Advances and limitations of epigenetic aging biomarkers
Introduction
The conceptualization of aging has undergone a profound paradigm shift over the past decade, transitioning from the view of an inevitable, unquantifiable decline to a measurable, and potentially modifiable, biological process. At the vanguard of this transformation are epigenetic clocks - multivariate machine learning algorithms that estimate biological age based on the methylation status of specific cytosine-phosphate-guanine (CpG) dinucleotides across the genome 122. By differentiating a subject's molecular biological age from their chronological age, these biomarkers have provided researchers with unprecedented predictive insights into age-related morbidities, healthspan, and all-cause mortality 123. Over the last decade, advancements in high-throughput microarray technologies have enabled gerontologists to sequence specific loci on the epigenome and track the velocity of human decline 46.
However, as the application of epigenetic clocks expands beyond the confines of observational epidemiology and enters the realms of interventional gerontology and the commercial direct-to-consumer (DTC) market, significant methodological and conceptual vulnerabilities have been exposed. Recent literature from 2023 through 2025 underscores a growing schism between the rigorous standards required for clinical diagnostics and the premature commercialization of these tools 58. Furthermore, as researchers attempt to utilize these clocks to validate anti-aging interventions, fundamental questions regarding dataset bias, technical noise, cellular heterogeneity, and the underlying biological mechanisms driving these methylation changes have come to the forefront 678. This report provides an exhaustive analysis of the contemporary epigenetic landscape, prioritizing recent advancements in single-cell resolution, principal component mathematical smoothing, and the critical evaluation of interventional and commercial applications.
The Biological Mechanisms of Epigenetic Aging: Stochastic Drift Versus Programmed Development
Before evaluating the algorithmic tools used to measure epigenetic aging, it is necessary to examine the underlying biological phenomena they capture. A persistent debate within molecular gerontology centers on whether age-related DNA methylation changes are the result of stochastic, entropic damage accumulation or if they represent the tail-end of a highly regulated, programmed developmental cascade. The exact contribution of each process determines how future anti-aging interventions must be designed: if aging is stochastic, interventions must focus on DNA repair, but if it is programmed, systemic cellular reprogramming is required 69.
Recent 2024 publications highlight this dichotomy with advanced computational modeling. Research by Meyer et al. posits that the accumulation of stochastic variation - often referred to as epigenetic drift - is mathematically sufficient to generate highly accurate aging clocks 9. By demonstrating that purely simulated data, assuming a "ground state at age zero" followed by random stochastic alterations over time, can yield predictive models indistinguishable from empirical first- and second-generation clocks, Meyer et al. argue that programmed aging theories may be unnecessary to explain the predictive efficacy of these biomarkers 9. Under this framework, the epigenome simply loses its regulatory fidelity over time due to the accumulation of random "noise," which machine learning algorithms synthesize into a measurable chronological signal.
Conversely, Tarkhov et al. advocate for a dual-nature model of epigenetic aging, arguing that it encompasses both stochastic elements and co-regulated, programmed changes 910.
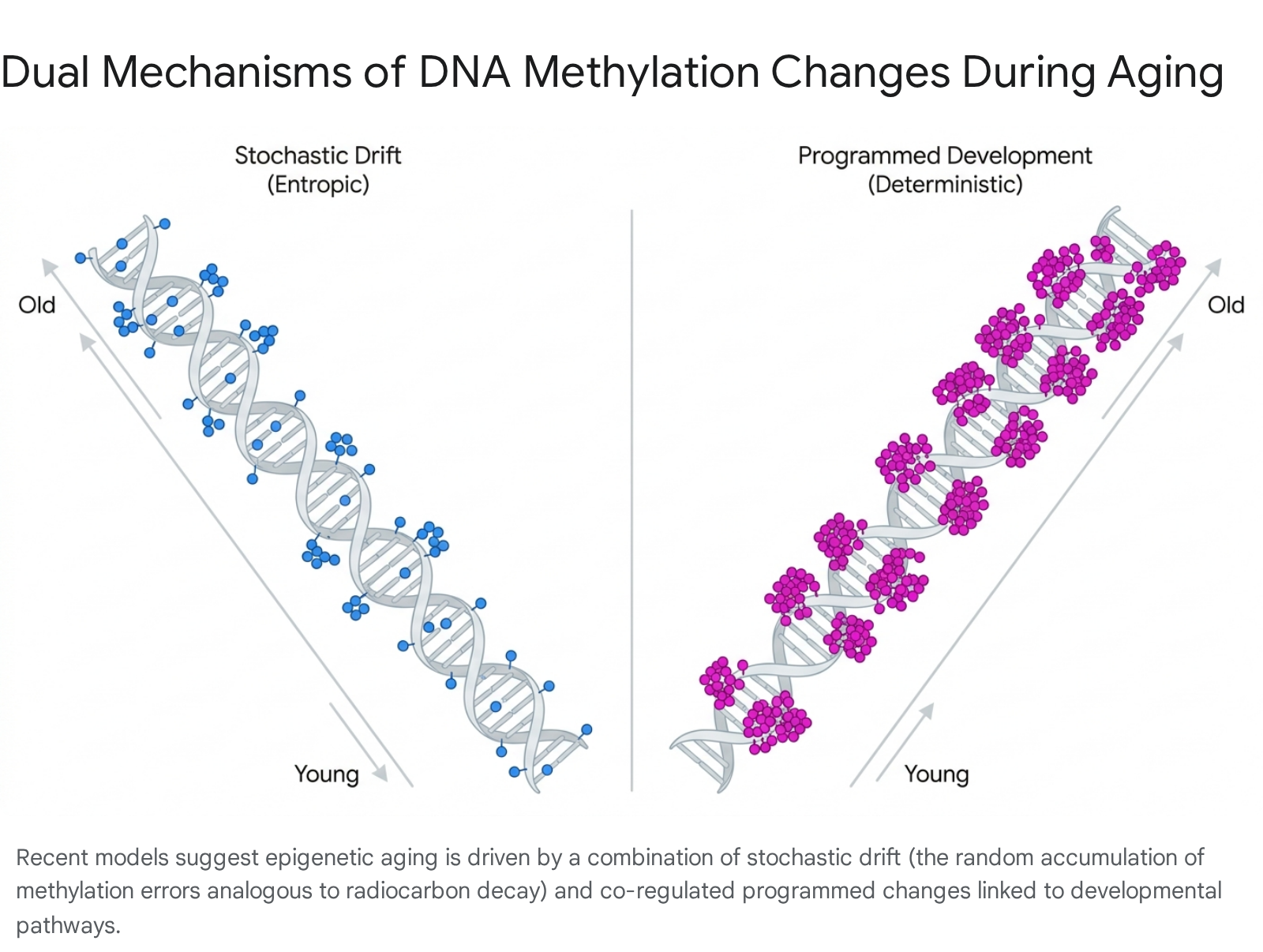
At single-cell resolution, Tarkhov et al. observed a simultaneous loss of regulation at both the epigenetic and transcriptional levels, which they modeled as a stochastic process akin to radiocarbon decay 9. However, their analysis also identified specific clusters of CpG sites that exhibited highly structured, co-regulated coordination patterns consistent with programmed developmental networks. This dual-nature perspective suggests that while stochastic damage accounts for a significant portion of the epigenome's broad degradation - specifically the general loss of constitutive heterochromatin - specific gene networks, particularly those involved in tissue differentiation and immune maturation, continue to follow a deterministic trajectory well into late life 911.
The Evolution and Taxonomy of Epigenetic Clocks
To accurately assess biological aging, researchers have developed successive generations of epigenetic clocks, each characterized by distinct training methodologies, CpG site selections, and primary clinical utilities. The transition through these generations reflects a conceptual shift from merely guessing a patient's chronological age to quantifying their physiological deterioration and mortality risk.
First-Generation Clocks: Chronological Age Predictors
The foundational first-generation algorithms, prominently featuring Horvath's Pan-Tissue clock and Hannum's Blood clock, were optimized strictly to predict chronological age using penalized regression methods, specifically elastic net regression 21115. Horvath's clock, published in 2013, represented a landmark achievement by utilizing 353 CpG sites - 193 positively correlated and 160 negatively correlated with age - across 51 tissue types, demonstrating remarkable cross-tissue applicability that extended even to in vitro analyses and other mammalian species 21512. Hannum's clock utilized 71 CpG sites specific to whole blood derived from adult populations 1512. While these clocks exhibit a high Pearson correlation with chronological age (often exceeding r = 0.96 with a mean absolute error of roughly 3 to 4 years), they possess an inherent algorithmic limitation 412. Because they are trained to perfectly mirror chronological time, any deviation in the form of epigenetic age acceleration is mathematically treated as an error by the algorithm rather than a true biological signal 24. Consequently, they are inherently less sensitive to lifestyle interventions, underlying morbidity risk, and the dynamic, variable aspects of aging 2515.
Second-Generation Clocks: Phenotypic and Mortality Predictors
Recognizing the limitations of chronological training, researchers developed second-generation clocks - such as PhenoAge and GrimAge - which are calibrated against physiological morbidity, composite healthspan indicators, and all-cause mortality rather than age alone 61517. PhenoAge incorporates 513 CpG sites associated with nine clinical biomarkers of physiological dysregulation (including albumin, creatinine, and C-reactive protein), offering superior predictions for age-related diseases and physical frailty 213. GrimAge, and its successor GrimAge2, represent a more sophisticated multi-omic approach by utilizing DNA methylation surrogates for plasma proteins - such as PAI-1, leptin, cystatin C, and adrenomedullin - as well as self-reported smoking pack-years 2617. Across vast epidemiological cohorts, GrimAge has consistently outperformed first-generation clocks in predicting cardiovascular disease, cancer onset, and mortality independent of traditional risk factors like BMI or socioeconomic status 151719.
Third-Generation Clocks: Quantifying the Pace of Aging
The most recent paradigm shift is represented by third-generation measures like DunedinPACE. Instead of estimating a static biological age in years, DunedinPACE fundamentally measures the current velocity of aging 1713. Developed from the Dunedin longitudinal birth cohort study, it tracks the rate of physiological decline across 19 organ function biomarkers measured repeatedly over several decades 17. The algorithm then isolates the specific methylation signature associated with faster or slower systemic decline. It outputs a score representing biological years aged per calendar year; for example, a score of 0.85 indicates the individual is aging 15% slower than the chronological average, whereas a score of 1.2 denotes a 20% acceleration 1720. This pace-of-aging approach is highly responsive to short-to-medium-term perturbations, making it currently the most clinically useful tool for tracking the efficacy of lifestyle modifications or evaluating gerotherapeutics in trial settings 151721.
Table 1: Comparison of Epigenetic Clock Generations (2024-2025 Consensus)
| Generation | Example Clocks | Primary Tissue Source | CpG Site Count | Primary Training Target | 2024/2025 Clinical & Research Accuracy Profile |
|---|---|---|---|---|---|
| First Generation | Horvath Pan-Tissue, Hannum | Multi-tissue (Horvath), Whole Blood (Hannum) | 353 (Horvath), 71 (Hannum) | Chronological Age | High accuracy for chronological age estimation (r > 0.96). Lower sensitivity to capturing healthspan, predicting disease risk, or responding to specific behavioral interventions. 111512 |
| Second Generation | PhenoAge, GrimAge, GrimAge2, CheekAge | Blood, Buccal (CheekAge) | 513 (PhenoAge), 1,030 (GrimAge) | Mortality, Clinical Biomarkers, Healthspan | Superior prediction of all-cause mortality, cardiovascular disease, and metabolic dysfunction. Highly effective for epidemiological disease forecasting. 2151713 |
| Third Generation | DunedinPACE | Blood, Saliva, Buccal | 173 | Velocity/Pace of Aging (Longitudinal decline) | Measures instantaneous rate of aging (biological years per calendar year). Highest sensitivity to caloric restriction and lifestyle interventions in clinical trials. 1517 |
Tissue Specificity and the "Tissue Issue" in Methylation Assessment
The accuracy and utility of epigenetic clocks are intrinsically tied to the tissue from which the DNA is extracted. Epigenetic markers dictate cellular differentiation; therefore, DNA methylation patterns are highly tissue-specific 622. Studies analyzing human samples from the Genotype-Tissue Expression (GTEx) project across nine distinct tissue types - including lung, colon, prostate, ovary, breast, kidney, testis, skeletal muscle, and whole blood - reveal that mean DNAm age estimates vary substantially across tissues even within the exact same individual 1123.
This variability creates a significant "tissue issue" when clocks trained on one biological medium are applied to another. While Horvath's original clock was designed as a pan-tissue estimator, subsequent, more powerful clocks like Hannum, PhenoAge, and GrimAge were trained almost exclusively on immune cells derived from whole blood 1123. The application of these blood-derived epigenetic clocks to oral-based tissues - such as saliva or buccal epithelial swabs - frequently results in extreme estimation errors. Recent comparative research assessing paired samples from 83 individuals demonstrated that applying blood-based clocks to buccal tissue can yield discrepancies of almost 30 years in biological age estimation, even after controlling for cellular proportions and other technical covariates 1324.
Buccal cells evaluated using PhenoAge or GrimAge show vast overestimations for young samples, rendering the outputs effectively meaningless for health tracking 20. To counter this, novel clocks like the saliva-specific 10-CpG clock, PedBE (for pediatric buccal samples), and CheekAge (a next-generation buccal clock trained on lifestyle factors) have been developed, yet achieving perfect parity across tissue domains remains an ongoing methodological challenge 121325. In forensic and clinical settings, failing to match the clock algorithm strictly to its training tissue will generate systemic bias and false data 323.
Overcoming Technical Noise: The Paradigm of Principal Component (PC) Clocks
While the predictive power of epigenetic clocks at the population level is well-established, their utility at the individual level has been historically hindered by profound measurement unreliability 72627. Conventional CpG-based clocks suffer from substantial technical noise inherent to microarray platforms (such as the Illumina Infinium EPIC arrays) and the biological volatility of the methylome under normal homeostatic stress 4728. In technical replicates - where the exact same blood sample is divided, processed on different arrays, and tested twice - first-generation clocks have exhibited extreme random deviations, frequently generating age differences ranging from 3 to 9 years 42627. In longitudinal studies, this noise creates false-positive and false-negative results, leading researchers to erroneously conclude that an intervention successfully reversed aging when the shift was purely a statistical artifact 57.
To resolve this critical flaw, researchers developed Principal Component (PC) clocks (e.g., PC-Horvath, PC-PhenoAge, PC-GrimAge) in 2022 and 2023, which now represent the standard for robust longitudinal analysis 32714. Instead of relying on the raw, noisy methylation beta-values of individual CpG sites, this advanced methodological framework utilizes Principal Component Analysis (PCA) to extract the shared, underlying covariance across tens of thousands of highly collinear CpG sites 141516. Because technical microarray noise and batch effects are typically random and uncorrelated across different probes, the PCA mathematical smoothing effectively filters them out, capturing only the robust, genuine age-related biological signal 151718.
The integration of PC clocks has revolutionized the statistical reliability of longitudinal aging research. Studies utilizing Intraclass Correlation Coefficients (ICC) - a rigorous metric of test-retest reliability - demonstrate that while traditional clocks often score poorly, PC clocks routinely achieve ICCs greater than 0.90, indicating excellent, clinical-grade stability 2734.
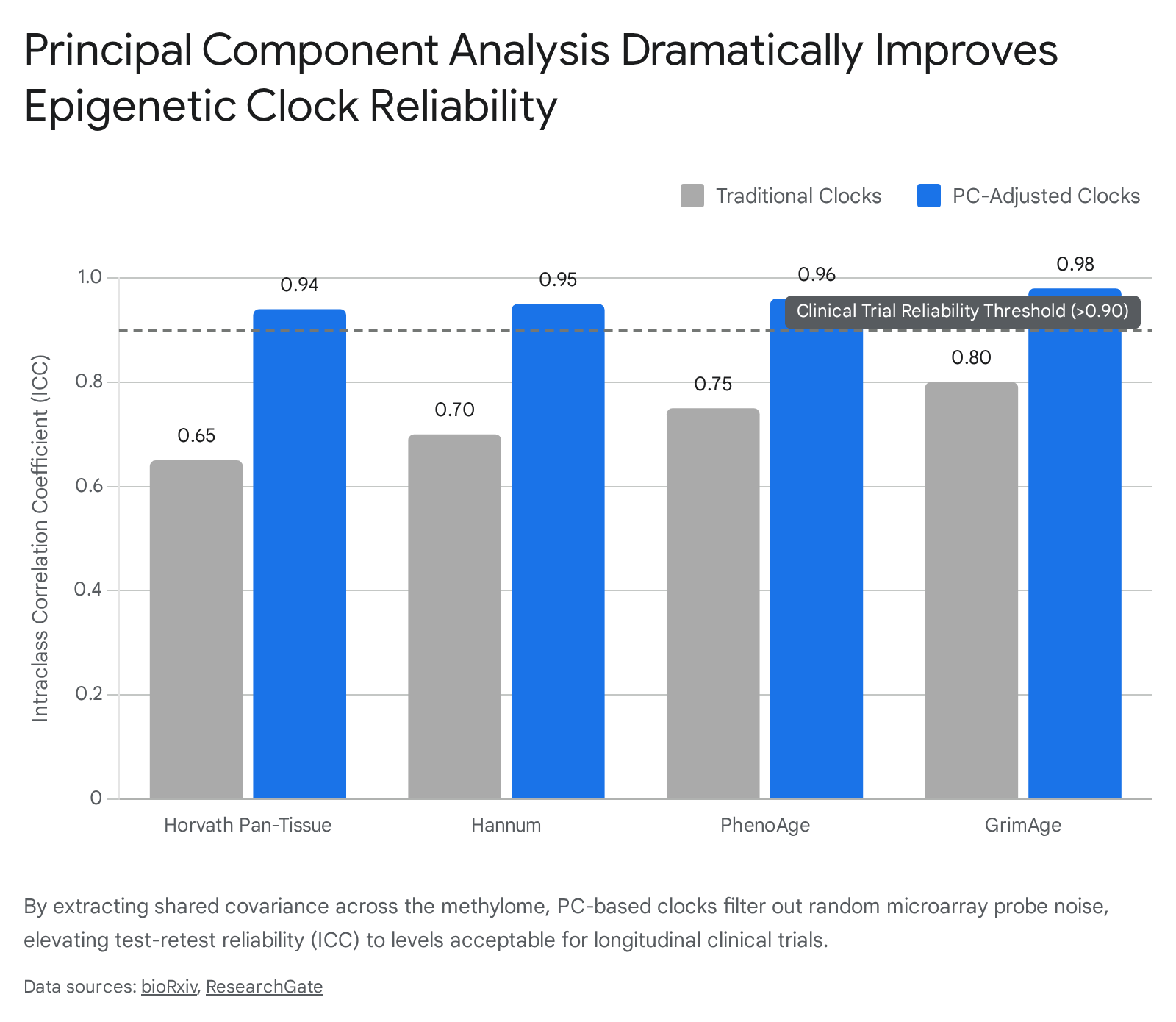
By drastically reducing intra-individual longitudinal variability, PC clocks enhance the statistical power to detect genuine biological changes over time. Pre-trial power analyses dictate that replacing standard clocks with PC clocks reduces the requisite sample size for clinical aging interventions by up to 10-fold 71734. Consequently, 2024 and 2025 consensus guidelines strongly recommend that any interventional trial seeking to validate a gerotherapeutic must utilize PC-adjusted clocks or mathematically stable third-generation metrics like DunedinPACE to avoid reporting illusory age reversals 5718.
High-Resolution Profiling: The Emergence of Single-Cell Epigenetic Clocks
Despite the mathematical improvements provided by PCA, traditional clocks share a fundamental biological limitation: they rely entirely on bulk tissue sequencing. A standard blood sample contains a highly heterogeneous mixture of cells - CD4+ T cells, CD8+ T cells, B cells, monocytes, and natural killer cells - each possessing unique methylation landscapes and aging at vastly divergent trajectories 22219. Bulk sequencing averages these disparate signals together, obscuring vital cell-intrinsic aging processes and making it difficult to determine whether a change in a patient's biological age is due to true molecular rejuvenation or merely a systemic shift in the relative proportions of different immune cells (e.g., a sudden influx of young, naive T cells following a viral infection) 819.
To achieve unprecedented granularity, researchers in 2024 and 2025 successfully developed single-cell epigenetic and transcriptomic clocks, representing a massive technological leap forward. The primary obstacle to single-cell methylation clocks was the intrinsically sparse and binarized nature of single-cell DNAm profiles. Because individual diploid cells only have two alleles, any specific CpG site is either 0%, 50%, or 100% methylated 820. Traditional bulk clocks assume continuous methylation values (ranging from 0 to 1) averaged across millions of cells; when applied directly to single cells, these standard linear regression models entirely fail 810.
Advanced computational frameworks like scAge and estiMAge have successfully bypassed this limitation 810. The scAge algorithm trains a linear model on bulk tissue reference data to establish a probabilistic baseline, then intersects this framework with the sparse reads of a single cell to compute a comprehensive likelihood profile across a range of ages. It then assigns the age of maximum likelihood as the cell's final epigenetic age 8. This has yielded remarkable insights previously hidden from gerontologists: scAge revealed that muscle stem cells exhibit severely attenuated epigenetic aging compared to surrounding somatic tissue, and it successfully tracked a natural, highly stratified cellular rejuvenation event that occurs during early mouse embryogenesis during the gastrulation phase 820. Furthermore, frameworks like estiMAge utilize imputation to reconstruct missing CpG values by exploiting redundancy within the methylome, allowing traditional bulk clocks to maintain predictive capability down to the single-hepatocyte level 10.
Simultaneously, single-cell transcriptomic and multi-omic immune clocks like sc-ImmuAging and scAgeClock have mapped the distinct aging trajectories of peripheral blood mononuclear cells 2213839. Research utilizing sc-ImmuAging (which analyzed cell types from over 1,000 healthy individuals) demonstrated that CD8+ naive T cells exhibit an epigenetic and transcriptomic age up to 20 years younger than effector memory T cells within the exact same individual 1521. Furthermore, these high-resolution models have uncovered precise, dynamic clinical events: severe COVID-19 infection induces profound, temporary age acceleration specifically localized within the monocyte population, while BCG vaccination can trigger distinct transcriptomic rejuvenation signatures within CD8+ T cells in patients with strong baseline interferon responses 21938. By isolating systemic composition shifts from true intrinsic cellular aging, single-cell clocks are poised to become the ultimate precision tool for evaluating highly targeted longevity therapeutics at the molecular level.
Dataset Bias and the Generalizability Crisis in Non-Western Demographics
As the clinical and commercial footprint of epigenetic clocks expands, a severe epidemiological limitation has been exposed: foundational epigenetic clocks suffer from profound dataset bias, exhibiting alarmingly poor generalizability outside of Western demographics 62522. Standard models, including Horvath's Pan-Tissue, Hannum, and early PhenoAge iterations, were trained predominantly on DNA methylation data derived from individuals of European and, to a lesser extent, Hispanic ancestry 222342.
When these Euro-centric clocks are applied to genetically diverged cohorts - including African, African-American, and East Asian populations - they frequently produce erratic and highly discordant age acceleration estimates 62342. For instance, multiple studies utilizing second-generation clocks routinely report that Black Americans exhibit significantly accelerated biological aging (higher GrimAge and DunedinPoAm scores) compared to White Americans, while simultaneously showing lower Hannum age acceleration 6. This acceleration phenomenon persists even after strictly controlling for socioeconomic status, access to education, and poverty levels 6. While some researchers previously hypothesized this strictly reflected the biological embedding of systemic discrimination and lifetime psychosocial stress, recent deep-genomic analyses from 2024 and 2025 reveal a starkly different primary driver: uncorrected genetic confounding 2823.
The underlying mechanism of this dataset bias is the prevalence of Methylation Quantitative Trait Loci (meQTLs) - common single-nucleotide polymorphisms (cSNPs) in the genome that structurally alter or dictate the baseline methylation levels of adjacent CpG sites 62823. Extensive genome-wide association studies (GWAS) have demonstrated that between 22% and 76% of the specific CpG sites utilized in standard epigenetic clocks are directly influenced by known meQTLs 2842. Because the allele frequencies of these meQTLs vary significantly across global populations due to deep ancestral divergence and varying degrees of heterozygosity, an individual of African or East Asian descent might possess a genetic variant that naturally elevates baseline methylation at a specific clock CpG site. The Euro-centric algorithmic model, entirely blind to this genetic variation, misinterprets the inherently high methylation level as severe "accelerated aging" 2342. As Gopalan et al. (2026) articulated following analyses of African populations (including the Himba, Baka, and San cohorts), "the same level of DNA methylation can translate to a very different age prediction depending on your genotype" 2342.
Recognizing that applying flawed clocks to diverse populations risks entrenching healthcare disparities and misdirecting clinical care, the scientific community is rapidly attempting to correct this ancestral bias 624. Solutions currently underway include the development of deep neural network (DNN) transfer learning frameworks designed to mathematically adapt existing clocks to underrepresented cohorts 25. More fundamentally, researchers are now actively excluding known cis-meQTL CpG sites during the algorithmic training phase to ensure the resulting clock captures purely epigenetic phenomena and remains ancestry-agnostic 6. Furthermore, efforts are yielding highly accurate, population-specific models. For example, the EAS clock and iCAS-DNAmAge have been trained explicitly on East Asian cohorts (South Korean, Taiwanese, and Han Chinese individuals) to provide culturally and genetically calibrated health predictors, successfully demonstrating that accurate localized modeling can vastly outperform general models in detecting diabetes and hypertension risks in these demographics 2244.
Evaluating Interventional Studies: Biomarker Gaming Versus Systemic Reversal
The ultimate aspiration of the longevity biotechnology sector is to utilize epigenetic clocks as surrogate endpoints in clinical trials, thereby avoiding the decades-long observational wait required to prove actual lifespan extension in human subjects. However, evaluating the success of these interventional studies requires an intense, critical distinction between genuinely reversing systemic biological aging and simply "gaming" a highly specific biomarker score.
This tension is perfectly encapsulated by the Thymus Regeneration, Immunorestoration, and Insulin Mitigation (TRIIM) trial, and its ongoing extension, TRIIM-X. Led by Dr. Greg Fahy, the original 2019 TRIIM study administered a targeted gerotherapeutic cocktail of recombinant human growth hormone (rhGH), dehydroepiandrosterone (DHEA), and metformin to nine healthy older men for 12 months 254647. The results were heralded as a historic biomedical breakthrough: participants demonstrated significant thymic fat replacement by functional tissue, broadly improved immune profiles, and a striking reduction in epigenetic age of approximately 2.5 years across four distinct clocks (Horvath, Hannum, PhenoAge, GrimAge), a finding that persisted six months post-trial 254748. The ongoing TRIIM-X trial (NCT04375657), which expanded the cohort to include women, a broader age range (40-80), and an active control group, has reportedly reproduced these age-reversal findings across 21 different epigenetic clock models, alongside observed reductions in systemic inflammation markers like C-reactive protein 47495051.
Despite the widespread enthusiasm, rigorous methodological critiques have been levied against interpreting these results as definitive "systemic age reversal." The primary concern is the phenomenon of "teaching to the test" - a manifestation of Goodhart's Law, which dictates that when a measure becomes a target, it ceases to be a good measure 265354. Because first- and second-generation clocks rely heavily on immune cell composition data - particularly the ratio of naive CD8+ T cells to terminally differentiated effector memory T cells - interventions like the TRIIM protocol that specifically stimulate thymic output will inevitably alter the exact cell populations the clocks measure 1927. While regenerating the thymus and improving immune profiles is undeniably a positive health outcome that decreases mortality risk, critics argue it represents the localized repair of a specific downstream symptom of aging, not the reversal of the foundational, systemic organismal aging process itself 27.
This distinction is crucial for the future of longevity medicine. If a patient takes a pharmaceutical that artificially alters the methylation state of 50 specific CpG sites utilized by a clock, their biological age score will plummet. However, if their vascular stiffness, mitochondrial dysfunction, amyloid-beta accumulation, and proteostatic collapse remain entirely unaddressed, their true systemic aging trajectory remains unaltered 27. In contrast, large-scale lifestyle interventions, such as the CALERIE trial evaluating continuous caloric restriction, utilize third-generation pace-of-aging metrics (DunedinPACE) alongside expansive biomarker panels to prove generalized, multi-systemic deceleration 2128. The consensus among leading gerontologists is shifting away from the hyperbole of "full reversibility" and "youth restoration" 2757. Instead, the scientific reality is that aging is a dynamic, modifiable condition that can be decelerated, attenuated, or partially mitigated 27. Future interventional trials must rely on expansive multi-omic approaches - combining epigenetic pace-of-aging clocks, comprehensive proteomic panels, metabolomics, functional physical assessments, and single-cell readouts - to definitively prove that an intervention is altering the underlying molecular drivers of aging rather than merely superficially manipulating a proxy biomarker equation 17212153.
The Direct-to-Consumer Market Versus Clinical and Research Utility
The rapid evolution and media popularization of epigenetic clocks have catalyzed a highly lucrative Direct-to-Consumer (DTC) longevity market. Dozens of commercial entities are aggressively commercializing biological age tests, frequently bundling them with proprietary lifestyle coaching, nutritional supplements, and generalized "biohacking" protocols marketed as capable of reversing aging 582458. However, peer-reviewed consensus from researchers and physicians strictly warns that while these tools are profoundly useful at the epidemiological research level, they currently fail to meet the rigorous reproducibility and accuracy standards required for individual clinical diagnostics or short-term consumer health tracking 482429.
A primary public misconception perpetuated by DTC marketing is that consumers can utilize commercially available biological age tests to track short-term, month-to-month health changes following a new diet, exercise routine, or supplement regimen. This is scientifically unfounded and practically impossible. Due to the inherent biological volatility of the methylome and the standard error of the predictive algorithms (typically ± 3 to 5 years), it requires a minimum of 6 to 12 months for genuine, intervention-driven epigenetic changes to meaningfully separate from background technical noise 17. Furthermore, DTC companies routinely encounter severe logistical barriers, most notably the "tissue issue" 22. To maximize consumer convenience and minimize shipping restrictions, commercial tests frequently rely on easily obtained biological samples like saliva or buccal (cheek) swabs. However, they subsequently analyze this DNA using foundational algorithms (like the standard GrimAge or PhenoAge) that were trained exclusively on blood 4. Because different tissues age at vastly different rates and possess distinct epigenetic profiles, applying a blood-based algorithm to buccal tissue yields wildly inaccurate and effectively meaningless age estimates - sometimes generating biological age discrepancies of 20 to 30 years from reality 3424.
Beyond severe technical inaccuracies, the rapid commercialization of these tests raises significant ethical, legal, and social issues (ELSI). In the absence of clinical context, professional medical guidance, or universally established diagnostic cut-offs, returning a probabilistic "accelerated aging" score to a consumer can induce severe psychological distress and health anxiety 45830. It risks pathologizing normal biological variation and driving individuals toward unproven, out-of-pocket, over-medicalized wellness interventions 45830. Moreover, because epigenetic clocks explicitly capture the biological embedding of environmental stress, toxic exposure, and socio-economic deprivation, legal and policy experts fear the data could eventually be weaponized by life insurance underwriters or employers. Using epigenetic aging scores to set insurance premiums or assess insurability would effectively penalize marginalized communities for structural disadvantages and life-course adversities beyond their control, transforming deep societal inequity into a personalized financial liability 4824.
Table 2: Clinical and Research Use Cases vs. Direct-to-Consumer (DTC) Applications
| Context | Key Applications | Methodological Strengths & Valid Uses | Limitations & Critical Risks |
|---|---|---|---|
| Research & Population Studies | Epidemiological forecasting, evaluating environmental exposures, understanding social determinants of health. | Excellent at predicting population-level mortality and disease risk. Highly effective at uncovering how trauma, poverty, and pollution biologically embed over time. 3424 | Subject to severe dataset bias if applied to non-European cohorts without algorithmic adjustment. Captures broad correlations rather than individualized causal mechanisms. 624 |
| Clinical Trials (Gerotherapeutics) | Assessing the longitudinal efficacy of anti-aging interventions (e.g., TRIIM-X, rapamycin, caloric restriction). | Valid and highly useful when utilizing PC-adjusted clocks or DunedinPACE over long durations (12+ months) to overcome statistical noise. 71727 | Risk of "biomarker gaming" - altering specific CpGs or immune cell ratios without genuinely reversing systemic physiological decline. 52627 |
| Clinical Diagnostics (Individual Care) | Personalized medicine, stratifying patient risk for specific age-related morbidities alongside traditional labs. | Currently strictly research-use-only. Strong potential for future integration alongside standard clinical biomarkers (e.g., lipid panels, HbA1c) to inform preventative care. 5861 | Fails standard clinical diagnostic criteria for reproducibility. High intra-individual variability. Lacks universally established "cut-off" thresholds for pathology. High cost ($240-$512 vs standard $10 tests). 424 |
| Direct-to-Consumer (DTC) Market | Commercial wellness tracking, biohacking, and proprietary supplement marketing. | Accessible introduction to longevity science for the general public. Can serve as a powerful behavioral motivational tool for adopting healthier lifestyle habits. 5862 | Severe limitations: Cannot reliably track short-term (<6 months) changes. Rampant tissue-mismatch errors (buccal DNA analyzed with blood clocks). Risks inducing severe health anxiety. Danger of future epigenetic insurance discrimination. 34817 |
Conclusion
Epigenetic clocks represent a monumental leap forward in our ability to quantify the biological wear-and-tear of human existence. The field's progression from simple chronological estimators to highly sophisticated, single-cell and principal component-adjusted mortality predictors highlights a rapid, transformative maturation. Tools like sc-ImmuAging and DunedinPACE have provided researchers with unprecedented molecular resolution, allowing gerontologists to peer into the distinct aging dynamics of individual cellular subtypes and accurately track the instantaneous velocity of human decline.
However, as the underlying science advances, so too must the rigorous scrutiny with which we interpret its computational outputs. The documented failure of foundational clocks to generalize across global, diverse populations highlights an urgent need to decouple genetic variance (meQTLs) from true epigenetic deterioration, ensuring that the diagnostic benefits of longevity science are equitably distributed across all demographics. Furthermore, as bold interventional trials like TRIIM-X continue to push the boundaries of medical possibility, researchers must remain deeply vigilant against the allure of "biomarker gaming," ensuring that experimental treatments genuinely reverse the complex, upstream systemic drivers of aging rather than merely cosmetically altering the inputs of a diagnostic test score. Ultimately, until the substantial technical noise, tissue specificities, and profound biological complexities of the methylome are fully resolved, epigenetic clocks must be treated with deep caution in the commercial consumer market, remaining securely within the domain of rigorous, long-term scientific inquiry.