asset | Therapeutic Target / Modality | Indication | Highest Phase (2025-2026 Status) | Key Developments, Readouts, and Strategic Delays |
|---|---|---|---|---|---|
| Insilico Medicine | INS018_055 (Rentosertib) | TNIK Inhibitor (Small Molecule) | Idiopathic Pulmonary Fibrosis (IPF) | Phase IIb/III prep | Landmark Success: Published positive Phase IIa data in Nature Medicine (2025) showing dose-dependent FVC improvement. First fully AI-discovered target & molecule to reach this stage 457. |
| Recursion Pharma | REC-4881 | MEK1/2 Inhibitor | Familial Adenomatous Polyposis (FAP) | Phase II | Advancing: Strong Phase II efficacy (43% median polyp reduction at 12 weeks, deepening to 53%). Initiating FDA engagement for a registrational pathway in 2H26 14. |
| Recursion Pharma | REC-994 | Superoxide Scavenger | Cerebral Cavernous Malformation | Discontinued (Phase II) | Scientific Failure: Terminated May 2025. While MRI initially showed lesion reduction, long-term extension data lacked functional clinical improvements 1331. |
| Exscientia | EXS4318 | PKC-theta Inhibitor | Immunology / Inflammation | Discontinued (Phase I) | Strategic Failure: BMS discontinued the program in late 2024 to achieve $1.5B in corporate cost savings, despite the asset displaying a positive early Phase I safety profile 1535. |
| Exscientia | DSP-1181 | Serotonin Receptor (5-HT1A) Agonist | Obsessive-Compulsive Disorder (OCD) | Phase I (Status Unclear) | Stalled: Celebrated in 2020 as the first AI-designed molecule in human trials, but has vanished from recent clinical updates without published efficacy data as of 2026 45. |
| Relay Therapeutics | RLY-2608 (zovegalisib) | Mutant-selective PI3Kα Inhibitor | HR+/HER2- Advanced Breast Cancer | Phase I/II | Advancing: Granted FDA Breakthrough Therapy designation. Demonstrated strong median PFS of 10.3 months. Pivotal triplet data and frontline Phase III plans expected in 2026 . |
| Absci | ABS-201 | Anti-PRLR Antibody | Alopecia / Endometriosis | Phase I/IIa | Advancing: Accelerated Phase 1/2a initiation for alopecia. Expanding clinical strategy to endometriosis, with a Phase II trial expected to initiate in Q4 2026 . |
| Verge Genomics | VRG50635 | PIKfyve Inhibitor | Amyotrophic Lateral Sclerosis (ALS) | Phase I/II | Advancing: Utilized the CONVERGE all-in-human multi-omics platform (bypassing traditional animal models for target ID) to transition from research to clinic in just four years 4142. |
5. The "Garbage In, Garbage Out" Paradigm: Solving Biological Data Bottlenecks
While algorithmic architectures such as diffusion models and biomolecular transformers have matured at a staggering pace, their real-world utility remains fundamentally tethered to, and bounded by, the quality of their underlying training data. The "garbage in, garbage out" (GIGO) principle is universally acknowledged as the single largest bottleneck hindering the translation of computational models into physical clinical success 1644. As noted in recent literature, modern drug discovery is not hampered by a lack of raw data, but by its inconsistent quality 45.
5.1 The Limitations and Biases of Public Datasets
Historically, AI models in drug discovery have been trained on sprawling public bioactivity databases, such as ChEMBL and PubChem, which aggregate data sourced from academic literature and clinical outcome registries 4445. However, these datasets harbor deep structural deficiencies that become magnified at scale. Academic literature is overwhelmingly biased toward positive results; the conventions of scientific publishing mean that failed synthesis routes, inactive chemical scaffolds, and highly toxic compound profiles are systematically underreported 44. A generative model trained predominantly on this positively biased data learns an unrealistically optimistic topology of chemical-biological space. It overstates the density of active scaffolds and routinely proposes molecules that ultimately fail during physical lab validation 44.
Furthermore, public biological data is rife with experimental noise. Variations in assay conditions across different academic laboratories, miscalibrated lab equipment, and differing reporting standards introduce severe batch effects. When AI models ingest this uncurated noise, they inevitably identify false correlations. Companies like Lead AI have published peer-reviewed research in journals such as Molecules, demonstrating that deploying "selective cleaning" methodologies to remove experimental batch effects from databases can dramatically enhance machine learning accuracy for target classes like MDM2 inhibitors 45.
5.2 The Transition to Proprietary Data and Federated Learning Solutions
To circumvent the limitations of public datasets, the biopharmaceutical sector is aggressively pursuing two distinct data strategies. The first involves building proprietary, closed-loop data generation factories. Organizations like Recursion and Verge Genomics have invested hundreds of millions of dollars into automated, high-throughput wet labs that generate massive, highly standardized datasets - such as human multi-omics and phenomics - specifically tailored to train their internal AI models, thereby creating an impenetrable intellectual property moat 4144.
The second, arguably more transformative strategy, is the adoption of Federated Learning. The reality of the pharmaceutical industry is that no single entity - not even the largest multinational conglomerates - possesses enough diverse, high-quality chemical-biological data to train truly universal foundational models 17. Federated learning solves this by allowing multiple institutions to collaboratively train a shared AI model without ever exchanging or centralizing raw, confidential proprietary data 1718.
In a federated architecture, proprietary molecular data never leaves the sponsor's secure environment. Instead, localized models are trained on-site within a hub-and-spoke network, and only the encrypted mathematical learnings - or model weights - are aggregated to fine-tune the central OpenFold3 model. The landmark MELLODDY project (Machine Learning Ledger Orchestration for Drug Discovery) provided the initial operational proof-of-concept for this approach. The project successfully demonstrated that 10 competing pharmaceutical companies, possessing an unprecedented aggregated dataset of over 2.6 billion confidential experimental activity points across 21 million molecules, could securely achieve predictive improvements on regression models without exposing their respective molecular libraries 18.
In late 2025 and early 2026, this concept evolved significantly with the formation of the AI Structural Biology (AISB) Network. Facilitated by the federated computing platform Apheris, major industry players - including Bristol Myers Squibb, Takeda, Astex Pharmaceuticals, AbbVie, and Johnson & Johnson - pooled metadata derived from several thousand experimentally determined protein - small molecule structures 1748. Working in collaboration with Columbia University, this highly confidential consortium is fine-tuning OpenFold3, an open-source reproduction of AlphaFold 3 4849. By leveraging a federated architecture, OpenFold3 accesses the richest, most diverse training dataset of proprietary protein-ligand interactions ever assembled, with the ultimate goal of achieving predictive precision on par with X-ray crystallography while rigidly protecting the trade secrets of all participants 174849.
6. Evolving Regulatory Frameworks: FDA and EMA 2026 Guidelines
As AI-designed drugs advanced rapidly from preclinical curiosities into Phase II and Phase III pivotal trials, global regulatory authorities were forced to rapidly adapt to a paradigm for which traditional frameworks were ill-equipped. A fragmented, inconsistent regulatory landscape posed a severe threat to global drug development timelines. In a watershed moment for transatlantic regulatory convergence, the U.S. Food and Drug Administration (FDA) and the European Medicines Agency (EMA) jointly published the "Guiding Principles of Good AI Practice in Drug Development" on January 14, 2026 501952.
While these ten principles are not legally binding statutes, they serve as the foundational expectations against which regulators will evaluate New Drug Applications (NDAs) and Biologics License Applications (BLAs) that feature AI-generated evidence. Crucially, they mark a definitive shift in regulatory philosophy: moving away from merely auditing the final clinical output of drug discovery toward actively regulating the computational process and training methodologies utilized to generate those outputs 5019.
6.1 Core Regulatory Principles for AI in Biopharma
The joint FDA/EMA framework mandates a shift toward transparent, risk-based AI deployment throughout the product lifecycle. The critical directives include: * Human-Centric by Design: The agencies dictate that AI must serve strictly as an assistive, advisory tool rather than an autonomous decision-maker. Absolute human oversight and accountability must be embedded throughout the pipeline, ensuring that AI-generated clinical trial designs, biomarker selections, or toxicity profiles are ultimately reviewed and validated by human clinical experts 5020. * Risk-Based Approach: The intensity of validation required for an AI model must be directly proportional to its potential impact on patient safety. For instance, an algorithm predicting early-stage molecular toxicity requires less rigorous regulatory scrutiny than an AI tool utilized to stratify human patients for a pivotal oncology trial 5052. * Data Governance and Explainability: Algorithms functioning as uninterpretable "black boxes" are increasingly unacceptable for regulatory-critical applications. Sponsors are now expected to maintain comprehensive documentation regarding data provenance, model limitations, and explicit efforts to mitigate bias within training datasets to ensure generalizability across diverse real-world patient populations 35021. * Clear Context of Use: Sponsors must explicitly define the boundaries within which an AI model operates. Applying an AI tool validated for extrapolating efficacy in adult populations to pediatric cohorts, without conducting extensive re-validation, constitutes a violation of the established context of use 5021.
The implementation of these rigorous principles sets a substantially higher barrier to entry for pure-play technology companies attempting to disrupt the pharmaceutical space. Regulatory bodies now demand deep mechanistic biological evidence alongside strict adherence to GxP (Good Clinical/Manufacturing Practice) compliance, effectively filtering out AI startups that possess advanced algorithms but lack robust clinical integration and translational medicine capabilities 652.
7. Geographical Diversity: The Ascendance of the Asian Ecosystem
The global center of gravity in AI-driven drug discovery is undergoing a profound structural realignment. While North America and Europe remain dominant in foundational model development and massive capital allocation, the Asian biotechnology ecosystem - particularly in China - has rapidly matured and scaled. By 2026, the Chinese pharmaceutical industry definitively transitioned from a historical era defined by domestic policy backing and generic drug manufacturing into an efficiency-focused, AI-native innovation hub 2223.
This geographical shift is characterized by intense R&D investments and the emergence of highly differentiated, specialized business models that cater to distinct segments of the global drug discovery value chain.
7.1 Divergent Business Models: XtalPi vs. Insilico Medicine
The maturity of the Asian ecosystem is clearly exemplified by the divergent, yet highly successful, commercial models of two regional heavyweights: XtalPi and Insilico Medicine.
The Infrastructure and PaaS Model (XtalPi): XtalPi has strategically positioned itself as the foundational infrastructure provider for AI-driven drug discovery, operating a Platform-as-a-Service (PaaS) model. By heavily integrating AI-driven molecular simulation algorithms with sprawling automated robotic laboratories in Zhangjiang Science City, XtalPi focuses on execution speed and physical validation 7823. Crucially, XtalPi's revenue is tied directly to research contracts and technology delivery rather than assuming the massive, binary clinical risk of human drug approvals. This low-risk, high-efficiency commercial model allowed XtalPi to report a remarkable 201.2% year-on-year revenue increase in 2025, reaching $111 million (802.6 million yuan), and making it the first profitable "AI for Science" company listed under Hong Kong's new Chapter 18C rules 7823.
The AI-Biotech Pipeline Model (Insilico Medicine): Conversely, Insilico Medicine operates an ambitious, high-risk, high-reward pipeline model. Utilizing its proprietary end-to-end generative platform, Pharma.AI, Insilico acts as a full-stack clinical biotechnology company. It absorbs massive internal R&D costs to advance wholly owned assets (such as INS018_055) entirely through clinical trials 723. While this strategy resulted in a reported adjusted net loss of $43.8 million in 2025 - with R&D spending equivalent to a staggering 145% of its annual revenue - it positions the company to capture the exponentially higher pharmaceutical upside if its assets successfully secure regulatory approval 723.
The robust technological foundation established in Asia has triggered a wave of strategic M&A and cross-border licensing. Multinational pharmaceutical companies are increasingly pivoting from basic fee-for-service agreements toward shared-risk, milestone-based co-development deals with Chinese AI platforms, leveraging these ecosystems to execute complex clinical trials and enhance development efficiency across vast Asian patient populations 22.
8. Conclusion
As the biopharmaceutical industry progresses through 2026, artificial intelligence has definitively ceased to be an optional technological overlay; it has evolved into the core operating system of modern drug discovery 124. The field has moved past the naive, early-decade assumption that algorithms could wholly bypass the chaotic complexities of human biology. Instead, the most successful applications tightly couple advanced mathematical architectures - such as Diffusion Models for structural complex assembly and GNNs for chemical property optimization - with rigorous, automated wet-lab validation and human oversight.
The clinical milestones achieved by candidates like INS018_055 provide empirical proof-of-concept that AI can both discover novel biological targets and design viable therapeutic molecules in a fraction of traditional timelines 47. However, the sobering failures and strategic terminations of other assets in late-stage trials reinforce the reality that clinical efficacy remains inextricably linked to underlying biological causality - a domain where computational models are still learning to extrapolate 2913.
Looking toward the remainder of the decade, competitive advantage in AI-driven drug discovery will no longer stem from the sheer novelty of the algorithm, as generative models rapidly commoditize and open-source alternatives proliferate. Instead, market supremacy will be dictated by access to high-quality, proprietary biological data, the successful implementation of secure federated learning networks, and the ability to seamlessly integrate computational predictions with the stringent, newly unified FDA and EMA regulatory frameworks. Organizations that master this triad of data superiority, flawless wet-lab execution, and regulatory compliance are poised to define the next generation of precision medicine.